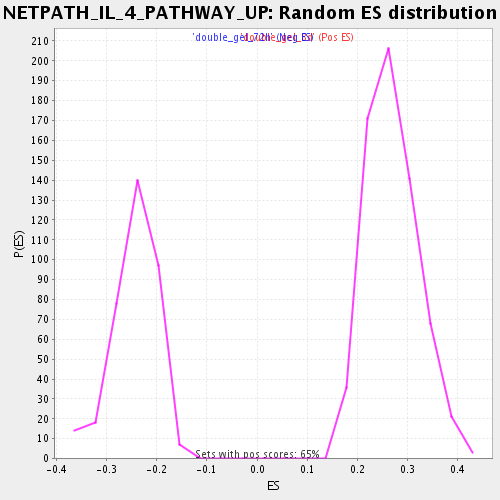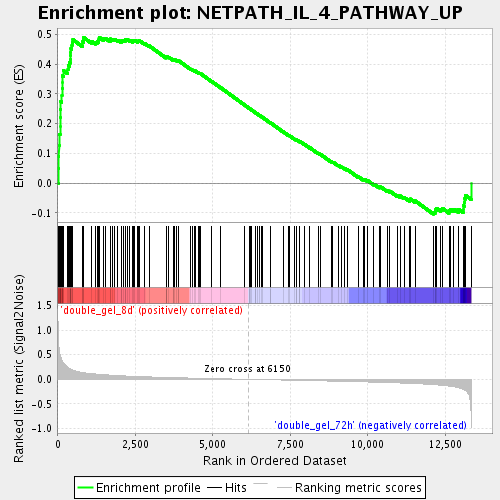
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | NETPATH_IL_4_PATHWAY_UP |
| Enrichment Score (ES) | 0.49005648 |
| Normalized Enrichment Score (NES) | 1.8252337 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.028663928 |
| FWER p-Value | 0.93 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 10 | 0.815 | 0.0483 | Yes |
| 2 | CYP1A1 | CYP1A1 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 1 | 18 | 0.708 | 0.0903 | Yes |
| 3 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 35 | 0.642 | 0.1277 | Yes |
| 4 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 42 | 0.616 | 0.1642 | Yes |
| 5 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 73 | 0.484 | 0.1911 | Yes |
| 6 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 76 | 0.477 | 0.2196 | Yes |
| 7 | LOX | LOX Entrez, Source | lysyl oxidase | 82 | 0.465 | 0.2472 | Yes |
| 8 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 91 | 0.451 | 0.2737 | Yes |
| 9 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 130 | 0.390 | 0.2943 | Yes |
| 10 | F13A1 | F13A1 Entrez, Source | coagulation factor XIII, A1 polypeptide | 135 | 0.379 | 0.3167 | Yes |
| 11 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 142 | 0.371 | 0.3385 | Yes |
| 12 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 145 | 0.367 | 0.3604 | Yes |
| 13 | SLC22A4 | SLC22A4 Entrez, Source | solute carrier family 22 (organic cation transporter), member 4 | 182 | 0.335 | 0.3778 | Yes |
| 14 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 323 | 0.239 | 0.3816 | Yes |
| 15 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 335 | 0.235 | 0.3949 | Yes |
| 16 | TMOD1 | TMOD1 Entrez, Source | tropomodulin 1 | 370 | 0.220 | 0.4055 | Yes |
| 17 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 399 | 0.209 | 0.4160 | Yes |
| 18 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 406 | 0.208 | 0.4280 | Yes |
| 19 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 412 | 0.206 | 0.4400 | Yes |
| 20 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 418 | 0.205 | 0.4520 | Yes |
| 21 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 451 | 0.195 | 0.4613 | Yes |
| 22 | JUN | JUN Entrez, Source | jun oncogene | 458 | 0.194 | 0.4725 | Yes |
| 23 | FBN1 | FBN1 Entrez, Source | fibrillin 1 | 483 | 0.185 | 0.4818 | Yes |
| 24 | GCNT1 | GCNT1 Entrez, Source | glucosaminyl (N-acetyl) transferase 1, core 2 (beta-1,6-N-acetylglucosaminyltransferase) | 800 | 0.140 | 0.4663 | Yes |
| 25 | GUCY1B3 | GUCY1B3 Entrez, Source | guanylate cyclase 1, soluble, beta 3 | 801 | 0.140 | 0.4746 | Yes |
| 26 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 825 | 0.137 | 0.4811 | Yes |
| 27 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 830 | 0.136 | 0.4890 | Yes |
| 28 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 1086 | 0.116 | 0.4766 | Yes |
| 29 | RASA2 | RASA2 Entrez, Source | RAS p21 protein activator 2 | 1216 | 0.109 | 0.4734 | Yes |
| 30 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 1268 | 0.106 | 0.4759 | Yes |
| 31 | ARG1 | ARG1 Entrez, Source | arginase, liver | 1316 | 0.103 | 0.4785 | Yes |
| 32 | IL13RA2 | IL13RA2 Entrez, Source | interleukin 13 receptor, alpha 2 | 1324 | 0.103 | 0.4842 | Yes |
| 33 | IL10 | IL10 Entrez, Source | interleukin 10 | 1329 | 0.102 | 0.4901 | Yes |
| 34 | SULF1 | SULF1 Entrez, Source | sulfatase 1 | 1470 | 0.095 | 0.4852 | No |
| 35 | PDZK1IP1 | PDZK1IP1 Entrez, Source | PDZK1 interacting protein 1 | 1523 | 0.093 | 0.4869 | No |
| 36 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1690 | 0.085 | 0.4794 | No |
| 37 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 1691 | 0.085 | 0.4845 | No |
| 38 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 1774 | 0.082 | 0.4832 | No |
| 39 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1840 | 0.078 | 0.4830 | No |
| 40 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 1934 | 0.075 | 0.4805 | No |
| 41 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 2045 | 0.071 | 0.4764 | No |
| 42 | SCGB3A1 | SCGB3A1 Entrez, Source | secretoglobin, family 3A, member 1 | 2064 | 0.071 | 0.4793 | No |
| 43 | SAMSN1 | SAMSN1 Entrez, Source | SAM domain, SH3 domain and nuclear localisation signals, 1 | 2103 | 0.070 | 0.4806 | No |
| 44 | ITGAM | ITGAM Entrez, Source | integrin, alpha M (complement component 3 receptor 3 subunit) | 2172 | 0.068 | 0.4796 | No |
| 45 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 2181 | 0.068 | 0.4831 | No |
| 46 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 2254 | 0.066 | 0.4815 | No |
| 47 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 2324 | 0.064 | 0.4802 | No |
| 48 | PAWR | PAWR Entrez, Source | PRKC, apoptosis, WT1, regulator | 2421 | 0.061 | 0.4766 | No |
| 49 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 2424 | 0.061 | 0.4801 | No |
| 50 | AGC1 | AGC1 Entrez, Source | aggrecan 1 (chondroitin sulfate proteoglycan 1, large aggregating proteoglycan, antigen identified by monoclonal antibody A0122) | 2474 | 0.060 | 0.4801 | No |
| 51 | PSCD1 | PSCD1 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 1(cytohesin 1) | 2579 | 0.058 | 0.4757 | No |
| 52 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 2597 | 0.058 | 0.4778 | No |
| 53 | FN1 | FN1 Entrez, Source | fibronectin 1 | 2637 | 0.057 | 0.4783 | No |
| 54 | TLR4 | TLR4 Entrez, Source | toll-like receptor 4 | 2800 | 0.053 | 0.4692 | No |
| 55 | IFNG | IFNG Entrez, Source | interferon, gamma | 2940 | 0.050 | 0.4617 | No |
| 56 | UPP1 | UPP1 Entrez, Source | uridine phosphorylase 1 | 3489 | 0.039 | 0.4225 | No |
| 57 | IL13 | IL13 Entrez, Source | interleukin 13 | 3501 | 0.039 | 0.4240 | No |
| 58 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 3509 | 0.038 | 0.4258 | No |
| 59 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 3568 | 0.037 | 0.4236 | No |
| 60 | SERPINB3 | SERPINB3 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 3 | 3724 | 0.035 | 0.4140 | No |
| 61 | SPINT2 | SPINT2 Entrez, Source | serine peptidase inhibitor, Kunitz type, 2 | 3725 | 0.035 | 0.4161 | No |
| 62 | MAL | MAL Entrez, Source | mal, T-cell differentiation protein | 3748 | 0.034 | 0.4165 | No |
| 63 | MAOA | MAOA Entrez, Source | monoamine oxidase A | 3833 | 0.033 | 0.4121 | No |
| 64 | CTSL | CTSL Entrez, Source | cathepsin L | 3892 | 0.032 | 0.4096 | No |
| 65 | ST8SIA1 | ST8SIA1 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 | 3901 | 0.032 | 0.4109 | No |
| 66 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 4288 | 0.026 | 0.3832 | No |
| 67 | HPCAL1 | HPCAL1 Entrez, Source | hippocalcin-like 1 | 4353 | 0.025 | 0.3799 | No |
| 68 | ALOX15 | ALOX15 Entrez, Source | arachidonate 15-lipoxygenase | 4419 | 0.023 | 0.3763 | No |
| 69 | IL2RB | IL2RB Entrez, Source | interleukin 2 receptor, beta | 4429 | 0.023 | 0.3771 | No |
| 70 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 4537 | 0.022 | 0.3703 | No |
| 71 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 4567 | 0.022 | 0.3694 | No |
| 72 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 4596 | 0.021 | 0.3685 | No |
| 73 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 4953 | 0.016 | 0.3425 | No |
| 74 | SOCS3 | SOCS3 Entrez, Source | suppressor of cytokine signaling 3 | 5260 | 0.012 | 0.3200 | No |
| 75 | IL10RA | IL10RA Entrez, Source | interleukin 10 receptor, alpha | 6007 | 0.002 | 0.2637 | No |
| 76 | CART1 | CART1 Entrez, Source | cartilage paired-class homeoprotein 1 | 6181 | -0.000 | 0.2506 | No |
| 77 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 6193 | -0.000 | 0.2498 | No |
| 78 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 6213 | -0.001 | 0.2484 | No |
| 79 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 6260 | -0.001 | 0.2450 | No |
| 80 | KCNK3 | KCNK3 Entrez, Source | potassium channel, subfamily K, member 3 | 6369 | -0.003 | 0.2370 | No |
| 81 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 6392 | -0.003 | 0.2355 | No |
| 82 | GAS6 | GAS6 Entrez, Source | growth arrest-specific 6 | 6432 | -0.004 | 0.2328 | No |
| 83 | EDNRA | EDNRA Entrez, Source | endothelin receptor type A | 6497 | -0.004 | 0.2282 | No |
| 84 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 6558 | -0.005 | 0.2240 | No |
| 85 | EEF1A2 | EEF1A2 Entrez, Source | eukaryotic translation elongation factor 1 alpha 2 | 6587 | -0.006 | 0.2222 | No |
| 86 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 6850 | -0.009 | 0.2029 | No |
| 87 | LRRN3 | LRRN3 Entrez, Source | leucine rich repeat neuronal 3 | 6873 | -0.009 | 0.2017 | No |
| 88 | TFEC | TFEC Entrez, Source | transcription factor EC | 7275 | -0.014 | 0.1722 | No |
| 89 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 7453 | -0.016 | 0.1598 | No |
| 90 | CHST10 | CHST10 Entrez, Source | carbohydrate sulfotransferase 10 | 7458 | -0.016 | 0.1605 | No |
| 91 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 7648 | -0.019 | 0.1473 | No |
| 92 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 7684 | -0.019 | 0.1458 | No |
| 93 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 7715 | -0.020 | 0.1447 | No |
| 94 | AHR | AHR Entrez, Source | aryl hydrocarbon receptor | 7804 | -0.021 | 0.1393 | No |
| 95 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 7809 | -0.021 | 0.1402 | No |
| 96 | CCL22 | CCL22 Entrez, Source | chemokine (C-C motif) ligand 22 | 7969 | -0.023 | 0.1296 | No |
| 97 | SLC39A8 | SLC39A8 Entrez, Source | solute carrier family 39 (zinc transporter), member 8 | 8133 | -0.026 | 0.1188 | No |
| 98 | BCL2L11 | BCL2L11 Entrez, Source | BCL2-like 11 (apoptosis facilitator) | 8410 | -0.030 | 0.0996 | No |
| 99 | VDR | VDR Entrez, Source | vitamin D (1,25- dihydroxyvitamin D3) receptor | 8460 | -0.030 | 0.0977 | No |
| 100 | XCL1 | XCL1 Entrez, Source | chemokine (C motif) ligand 1 | 8819 | -0.035 | 0.0727 | No |
| 101 | SOCS1 | SOCS1 Entrez, Source | suppressor of cytokine signaling 1 | 8877 | -0.036 | 0.0706 | No |
| 102 | HSPB8 | HSPB8 Entrez, Source | heat shock 22kDa protein 8 | 9051 | -0.038 | 0.0598 | No |
| 103 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9154 | -0.040 | 0.0544 | No |
| 104 | NFKB1 | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 9252 | -0.041 | 0.0496 | No |
| 105 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 9332 | -0.042 | 0.0461 | No |
| 106 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 9701 | -0.049 | 0.0212 | No |
| 107 | IGHE | IGHE Entrez, Source | immunoglobulin heavy constant epsilon | 9859 | -0.051 | 0.0124 | No |
| 108 | NKG7 | NKG7 Entrez, Source | natural killer cell group 7 sequence | 9908 | -0.052 | 0.0118 | No |
| 109 | GJA5 | GJA5 Entrez, Source | gap junction protein, alpha 5, 40kDa (connexin 40) | 9982 | -0.053 | 0.0095 | No |
| 110 | IL1RN | IL1RN Entrez, Source | interleukin 1 receptor antagonist | 10196 | -0.057 | -0.0032 | No |
| 111 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 10373 | -0.060 | -0.0130 | No |
| 112 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 10417 | -0.061 | -0.0126 | No |
| 113 | CD86 | CD86 Entrez, Source | CD86 molecule | 10637 | -0.065 | -0.0252 | No |
| 114 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 10706 | -0.066 | -0.0264 | No |
| 115 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 10964 | -0.073 | -0.0415 | No |
| 116 | RDX | RDX Entrez, Source | radixin | 11042 | -0.074 | -0.0429 | No |
| 117 | PMCH | PMCH Entrez, Source | pro-melanin-concentrating hormone | 11172 | -0.078 | -0.0480 | No |
| 118 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 11362 | -0.083 | -0.0573 | No |
| 119 | CTSC | CTSC Entrez, Source | cathepsin C | 11373 | -0.083 | -0.0531 | No |
| 120 | DNAJC3 | DNAJC3 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 3 | 11530 | -0.087 | -0.0596 | No |
| 121 | CH25H | CH25H Entrez, Source | cholesterol 25-hydroxylase | 12118 | -0.108 | -0.0976 | No |
| 122 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12174 | -0.110 | -0.0951 | No |
| 123 | IL13RA1 | IL13RA1 Entrez, Source | interleukin 13 receptor, alpha 1 | 12191 | -0.111 | -0.0896 | No |
| 124 | CASP2 | CASP2 Entrez, Source | caspase 2, apoptosis-related cysteine peptidase (neural precursor cell expressed, developmentally down-regulated 2) | 12210 | -0.112 | -0.0843 | No |
| 125 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 12352 | -0.119 | -0.0878 | No |
| 126 | MET | MET Entrez, Source | met proto-oncogene (hepatocyte growth factor receptor) | 12396 | -0.121 | -0.0838 | No |
| 127 | SELE | SELE Entrez, Source | selectin E (endothelial adhesion molecule 1) | 12624 | -0.136 | -0.0928 | No |
| 128 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 12659 | -0.138 | -0.0871 | No |
| 129 | RASGRP1 | RASGRP1 Entrez, Source | RAS guanyl releasing protein 1 (calcium and DAG-regulated) | 12782 | -0.150 | -0.0873 | No |
| 130 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12922 | -0.168 | -0.0878 | No |
| 131 | CA2 | CA2 Entrez, Source | carbonic anhydrase II | 13081 | -0.204 | -0.0875 | No |
| 132 | VCAM1 | VCAM1 Entrez, Source | vascular cell adhesion molecule 1 | 13085 | -0.205 | -0.0754 | No |
| 133 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13114 | -0.214 | -0.0647 | No |
| 134 | RIPK2 | RIPK2 Entrez, Source | receptor-interacting serine-threonine kinase 2 | 13131 | -0.220 | -0.0527 | No |
| 135 | LIFR | LIFR Entrez, Source | leukemia inhibitory factor receptor alpha | 13146 | -0.224 | -0.0402 | No |
| 136 | EDG1 | EDG1 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 1 | 13341 | -0.915 | 0.0001 | No |