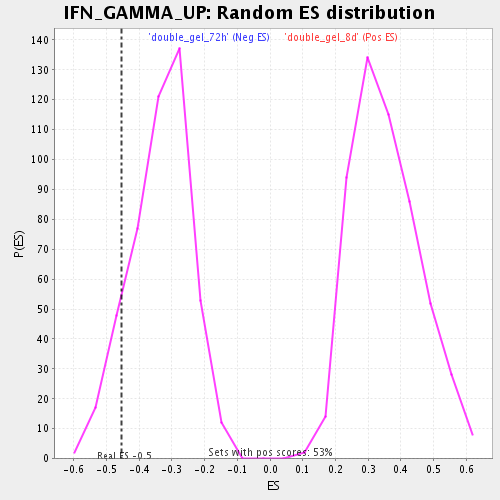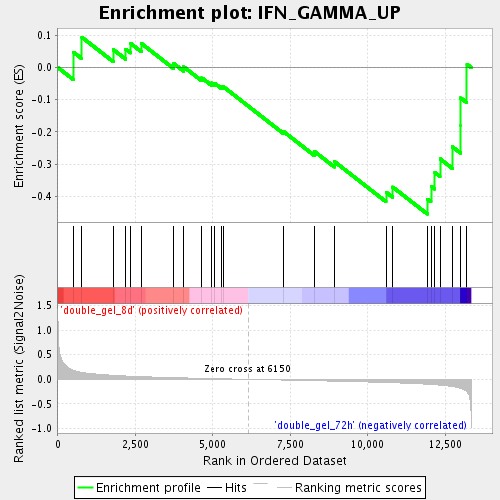
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | IFN_GAMMA_UP |
| Enrichment Score (ES) | -0.4551712 |
| Normalized Enrichment Score (NES) | -1.3560561 |
| Nominal p-value | 0.113490365 |
| FDR q-value | 0.4960169 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 496 | 0.183 | 0.0481 | No |
| 2 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 772 | 0.143 | 0.0942 | No |
| 3 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1791 | 0.081 | 0.0555 | No |
| 4 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 2190 | 0.067 | 0.0571 | No |
| 5 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 2350 | 0.063 | 0.0748 | No |
| 6 | CASP8 | CASP8 Entrez, Source | caspase 8, apoptosis-related cysteine peptidase | 2697 | 0.055 | 0.0746 | No |
| 7 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 3738 | 0.035 | 0.0127 | No |
| 8 | PPP5C | PPP5C Entrez, Source | protein phosphatase 5, catalytic subunit | 4060 | 0.029 | 0.0023 | No |
| 9 | BAK1 | BAK1 Entrez, Source | BCL2-antagonist/killer 1 | 4646 | 0.020 | -0.0321 | No |
| 10 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 4953 | 0.016 | -0.0478 | No |
| 11 | COL16A1 | COL16A1 Entrez, Source | collagen, type XVI, alpha 1 | 5051 | 0.015 | -0.0482 | No |
| 12 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 5268 | 0.012 | -0.0590 | No |
| 13 | EIF2B1 | EIF2B1 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 1 alpha, 26kDa | 5340 | 0.011 | -0.0594 | No |
| 14 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 7272 | -0.014 | -0.1980 | No |
| 15 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 8279 | -0.028 | -0.2606 | No |
| 16 | ADAR | ADAR Entrez, Source | adenosine deaminase, RNA-specific | 8925 | -0.036 | -0.2920 | No |
| 17 | BBC3 | BBC3 Entrez, Source | BCL2 binding component 3 | 10590 | -0.064 | -0.3871 | No |
| 18 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 10803 | -0.068 | -0.3710 | No |
| 19 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 11925 | -0.100 | -0.4085 | Yes |
| 20 | MAP3K10 | MAP3K10 Entrez, Source | mitogen-activated protein kinase kinase kinase 10 | 12042 | -0.105 | -0.3683 | Yes |
| 21 | PSME1 | PSME1 Entrez, Source | proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) | 12165 | -0.110 | -0.3261 | Yes |
| 22 | VAT1 | VAT1 Entrez, Source | vesicle amine transport protein 1 homolog (T californica) | 12333 | -0.118 | -0.2836 | Yes |
| 23 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12731 | -0.144 | -0.2460 | Yes |
| 24 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 12996 | -0.183 | -0.1802 | Yes |
| 25 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 12999 | -0.184 | -0.0943 | Yes |
| 26 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 13202 | -0.257 | 0.0105 | Yes |