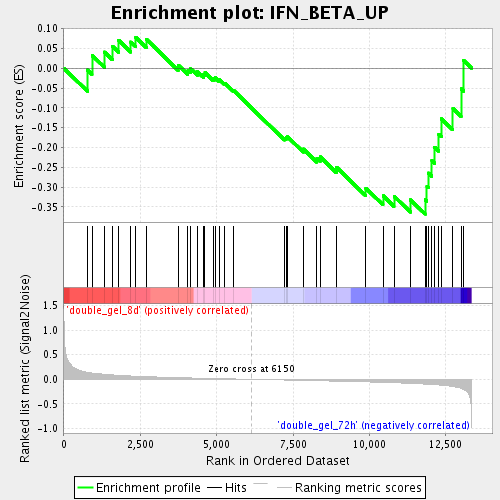
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | IFN_BETA_UP |
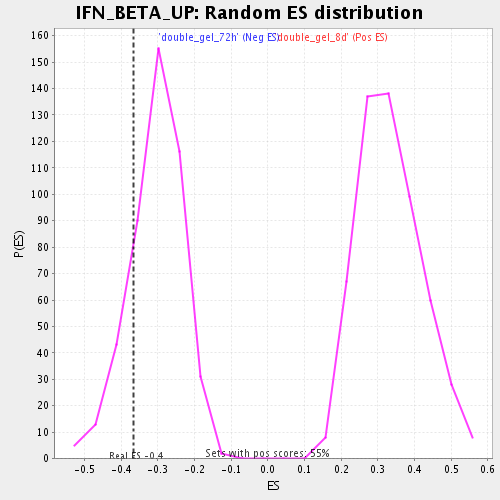
| Enrichment Score (ES) | -0.3678996 |
| Normalized Enrichment Score (NES) | -1.2095292 |
| Nominal p-value | 0.17802198 |
| FDR q-value | 0.6434348 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 772 | 0.143 | -0.0041 | No |
| 2 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 920 | 0.127 | 0.0329 | No |
| 3 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 1327 | 0.103 | 0.0411 | No |
| 4 | BCLAF1 | BCLAF1 Entrez, Source | BCL2-associated transcription factor 1 | 1588 | 0.090 | 0.0554 | No |
| 5 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1791 | 0.081 | 0.0708 | No |
| 6 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 2190 | 0.067 | 0.0663 | No |
| 7 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 2350 | 0.063 | 0.0783 | No |
| 8 | CASP8 | CASP8 Entrez, Source | caspase 8, apoptosis-related cysteine peptidase | 2697 | 0.055 | 0.0732 | No |
| 9 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 3738 | 0.035 | 0.0081 | No |
| 10 | PPP5C | PPP5C Entrez, Source | protein phosphatase 5, catalytic subunit | 4060 | 0.029 | -0.0050 | No |
| 11 | OAS1 | OAS1 Entrez, Source | 2',5'-oligoadenylate synthetase 1, 40/46kDa | 4150 | 0.028 | -0.0011 | No |
| 12 | CSRP3 | CSRP3 Entrez, Source | cysteine and glycine-rich protein 3 (cardiac LIM protein) | 4369 | 0.024 | -0.0084 | No |
| 13 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 4566 | 0.022 | -0.0150 | No |
| 14 | CD164 | CD164 Entrez, Source | CD164 molecule, sialomucin | 4616 | 0.021 | -0.0108 | No |
| 15 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 4892 | 0.017 | -0.0251 | No |
| 16 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 4953 | 0.016 | -0.0237 | No |
| 17 | EIF2AK2 | EIF2AK2 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 2 | 5091 | 0.014 | -0.0286 | No |
| 18 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 5268 | 0.012 | -0.0375 | No |
| 19 | AP3M2 | AP3M2 Entrez, Source | adaptor-related protein complex 3, mu 2 subunit | 5555 | 0.007 | -0.0562 | No |
| 20 | EPS15 | EPS15 Entrez, Source | epidermal growth factor receptor pathway substrate 15 | 7217 | -0.013 | -0.1760 | No |
| 21 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 7272 | -0.014 | -0.1748 | No |
| 22 | PPP3CA | PPP3CA Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, alpha isoform (calcineurin A alpha) | 7318 | -0.014 | -0.1728 | No |
| 23 | GMFB | GMFB Entrez, Source | glia maturation factor, beta | 7835 | -0.021 | -0.2035 | No |
| 24 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 8279 | -0.028 | -0.2264 | No |
| 25 | CTR9 | CTR9 Entrez, Source | Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 8385 | -0.029 | -0.2234 | No |
| 26 | ADAR | ADAR Entrez, Source | adenosine deaminase, RNA-specific | 8925 | -0.036 | -0.2501 | No |
| 27 | PTPN11 | PTPN11 Entrez, Source | protein tyrosine phosphatase, non-receptor type 11 (Noonan syndrome 1) | 9881 | -0.052 | -0.3024 | No |
| 28 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 10443 | -0.061 | -0.3215 | No |
| 29 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 10803 | -0.068 | -0.3226 | No |
| 30 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 11340 | -0.083 | -0.3317 | Yes |
| 31 | PARP4 | PARP4 Entrez, Source | poly (ADP-ribose) polymerase family, member 4 | 11822 | -0.096 | -0.3315 | Yes |
| 32 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 11880 | -0.098 | -0.2986 | Yes |
| 33 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 11925 | -0.100 | -0.2641 | Yes |
| 34 | MAP3K10 | MAP3K10 Entrez, Source | mitogen-activated protein kinase kinase kinase 10 | 12042 | -0.105 | -0.2334 | Yes |
| 35 | ZRF1 | ZRF1 Entrez, Source | zuotin related factor 1 | 12130 | -0.108 | -0.1989 | Yes |
| 36 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 12268 | -0.115 | -0.1659 | Yes |
| 37 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 12353 | -0.119 | -0.1274 | Yes |
| 38 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12731 | -0.144 | -0.1012 | Yes |
| 39 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 12999 | -0.184 | -0.0517 | Yes |
| 40 | FUBP1 | FUBP1 Entrez, Source | far upstream element (FUSE) binding protein 1 | 13083 | -0.205 | 0.0195 | Yes |