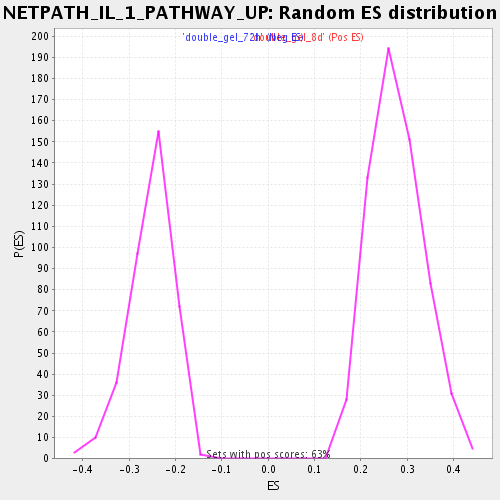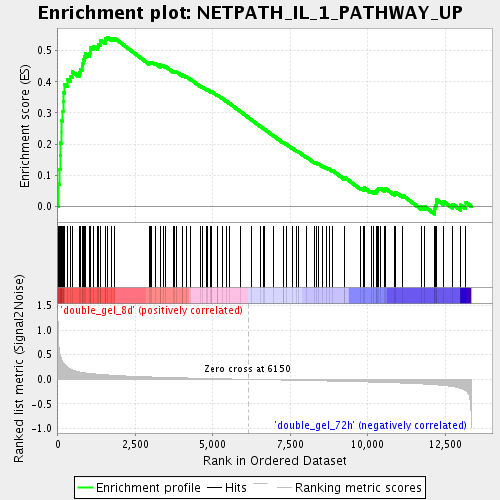
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | NETPATH_IL_1_PATHWAY_UP |
| Enrichment Score (ES) | 0.5428702 |
| Normalized Enrichment Score (NES) | 1.9559095 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.008621605 |
| FWER p-Value | 0.355 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 12 | 0.803 | 0.0725 | Yes |
| 2 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 61 | 0.555 | 0.1197 | Yes |
| 3 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 76 | 0.477 | 0.1622 | Yes |
| 4 | LOX | LOX Entrez, Source | lysyl oxidase | 82 | 0.465 | 0.2044 | Yes |
| 5 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 117 | 0.402 | 0.2387 | Yes |
| 6 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 118 | 0.402 | 0.2754 | Yes |
| 7 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 160 | 0.355 | 0.3047 | Yes |
| 8 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 171 | 0.348 | 0.3358 | Yes |
| 9 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 180 | 0.337 | 0.3660 | Yes |
| 10 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 219 | 0.308 | 0.3913 | Yes |
| 11 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 306 | 0.247 | 0.4074 | Yes |
| 12 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 414 | 0.206 | 0.4182 | Yes |
| 13 | JUN | JUN Entrez, Source | jun oncogene | 458 | 0.194 | 0.4326 | Yes |
| 14 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 685 | 0.152 | 0.4295 | Yes |
| 15 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 717 | 0.149 | 0.4408 | Yes |
| 16 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 785 | 0.141 | 0.4486 | Yes |
| 17 | GUCY1B3 | GUCY1B3 Entrez, Source | guanylate cyclase 1, soluble, beta 3 | 801 | 0.140 | 0.4602 | Yes |
| 18 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 825 | 0.137 | 0.4710 | Yes |
| 19 | STK39 | STK39 Entrez, Source | serine threonine kinase 39 (STE20/SPS1 homolog, yeast) | 864 | 0.133 | 0.4803 | Yes |
| 20 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 888 | 0.130 | 0.4904 | Yes |
| 21 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 1014 | 0.121 | 0.4920 | Yes |
| 22 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 1037 | 0.118 | 0.5012 | Yes |
| 23 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 1062 | 0.117 | 0.5100 | Yes |
| 24 | CCL7 | CCL7 Entrez, Source | chemokine (C-C motif) ligand 7 | 1159 | 0.111 | 0.5130 | Yes |
| 25 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 1285 | 0.105 | 0.5131 | Yes |
| 26 | MMP14 | MMP14 Entrez, Source | matrix metallopeptidase 14 (membrane-inserted) | 1312 | 0.104 | 0.5206 | Yes |
| 27 | HNMT | HNMT Entrez, Source | histamine N-methyltransferase | 1378 | 0.100 | 0.5248 | Yes |
| 28 | GJB1 | GJB1 Entrez, Source | gap junction protein, beta 1, 32kDa (connexin 32, Charcot-Marie-Tooth neuropathy, X-linked) | 1380 | 0.100 | 0.5338 | Yes |
| 29 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1541 | 0.092 | 0.5302 | Yes |
| 30 | ZFP36 | ZFP36 Entrez, Source | zinc finger protein 36, C3H type, homolog (mouse) | 1543 | 0.092 | 0.5385 | Yes |
| 31 | HMGCL | HMGCL Entrez, Source | 3-hydroxymethyl-3-methylglutaryl-Coenzyme A lyase (hydroxymethylglutaricaciduria) | 1594 | 0.089 | 0.5429 | Yes |
| 32 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 1734 | 0.083 | 0.5400 | No |
| 33 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 1832 | 0.079 | 0.5399 | No |
| 34 | IFNG | IFNG Entrez, Source | interferon, gamma | 2940 | 0.050 | 0.4608 | No |
| 35 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 2974 | 0.049 | 0.4628 | No |
| 36 | DCN | DCN Entrez, Source | decorin | 3029 | 0.048 | 0.4631 | No |
| 37 | CDK10 | CDK10 Entrez, Source | cyclin-dependent kinase (CDC2-like) 10 | 3149 | 0.045 | 0.4583 | No |
| 38 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 3298 | 0.043 | 0.4510 | No |
| 39 | CSF2 | CSF2 Entrez, Source | colony stimulating factor 2 (granulocyte-macrophage) | 3305 | 0.042 | 0.4544 | No |
| 40 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 3396 | 0.041 | 0.4513 | No |
| 41 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 3468 | 0.039 | 0.4495 | No |
| 42 | SERPINB3 | SERPINB3 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 3 | 3724 | 0.035 | 0.4334 | No |
| 43 | MMP3 | MMP3 Entrez, Source | matrix metallopeptidase 3 (stromelysin 1, progelatinase) | 3764 | 0.034 | 0.4336 | No |
| 44 | TNFRSF11B | TNFRSF11B Entrez, Source | tumor necrosis factor receptor superfamily, member 11b (osteoprotegerin) | 3828 | 0.033 | 0.4319 | No |
| 45 | GNB5 | GNB5 Entrez, Source | guanine nucleotide binding protein (G protein), beta 5 | 4018 | 0.030 | 0.4203 | No |
| 46 | CCL11 | CCL11 Entrez, Source | chemokine (C-C motif) ligand 11 | 4022 | 0.030 | 0.4228 | No |
| 47 | MMP10 | MMP10 Entrez, Source | matrix metallopeptidase 10 (stromelysin 2) | 4140 | 0.028 | 0.4166 | No |
| 48 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 4288 | 0.026 | 0.4078 | No |
| 49 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 4596 | 0.021 | 0.3865 | No |
| 50 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 4675 | 0.020 | 0.3825 | No |
| 51 | GAPDH | GAPDH Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase | 4797 | 0.018 | 0.3750 | No |
| 52 | AK3 | AK3 Entrez, Source | adenylate kinase 3 | 4820 | 0.018 | 0.3750 | No |
| 53 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 4934 | 0.016 | 0.3679 | No |
| 54 | ARSA | ARSA Entrez, Source | arylsulfatase A | 4941 | 0.016 | 0.3689 | No |
| 55 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 4953 | 0.016 | 0.3695 | No |
| 56 | MAPK10 | MAPK10 Entrez, Source | mitogen-activated protein kinase 10 | 5148 | 0.013 | 0.3561 | No |
| 57 | IL1A | IL1A Entrez, Source | interleukin 1, alpha | 5162 | 0.013 | 0.3563 | No |
| 58 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 5296 | 0.011 | 0.3473 | No |
| 59 | MMP12 | MMP12 Entrez, Source | matrix metallopeptidase 12 (macrophage elastase) | 5434 | 0.009 | 0.3378 | No |
| 60 | PIP5K1A | PIP5K1A Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, alpha | 5528 | 0.008 | 0.3315 | No |
| 61 | BMP1 | BMP1 Entrez, Source | bone morphogenetic protein 1 | 5892 | 0.004 | 0.3043 | No |
| 62 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 6260 | -0.001 | 0.2767 | No |
| 63 | LYPD3 | LYPD3 Entrez, Source | LY6/PLAUR domain containing 3 | 6537 | -0.005 | 0.2563 | No |
| 64 | UBD | UBD Entrez, Source | ubiquitin D | 6622 | -0.006 | 0.2505 | No |
| 65 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 6658 | -0.007 | 0.2485 | No |
| 66 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 6950 | -0.010 | 0.2274 | No |
| 67 | SHC1 | SHC1 Entrez, Source | SHC (Src homology 2 domain containing) transforming protein 1 | 7269 | -0.014 | 0.2047 | No |
| 68 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 7279 | -0.014 | 0.2053 | No |
| 69 | ACPP | ACPP Entrez, Source | acid phosphatase, prostate | 7380 | -0.015 | 0.1991 | No |
| 70 | GRAP2 | GRAP2 Entrez, Source | GRB2-related adaptor protein 2 | 7576 | -0.018 | 0.1860 | No |
| 71 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 7715 | -0.020 | 0.1773 | No |
| 72 | TMLHE | TMLHE Entrez, Source | trimethyllysine hydroxylase, epsilon | 7760 | -0.020 | 0.1759 | No |
| 73 | SOD3 | SOD3 Entrez, Source | superoxide dismutase 3, extracellular | 8030 | -0.024 | 0.1577 | No |
| 74 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 8279 | -0.028 | 0.1415 | No |
| 75 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 8340 | -0.028 | 0.1396 | No |
| 76 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 8408 | -0.029 | 0.1372 | No |
| 77 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 8552 | -0.031 | 0.1293 | No |
| 78 | HSP90AB1 | HSP90AB1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class B member 1 | 8666 | -0.033 | 0.1238 | No |
| 79 | C3 | C3 Entrez, Source | complement component 3 | 8749 | -0.034 | 0.1207 | No |
| 80 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 8852 | -0.035 | 0.1162 | No |
| 81 | GSTA3 | GSTA3 Entrez, Source | glutathione S-transferase A3 | 9247 | -0.041 | 0.0902 | No |
| 82 | NFKB1 | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 9252 | -0.041 | 0.0937 | No |
| 83 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 9770 | -0.050 | 0.0591 | No |
| 84 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 9869 | -0.051 | 0.0564 | No |
| 85 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 9884 | -0.052 | 0.0601 | No |
| 86 | C1R | C1R Entrez, Source | complement component 1, r subcomponent | 10107 | -0.055 | 0.0484 | No |
| 87 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 10192 | -0.057 | 0.0472 | No |
| 88 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10269 | -0.058 | 0.0467 | No |
| 89 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 10279 | -0.058 | 0.0514 | No |
| 90 | PRKCZ | PRKCZ Entrez, Source | protein kinase C, zeta | 10325 | -0.059 | 0.0534 | No |
| 91 | ARNT | ARNT Entrez, Source | aryl hydrocarbon receptor nuclear translocator | 10332 | -0.059 | 0.0583 | No |
| 92 | RPS21 | RPS21 Entrez, Source | ribosomal protein S21 | 10407 | -0.060 | 0.0582 | No |
| 93 | GTF2F1 | GTF2F1 Entrez, Source | general transcription factor IIF, polypeptide 1, 74kDa | 10529 | -0.063 | 0.0548 | No |
| 94 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 10574 | -0.064 | 0.0573 | No |
| 95 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 10870 | -0.070 | 0.0415 | No |
| 96 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10899 | -0.071 | 0.0458 | No |
| 97 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 11134 | -0.077 | 0.0351 | No |
| 98 | CFB | CFB Entrez, Source | complement factor B | 11731 | -0.093 | -0.0014 | No |
| 99 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 11841 | -0.097 | -0.0007 | No |
| 100 | ST3GAL4 | ST3GAL4 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 4 | 12158 | -0.110 | -0.0146 | No |
| 101 | PTPN21 | PTPN21 Entrez, Source | protein tyrosine phosphatase, non-receptor type 21 | 12166 | -0.110 | -0.0050 | No |
| 102 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 12195 | -0.111 | 0.0030 | No |
| 103 | CS | CS Entrez, Source | citrate synthase | 12219 | -0.112 | 0.0116 | No |
| 104 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 12229 | -0.113 | 0.0212 | No |
| 105 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 12451 | -0.124 | 0.0158 | No |
| 106 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 12736 | -0.145 | 0.0076 | No |
| 107 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 12996 | -0.183 | 0.0048 | No |
| 108 | SULT4A1 | SULT4A1 Entrez, Source | sulfotransferase family 4A, member 1 | 13166 | -0.233 | 0.0133 | No |