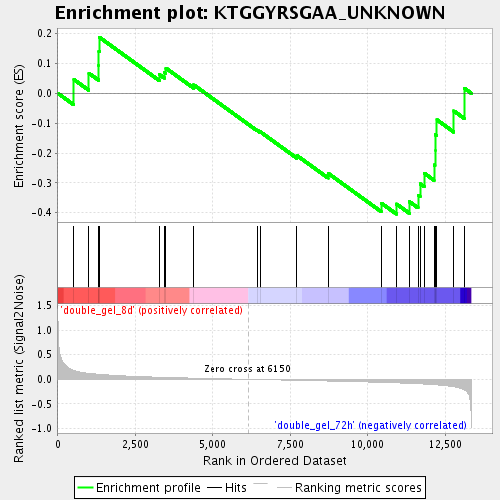
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | KTGGYRSGAA_UNKNOWN |
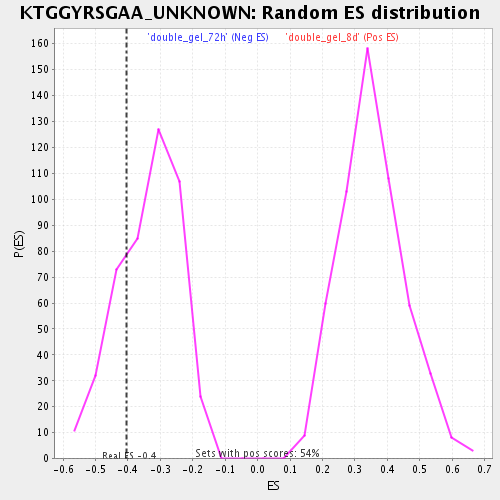
| Enrichment Score (ES) | -0.40460378 |
| Normalized Enrichment Score (NES) | -1.2029673 |
| Nominal p-value | 0.24183007 |
| FDR q-value | 0.64802706 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.0461 | No |
| 2 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 1001 | 0.121 | 0.0672 | No |
| 3 | DLGAP4 | DLGAP4 Entrez, Source | discs, large (Drosophila) homolog-associated protein 4 | 1317 | 0.103 | 0.0923 | No |
| 4 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 1323 | 0.103 | 0.1406 | No |
| 5 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1338 | 0.102 | 0.1878 | No |
| 6 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 3272 | 0.043 | 0.0630 | No |
| 7 | TUBA4 | TUBA4 Entrez, Source | tubulin, alpha 4 | 3435 | 0.040 | 0.0697 | No |
| 8 | GDI1 | GDI1 Entrez, Source | GDP dissociation inhibitor 1 | 3486 | 0.039 | 0.0843 | No |
| 9 | SH3KBP1 | SH3KBP1 Entrez, Source | SH3-domain kinase binding protein 1 | 4386 | 0.024 | 0.0281 | No |
| 10 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 6442 | -0.004 | -0.1245 | No |
| 11 | CNTFR | CNTFR Entrez, Source | ciliary neurotrophic factor receptor | 6526 | -0.005 | -0.1285 | No |
| 12 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 7715 | -0.020 | -0.2085 | No |
| 13 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 8719 | -0.034 | -0.2678 | No |
| 14 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 10434 | -0.061 | -0.3677 | No |
| 15 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10927 | -0.072 | -0.3708 | Yes |
| 16 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 11343 | -0.083 | -0.3629 | Yes |
| 17 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 11624 | -0.090 | -0.3416 | Yes |
| 18 | SEC61A1 | SEC61A1 Entrez, Source | Sec61 alpha 1 subunit (S. cerevisiae) | 11691 | -0.092 | -0.3033 | Yes |
| 19 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 11823 | -0.096 | -0.2676 | Yes |
| 20 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 12141 | -0.109 | -0.2399 | Yes |
| 21 | HTF9C | HTF9C Entrez, Source | - | 12171 | -0.110 | -0.1900 | Yes |
| 22 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12174 | -0.110 | -0.1379 | Yes |
| 23 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 12233 | -0.113 | -0.0890 | Yes |
| 24 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 12780 | -0.150 | -0.0591 | Yes |
| 25 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13114 | -0.214 | 0.0171 | Yes |