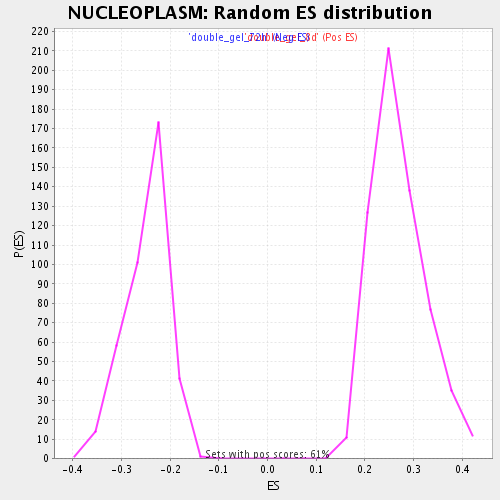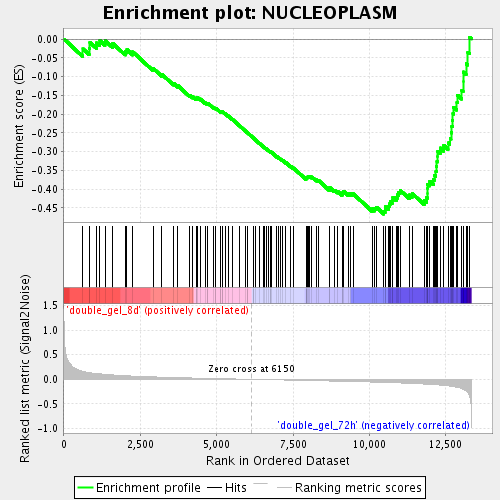
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | NUCLEOPLASM |
| Enrichment Score (ES) | -0.46644247 |
| Normalized Enrichment Score (NES) | -1.8723346 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.10313495 |
| FWER p-Value | 0.652 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GCM1 | GCM1 Entrez, Source | glial cells missing homolog 1 (Drosophila) | 623 | 0.162 | -0.0264 | No |
| 2 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 827 | 0.136 | -0.0243 | No |
| 3 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 850 | 0.134 | -0.0088 | No |
| 4 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1068 | 0.116 | -0.0104 | No |
| 5 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 1177 | 0.111 | -0.0044 | No |
| 6 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 1345 | 0.102 | -0.0040 | No |
| 7 | HDAC7A | HDAC7A Entrez, Source | histone deacetylase 7A | 1598 | 0.089 | -0.0117 | No |
| 8 | PTF1A | PTF1A Entrez, Source | pancreas specific transcription factor, 1a | 2008 | 0.073 | -0.0333 | No |
| 9 | MEN1 | MEN1 Entrez, Source | multiple endocrine neoplasia I | 2062 | 0.071 | -0.0282 | No |
| 10 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 2254 | 0.066 | -0.0343 | No |
| 11 | CDK9 | CDK9 Entrez, Source | cyclin-dependent kinase 9 (CDC2-related kinase) | 2915 | 0.051 | -0.0777 | No |
| 12 | CIB1 | CIB1 Entrez, Source | calcium and integrin binding 1 (calmyrin) | 3206 | 0.045 | -0.0940 | No |
| 13 | HCLS1 | HCLS1 Entrez, Source | hematopoietic cell-specific Lyn substrate 1 | 3587 | 0.037 | -0.1180 | No |
| 14 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 3726 | 0.035 | -0.1240 | No |
| 15 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 4109 | 0.029 | -0.1492 | No |
| 16 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 4217 | 0.027 | -0.1539 | No |
| 17 | TAF9 | TAF9 Entrez, Source | TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 32kDa | 4322 | 0.025 | -0.1585 | No |
| 18 | NAP1L2 | NAP1L2 Entrez, Source | nucleosome assembly protein 1-like 2 | 4354 | 0.025 | -0.1577 | No |
| 19 | PPARGC1B | PPARGC1B Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, beta | 4372 | 0.024 | -0.1559 | No |
| 20 | NKX2-5 | NKX2-5 Entrez, Source | NK2 transcription factor related, locus 5 (Drosophila) | 4463 | 0.023 | -0.1598 | No |
| 21 | HTATIP | HTATIP Entrez, Source | HIV-1 Tat interacting protein, 60kDa | 4633 | 0.021 | -0.1700 | No |
| 22 | TBP | TBP Entrez, Source | TATA box binding protein | 4690 | 0.020 | -0.1717 | No |
| 23 | ACTB | ACTB Entrez, Source | actin, beta | 4714 | 0.019 | -0.1709 | No |
| 24 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 4892 | 0.017 | -0.1822 | No |
| 25 | MCRS1 | MCRS1 Entrez, Source | microspherule protein 1 | 4951 | 0.016 | -0.1845 | No |
| 26 | SLU7 | SLU7 Entrez, Source | - | 4976 | 0.016 | -0.1844 | No |
| 27 | NCR1 | NCR1 Entrez, Source | natural cytotoxicity triggering receptor 1 | 5135 | 0.013 | -0.1946 | No |
| 28 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 5139 | 0.013 | -0.1931 | No |
| 29 | GTF3C2 | GTF3C2 Entrez, Source | general transcription factor IIIC, polypeptide 2, beta 110kDa | 5178 | 0.013 | -0.1943 | No |
| 30 | NCOA6 | NCOA6 Entrez, Source | nuclear receptor coactivator 6 | 5196 | 0.013 | -0.1940 | No |
| 31 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 5284 | 0.011 | -0.1991 | No |
| 32 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 5377 | 0.010 | -0.2048 | No |
| 33 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 5521 | 0.008 | -0.2146 | No |
| 34 | FOXE3 | FOXE3 Entrez, Source | forkhead box E3 | 5746 | 0.005 | -0.2309 | No |
| 35 | NAP1L4 | NAP1L4 Entrez, Source | nucleosome assembly protein 1-like 4 | 5935 | 0.003 | -0.2447 | No |
| 36 | RTCD1 | RTCD1 Entrez, Source | RNA terminal phosphate cyclase domain 1 | 5993 | 0.002 | -0.2487 | No |
| 37 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 6006 | 0.002 | -0.2494 | No |
| 38 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 6189 | -0.000 | -0.2631 | No |
| 39 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 6213 | -0.001 | -0.2647 | No |
| 40 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 6260 | -0.001 | -0.2680 | No |
| 41 | NFYC | NFYC Entrez, Source | nuclear transcription factor Y, gamma | 6397 | -0.003 | -0.2779 | No |
| 42 | NFYA | NFYA Entrez, Source | nuclear transcription factor Y, alpha | 6517 | -0.005 | -0.2863 | No |
| 43 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 6574 | -0.005 | -0.2899 | No |
| 44 | TAF11 | TAF11 Entrez, Source | TAF11 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 28kDa | 6619 | -0.006 | -0.2924 | No |
| 45 | TAF6 | TAF6 Entrez, Source | TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa | 6684 | -0.007 | -0.2964 | No |
| 46 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 6756 | -0.008 | -0.3008 | No |
| 47 | THRAP3 | THRAP3 Entrez, Source | thyroid hormone receptor associated protein 3 | 6772 | -0.008 | -0.3009 | No |
| 48 | SMAD4 | SMAD4 Entrez, Source | SMAD, mothers against DPP homolog 4 (Drosophila) | 6788 | -0.008 | -0.3010 | No |
| 49 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 6971 | -0.010 | -0.3135 | No |
| 50 | OGG1 | OGG1 Entrez, Source | 8-oxoguanine DNA glycosylase | 7030 | -0.011 | -0.3164 | No |
| 51 | CLOCK | CLOCK Entrez, Source | clock homolog (mouse) | 7102 | -0.012 | -0.3203 | No |
| 52 | HDAC3 | HDAC3 Entrez, Source | histone deacetylase 3 | 7165 | -0.013 | -0.3234 | No |
| 53 | SYMPK | SYMPK Entrez, Source | symplekin | 7266 | -0.014 | -0.3292 | No |
| 54 | INTS6 | INTS6 Entrez, Source | integrator complex subunit 6 | 7424 | -0.016 | -0.3390 | No |
| 55 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 7512 | -0.017 | -0.3435 | No |
| 56 | SUB1 | SUB1 Entrez, Source | SUB1 homolog (S. cerevisiae) | 7922 | -0.022 | -0.3716 | No |
| 57 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 7943 | -0.023 | -0.3702 | No |
| 58 | SAP18 | SAP18 Entrez, Source | Sin3A-associated protein, 18kDa | 7954 | -0.023 | -0.3680 | No |
| 59 | SMARCB1 | SMARCB1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 | 7987 | -0.023 | -0.3674 | No |
| 60 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 8006 | -0.024 | -0.3658 | No |
| 61 | PRM2 | PRM2 Entrez, Source | protamine 2 | 8050 | -0.024 | -0.3659 | No |
| 62 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 8111 | -0.025 | -0.3673 | No |
| 63 | HNRPK | HNRPK Entrez, Source | heterogeneous nuclear ribonucleoprotein K | 8260 | -0.027 | -0.3750 | No |
| 64 | PHB | PHB Entrez, Source | prohibitin | 8324 | -0.028 | -0.3761 | No |
| 65 | NUDT21 | NUDT21 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 21 | 8696 | -0.034 | -0.3999 | No |
| 66 | SMARCD2 | SMARCD2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | 8700 | -0.034 | -0.3958 | No |
| 67 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 8847 | -0.035 | -0.4023 | No |
| 68 | CPSF3 | CPSF3 Entrez, Source | cleavage and polyadenylation specific factor 3, 73kDa | 8954 | -0.037 | -0.4056 | No |
| 69 | IKBKAP | IKBKAP Entrez, Source | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase complex-associated protein | 9104 | -0.039 | -0.4119 | No |
| 70 | HDAC10 | HDAC10 Entrez, Source | histone deacetylase 10 | 9127 | -0.039 | -0.4086 | No |
| 71 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9154 | -0.040 | -0.4055 | No |
| 72 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 9321 | -0.042 | -0.4127 | No |
| 73 | DKC1 | DKC1 Entrez, Source | dyskeratosis congenita 1, dyskerin | 9386 | -0.043 | -0.4120 | No |
| 74 | PAX8 | PAX8 Entrez, Source | paired box gene 8 | 9468 | -0.044 | -0.4124 | No |
| 75 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 10088 | -0.055 | -0.4522 | No |
| 76 | GEMIN6 | GEMIN6 Entrez, Source | gem (nuclear organelle) associated protein 6 | 10170 | -0.056 | -0.4511 | No |
| 77 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 10227 | -0.057 | -0.4481 | No |
| 78 | COIL | COIL Entrez, Source | coilin | 10471 | -0.062 | -0.4586 | Yes |
| 79 | MED4 | MED4 Entrez, Source | mediator of RNA polymerase II transcription, subunit 4 homolog (S. cerevisiae) | 10512 | -0.062 | -0.4536 | Yes |
| 80 | GTF2F1 | GTF2F1 Entrez, Source | general transcription factor IIF, polypeptide 1, 74kDa | 10529 | -0.063 | -0.4468 | Yes |
| 81 | ATXN3 | ATXN3 Entrez, Source | ataxin 3 | 10628 | -0.065 | -0.4459 | Yes |
| 82 | MPG | MPG Entrez, Source | N-methylpurine-DNA glycosylase | 10650 | -0.065 | -0.4391 | Yes |
| 83 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 10677 | -0.066 | -0.4327 | Yes |
| 84 | PPIG | PPIG Entrez, Source | peptidylprolyl isomerase G (cyclophilin G) | 10743 | -0.067 | -0.4290 | Yes |
| 85 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 10765 | -0.068 | -0.4220 | Yes |
| 86 | GATAD2A | GATAD2A Entrez, Source | GATA zinc finger domain containing 2A | 10877 | -0.070 | -0.4214 | Yes |
| 87 | HDAC6 | HDAC6 Entrez, Source | histone deacetylase 6 | 10914 | -0.071 | -0.4150 | Yes |
| 88 | MKI67IP | MKI67IP Entrez, Source | MKI67 (FHA domain) interacting nucleolar phosphoprotein | 10951 | -0.072 | -0.4085 | Yes |
| 89 | NUFIP1 | NUFIP1 Entrez, Source | nuclear fragile X mental retardation protein interacting protein 1 | 11003 | -0.073 | -0.4030 | Yes |
| 90 | RBM39 | RBM39 Entrez, Source | RNA binding motif protein 39 | 11305 | -0.082 | -0.4153 | Yes |
| 91 | TFIP11 | TFIP11 Entrez, Source | tuftelin interacting protein 11 | 11393 | -0.084 | -0.4112 | Yes |
| 92 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 11792 | -0.095 | -0.4291 | Yes |
| 93 | ACTL6A | ACTL6A Entrez, Source | actin-like 6A | 11873 | -0.098 | -0.4226 | Yes |
| 94 | TAF2 | TAF2 Entrez, Source | TAF2 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 150kDa | 11894 | -0.099 | -0.4114 | Yes |
| 95 | SP140 | SP140 Entrez, Source | SP140 nuclear body protein | 11896 | -0.099 | -0.3988 | Yes |
| 96 | FUSIP1 | FUSIP1 Entrez, Source | FUS interacting protein (serine/arginine-rich) 1 | 11902 | -0.099 | -0.3865 | Yes |
| 97 | GTF3C1 | GTF3C1 Entrez, Source | general transcription factor IIIC, polypeptide 1, alpha 220kDa | 11971 | -0.102 | -0.3786 | Yes |
| 98 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 12100 | -0.107 | -0.3746 | Yes |
| 99 | NCBP1 | NCBP1 Entrez, Source | nuclear cap binding protein subunit 1, 80kDa | 12122 | -0.108 | -0.3624 | Yes |
| 100 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12174 | -0.110 | -0.3521 | Yes |
| 101 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 12189 | -0.111 | -0.3390 | Yes |
| 102 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 12208 | -0.112 | -0.3260 | Yes |
| 103 | SPTBN4 | SPTBN4 Entrez, Source | spectrin, beta, non-erythrocytic 4 | 12231 | -0.113 | -0.3133 | Yes |
| 104 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 12235 | -0.113 | -0.2991 | Yes |
| 105 | GTF2H4 | GTF2H4 Entrez, Source | general transcription factor IIH, polypeptide 4, 52kDa | 12316 | -0.117 | -0.2902 | Yes |
| 106 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 12430 | -0.123 | -0.2831 | Yes |
| 107 | ARID1B | ARID1B Entrez, Source | AT rich interactive domain 1B (SWI1-like) | 12572 | -0.133 | -0.2767 | Yes |
| 108 | NUP54 | NUP54 Entrez, Source | nucleoporin 54kDa | 12635 | -0.137 | -0.2639 | Yes |
| 109 | POLR3C | POLR3C Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide C (62kD) | 12669 | -0.139 | -0.2487 | Yes |
| 110 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 12694 | -0.141 | -0.2324 | Yes |
| 111 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 12718 | -0.143 | -0.2159 | Yes |
| 112 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12731 | -0.144 | -0.1984 | Yes |
| 113 | HES6 | HES6 Entrez, Source | hairy and enhancer of split 6 (Drosophila) | 12756 | -0.147 | -0.1814 | Yes |
| 114 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 12862 | -0.160 | -0.1689 | Yes |
| 115 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 12879 | -0.162 | -0.1494 | Yes |
| 116 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 13022 | -0.190 | -0.1358 | Yes |
| 117 | NUAK2 | NUAK2 Entrez, Source | NUAK family, SNF1-like kinase, 2 | 13065 | -0.201 | -0.1134 | Yes |
| 118 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 13067 | -0.201 | -0.0878 | Yes |
| 119 | NAP1L3 | NAP1L3 Entrez, Source | nucleosome assembly protein 1-like 3 | 13164 | -0.231 | -0.0654 | Yes |
| 120 | APTX | APTX Entrez, Source | aprataxin | 13219 | -0.270 | -0.0349 | Yes |
| 121 | ARNTL | ARNTL Entrez, Source | aryl hydrocarbon receptor nuclear translocator-like | 13280 | -0.345 | 0.0047 | Yes |