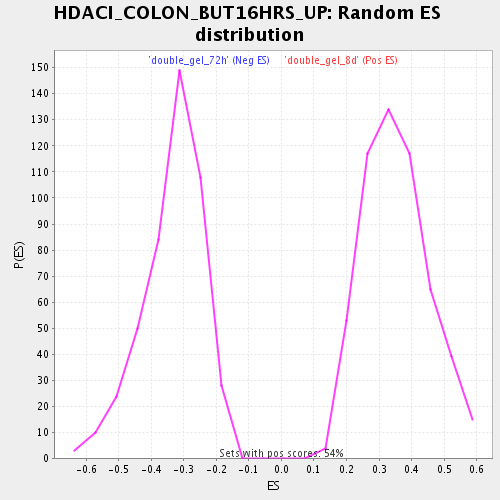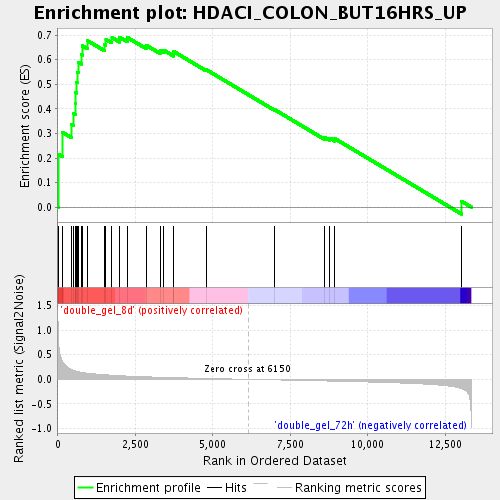
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HDACI_COLON_BUT16HRS_UP |
| Enrichment Score (ES) | 0.69151026 |
| Normalized Enrichment Score (NES) | 1.9663641 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.007877029 |
| FWER p-Value | 0.315 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 12 | 0.803 | 0.2149 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 142 | 0.371 | 0.3048 | Yes |
| 3 | GABBR1 | GABBR1 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 1 | 432 | 0.200 | 0.3370 | Yes |
| 4 | NEU1 | NEU1 Entrez, Source | sialidase 1 (lysosomal sialidase) | 510 | 0.181 | 0.3798 | Yes |
| 5 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 567 | 0.170 | 0.4213 | Yes |
| 6 | IL17RE | IL17RE Entrez, Source | interleukin 17 receptor E | 574 | 0.168 | 0.4661 | Yes |
| 7 | ZSWIM6 | ZSWIM6 Entrez, Source | zinc finger, SWIM-type containing 6 | 600 | 0.165 | 0.5085 | Yes |
| 8 | BIRC4 | BIRC4 Entrez, Source | baculoviral IAP repeat-containing 4 | 630 | 0.161 | 0.5495 | Yes |
| 9 | RHBDF1 | RHBDF1 Entrez, Source | rhomboid 5 homolog 1 (Drosophila) | 661 | 0.156 | 0.5893 | Yes |
| 10 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 756 | 0.144 | 0.6210 | Yes |
| 11 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 785 | 0.141 | 0.6567 | Yes |
| 12 | PTPN1 | PTPN1 Entrez, Source | protein tyrosine phosphatase, non-receptor type 1 | 959 | 0.124 | 0.6772 | Yes |
| 13 | ROCK1 | ROCK1 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 1 | 1513 | 0.093 | 0.6607 | Yes |
| 14 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 1550 | 0.092 | 0.6826 | Yes |
| 15 | CALCOCO1 | CALCOCO1 Entrez, Source | calcium binding and coiled-coil domain 1 | 1744 | 0.083 | 0.6904 | Yes |
| 16 | SPARCL1 | SPARCL1 Entrez, Source | SPARC-like 1 (mast9, hevin) | 1992 | 0.073 | 0.6915 | Yes |
| 17 | INPP5E | INPP5E Entrez, Source | inositol polyphosphate-5-phosphatase, 72 kDa | 2239 | 0.066 | 0.6908 | No |
| 18 | CRYL1 | CRYL1 Entrez, Source | crystallin, lambda 1 | 2849 | 0.052 | 0.6590 | No |
| 19 | CLU | CLU Entrez, Source | clusterin | 3299 | 0.043 | 0.6367 | No |
| 20 | MAP1LC3B | MAP1LC3B Entrez, Source | microtubule-associated protein 1 light chain 3 beta | 3406 | 0.040 | 0.6396 | No |
| 21 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 3736 | 0.035 | 0.6242 | No |
| 22 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 3739 | 0.035 | 0.6333 | No |
| 23 | DNM3 | DNM3 Entrez, Source | dynamin 3 | 4784 | 0.018 | 0.5599 | No |
| 24 | GNL1 | GNL1 Entrez, Source | guanine nucleotide binding protein-like 1 | 6990 | -0.011 | 0.3971 | No |
| 25 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 8603 | -0.032 | 0.2847 | No |
| 26 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 8778 | -0.035 | 0.2809 | No |
| 27 | TSNAX | TSNAX Entrez, Source | translin-associated factor X | 8938 | -0.037 | 0.2789 | No |
| 28 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 13029 | -0.193 | 0.0235 | No |