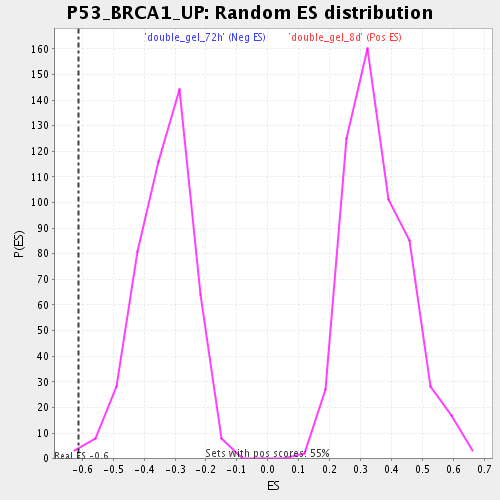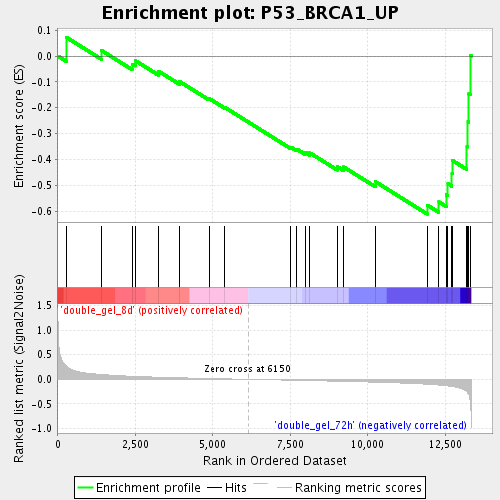
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | P53_BRCA1_UP |
| Enrichment Score (ES) | -0.6115242 |
| Normalized Enrichment Score (NES) | -1.8233182 |
| Nominal p-value | 0.0022123894 |
| FDR q-value | 0.112121865 |
| FWER p-Value | 0.841 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 269 | 0.266 | 0.0728 | No |
| 2 | CTSF | CTSF Entrez, Source | cathepsin F | 1408 | 0.098 | 0.0215 | No |
| 3 | COX6B2 | COX6B2 Entrez, Source | cytochrome c oxidase subunit VIb polypeptide 2 (testis) | 2398 | 0.062 | -0.0311 | No |
| 4 | PQLC3 | PQLC3 Entrez, Source | PQ loop repeat containing 3 | 2498 | 0.060 | -0.0177 | No |
| 5 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 3249 | 0.044 | -0.0587 | No |
| 6 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 3925 | 0.031 | -0.0985 | No |
| 7 | AK1 | AK1 Entrez, Source | adenylate kinase 1 | 4876 | 0.017 | -0.1638 | No |
| 8 | CTSH | CTSH Entrez, Source | cathepsin H | 5387 | 0.010 | -0.1986 | No |
| 9 | ZMAT3 | ZMAT3 Entrez, Source | zinc finger, matrin type 3 | 7516 | -0.017 | -0.3525 | No |
| 10 | PTPRV | PTPRV Entrez, Source | protein tyrosine phosphatase, receptor type, V (pseudogene) | 7703 | -0.019 | -0.3597 | No |
| 11 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 7976 | -0.023 | -0.3721 | No |
| 12 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 8130 | -0.026 | -0.3747 | No |
| 13 | ROBO1 | ROBO1 Entrez, Source | roundabout, axon guidance receptor, homolog 1 (Drosophila) | 9015 | -0.038 | -0.4279 | No |
| 14 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 9220 | -0.040 | -0.4291 | No |
| 15 | FXYD3 | FXYD3 Entrez, Source | FXYD domain containing ion transport regulator 3 | 10251 | -0.058 | -0.4863 | No |
| 16 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 11920 | -0.100 | -0.5767 | Yes |
| 17 | SRXN1 | SRXN1 Entrez, Source | sulfiredoxin 1 homolog (S. cerevisiae) | 12296 | -0.116 | -0.5644 | Yes |
| 18 | PHLDA3 | PHLDA3 Entrez, Source | pleckstrin homology-like domain, family A, member 3 | 12536 | -0.130 | -0.5369 | Yes |
| 19 | EPHX1 | EPHX1 Entrez, Source | epoxide hydrolase 1, microsomal (xenobiotic) | 12587 | -0.134 | -0.4939 | Yes |
| 20 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 12718 | -0.143 | -0.4537 | Yes |
| 21 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 12743 | -0.146 | -0.4048 | Yes |
| 22 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 13202 | -0.257 | -0.3496 | Yes |
| 23 | ENPP2 | ENPP2 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 2 (autotaxin) | 13234 | -0.281 | -0.2539 | Yes |
| 24 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 13262 | -0.313 | -0.1466 | Yes |
| 25 | APOBEC1 | APOBEC1 Entrez, Source | apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1 | 13309 | -0.437 | 0.0025 | Yes |