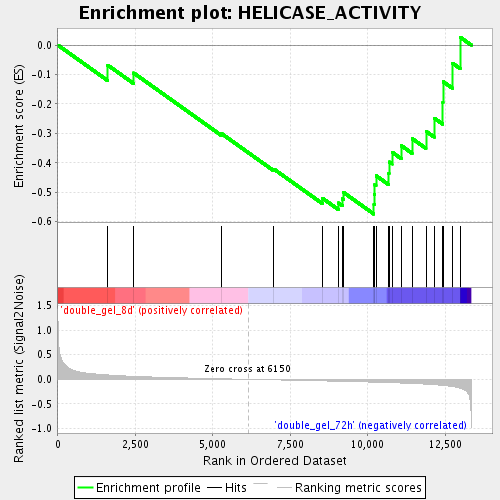
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | HELICASE_ACTIVITY |
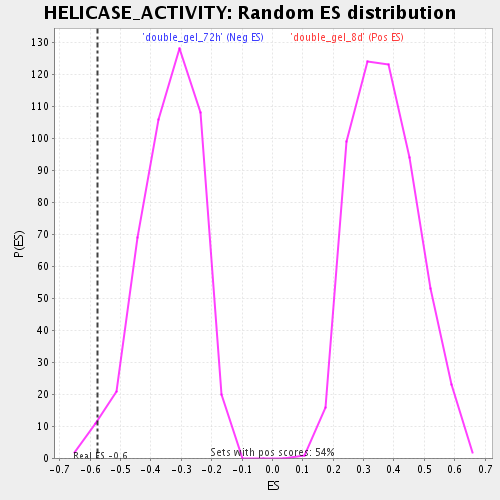
| Enrichment Score (ES) | -0.5744695 |
| Normalized Enrichment Score (NES) | -1.7007769 |
| Nominal p-value | 0.017204301 |
| FDR q-value | 0.17120112 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 1606 | 0.089 | -0.0679 | No |
| 2 | DDX25 | DDX25 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 25 | 2444 | 0.061 | -0.0946 | No |
| 3 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 5284 | 0.011 | -0.3010 | No |
| 4 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 6971 | -0.010 | -0.4215 | No |
| 5 | RECQL | RECQL Entrez, Source | RecQ protein-like (DNA helicase Q1-like) | 8536 | -0.031 | -0.5204 | No |
| 6 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 9054 | -0.038 | -0.5365 | No |
| 7 | DHX8 | DHX8 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 8 | 9187 | -0.040 | -0.5226 | No |
| 8 | SKIV2L | SKIV2L Entrez, Source | superkiller viralicidic activity 2-like (S. cerevisiae) | 9215 | -0.040 | -0.5007 | No |
| 9 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 10199 | -0.057 | -0.5409 | Yes |
| 10 | IGHMBP2 | IGHMBP2 Entrez, Source | immunoglobulin mu binding protein 2 | 10212 | -0.057 | -0.5081 | Yes |
| 11 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 10227 | -0.057 | -0.4752 | Yes |
| 12 | DHX16 | DHX16 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 16 | 10268 | -0.058 | -0.4439 | Yes |
| 13 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 10677 | -0.066 | -0.4355 | Yes |
| 14 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 10688 | -0.066 | -0.3972 | Yes |
| 15 | DDX56 | DDX56 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 56 | 10784 | -0.068 | -0.3640 | Yes |
| 16 | DDX20 | DDX20 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 | 11083 | -0.075 | -0.3417 | Yes |
| 17 | DDX18 | DDX18 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 18 | 11433 | -0.085 | -0.3178 | Yes |
| 18 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 11880 | -0.098 | -0.2929 | Yes |
| 19 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 12144 | -0.109 | -0.2480 | Yes |
| 20 | SUPV3L1 | SUPV3L1 Entrez, Source | suppressor of var1, 3-like 1 (S. cerevisiae) | 12423 | -0.123 | -0.1963 | Yes |
| 21 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 12430 | -0.123 | -0.1240 | Yes |
| 22 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12731 | -0.144 | -0.0611 | Yes |
| 23 | ASCC3L1 | ASCC3L1 Entrez, Source | activating signal cointegrator 1 complex subunit 3-like 1 | 12985 | -0.180 | 0.0268 | Yes |