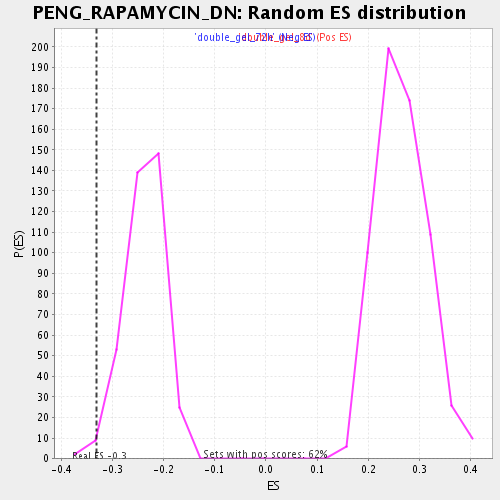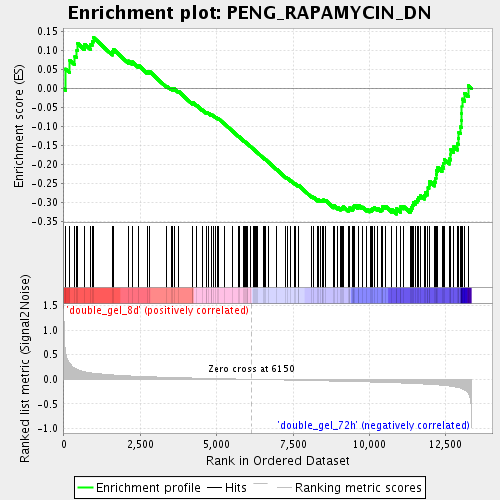
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | PENG_RAPAMYCIN_DN |
| Enrichment Score (ES) | -0.33097607 |
| Normalized Enrichment Score (NES) | -1.3883328 |
| Nominal p-value | 0.010638298 |
| FDR q-value | 0.4984215 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 61 | 0.555 | 0.0517 | No |
| 2 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 192 | 0.328 | 0.0751 | No |
| 3 | EMP3 | EMP3 Entrez, Source | epithelial membrane protein 3 | 358 | 0.227 | 0.0857 | No |
| 4 | P2RX5 | P2RX5 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 5 | 423 | 0.203 | 0.1014 | No |
| 5 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 450 | 0.195 | 0.1193 | No |
| 6 | ACVRL1 | ACVRL1 Entrez, Source | activin A receptor type II-like 1 | 678 | 0.153 | 0.1176 | No |
| 7 | HADH2 | HADH2 Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase, type II | 876 | 0.131 | 0.1160 | No |
| 8 | EIF2B2 | EIF2B2 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 2 beta, 39kDa | 924 | 0.127 | 0.1254 | No |
| 9 | PTPN1 | PTPN1 Entrez, Source | protein tyrosine phosphatase, non-receptor type 1 | 959 | 0.124 | 0.1354 | No |
| 10 | EXTL3 | EXTL3 Entrez, Source | exostoses (multiple)-like 3 | 1580 | 0.090 | 0.0976 | No |
| 11 | RTN4 | RTN4 Entrez, Source | reticulon 4 | 1617 | 0.089 | 0.1039 | No |
| 12 | PRDX3 | PRDX3 Entrez, Source | peroxiredoxin 3 | 2119 | 0.069 | 0.0730 | No |
| 13 | INPP5E | INPP5E Entrez, Source | inositol polyphosphate-5-phosphatase, 72 kDa | 2239 | 0.066 | 0.0707 | No |
| 14 | PSMD8 | PSMD8 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 8 | 2441 | 0.061 | 0.0616 | No |
| 15 | HBB | HBB Entrez, Source | hemoglobin, beta | 2720 | 0.055 | 0.0461 | No |
| 16 | RAB14 | RAB14 Entrez, Source | RAB14, member RAS oncogene family | 2806 | 0.053 | 0.0450 | No |
| 17 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 3372 | 0.041 | 0.0064 | No |
| 18 | IDH3A | IDH3A Entrez, Source | isocitrate dehydrogenase 3 (NAD+) alpha | 3505 | 0.039 | 0.0003 | No |
| 19 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 3568 | 0.037 | -0.0006 | No |
| 20 | GTF2A2 | GTF2A2 Entrez, Source | general transcription factor IIA, 2, 12kDa | 3615 | 0.037 | -0.0003 | No |
| 21 | CAPZA2 | CAPZA2 Entrez, Source | capping protein (actin filament) muscle Z-line, alpha 2 | 3750 | 0.034 | -0.0070 | No |
| 22 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 4212 | 0.027 | -0.0392 | No |
| 23 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 4217 | 0.027 | -0.0368 | No |
| 24 | BCAP31 | BCAP31 Entrez, Source | B-cell receptor-associated protein 31 | 4339 | 0.025 | -0.0434 | No |
| 25 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 4537 | 0.022 | -0.0561 | No |
| 26 | COX7B | COX7B Entrez, Source | cytochrome c oxidase subunit VIIb | 4657 | 0.020 | -0.0631 | No |
| 27 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 4681 | 0.020 | -0.0628 | No |
| 28 | PFKM | PFKM Entrez, Source | phosphofructokinase, muscle | 4734 | 0.019 | -0.0648 | No |
| 29 | ETFA | ETFA Entrez, Source | electron-transfer-flavoprotein, alpha polypeptide (glutaric aciduria II) | 4825 | 0.018 | -0.0698 | No |
| 30 | ARF1 | ARF1 Entrez, Source | ADP-ribosylation factor 1 | 4845 | 0.018 | -0.0694 | No |
| 31 | NMT1 | NMT1 Entrez, Source | N-myristoyltransferase 1 | 4905 | 0.017 | -0.0722 | No |
| 32 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 4968 | 0.016 | -0.0753 | No |
| 33 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 5035 | 0.015 | -0.0788 | No |
| 34 | CD47 | CD47 Entrez, Source | CD47 molecule | 5066 | 0.014 | -0.0796 | No |
| 35 | PGAM1 | PGAM1 Entrez, Source | phosphoglycerate mutase 1 (brain) | 5246 | 0.012 | -0.0920 | No |
| 36 | PIP5K1A | PIP5K1A Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, alpha | 5528 | 0.008 | -0.1125 | No |
| 37 | PPIB | PPIB Entrez, Source | peptidylprolyl isomerase B (cyclophilin B) | 5713 | 0.006 | -0.1258 | No |
| 38 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 5742 | 0.005 | -0.1274 | No |
| 39 | BMP1 | BMP1 Entrez, Source | bone morphogenetic protein 1 | 5892 | 0.004 | -0.1384 | No |
| 40 | PPIF | PPIF Entrez, Source | peptidylprolyl isomerase F (cyclophilin F) | 5913 | 0.003 | -0.1396 | No |
| 41 | POLR2F | POLR2F Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide F | 5958 | 0.003 | -0.1426 | No |
| 42 | JTB | JTB Entrez, Source | jumping translocation breakpoint | 5970 | 0.002 | -0.1432 | No |
| 43 | RTCD1 | RTCD1 Entrez, Source | RNA terminal phosphate cyclase domain 1 | 5993 | 0.002 | -0.1446 | No |
| 44 | CCNC | CCNC Entrez, Source | cyclin C | 5995 | 0.002 | -0.1445 | No |
| 45 | SLC1A5 | SLC1A5 Entrez, Source | solute carrier family 1 (neutral amino acid transporter), member 5 | 6099 | 0.001 | -0.1522 | No |
| 46 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 6191 | -0.000 | -0.1591 | No |
| 47 | TIMM44 | TIMM44 Entrez, Source | translocase of inner mitochondrial membrane 44 homolog (yeast) | 6231 | -0.001 | -0.1619 | No |
| 48 | CCT6A | CCT6A Entrez, Source | chaperonin containing TCP1, subunit 6A (zeta 1) | 6256 | -0.001 | -0.1636 | No |
| 49 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 6301 | -0.002 | -0.1667 | No |
| 50 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 6333 | -0.002 | -0.1688 | No |
| 51 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 6529 | -0.005 | -0.1831 | No |
| 52 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 6557 | -0.005 | -0.1846 | No |
| 53 | PSMC2 | PSMC2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 2 | 6594 | -0.006 | -0.1868 | No |
| 54 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 6700 | -0.007 | -0.1940 | No |
| 55 | ADRM1 | ADRM1 Entrez, Source | adhesion regulating molecule 1 | 6953 | -0.010 | -0.2121 | No |
| 56 | MAPKAPK3 | MAPKAPK3 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 3 | 7267 | -0.014 | -0.2344 | No |
| 57 | PTMS | PTMS Entrez, Source | parathymosin | 7316 | -0.014 | -0.2366 | No |
| 58 | VCP | VCP Entrez, Source | valosin-containing protein | 7423 | -0.016 | -0.2430 | No |
| 59 | SAFB | SAFB Entrez, Source | scaffold attachment factor B | 7539 | -0.017 | -0.2500 | No |
| 60 | MORF4L2 | MORF4L2 Entrez, Source | mortality factor 4 like 2 | 7580 | -0.018 | -0.2512 | No |
| 61 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 7672 | -0.019 | -0.2561 | No |
| 62 | DNAJA1 | DNAJA1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 1 | 7683 | -0.019 | -0.2549 | No |
| 63 | USP10 | USP10 Entrez, Source | ubiquitin specific peptidase 10 | 8103 | -0.025 | -0.2841 | No |
| 64 | CCT2 | CCT2 Entrez, Source | chaperonin containing TCP1, subunit 2 (beta) | 8167 | -0.026 | -0.2862 | No |
| 65 | GORASP2 | GORASP2 Entrez, Source | golgi reassembly stacking protein 2, 55kDa | 8308 | -0.028 | -0.2940 | No |
| 66 | PHB | PHB Entrez, Source | prohibitin | 8324 | -0.028 | -0.2923 | No |
| 67 | WDR18 | WDR18 Entrez, Source | WD repeat domain 18 | 8388 | -0.029 | -0.2941 | No |
| 68 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 8447 | -0.030 | -0.2955 | No |
| 69 | RNASEH1 | RNASEH1 Entrez, Source | ribonuclease H1 | 8474 | -0.031 | -0.2943 | No |
| 70 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 8504 | -0.031 | -0.2934 | No |
| 71 | EIF2B4 | EIF2B4 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 4 delta, 67kDa | 8566 | -0.032 | -0.2948 | No |
| 72 | PSMB5 | PSMB5 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 5 | 8833 | -0.035 | -0.3114 | No |
| 73 | RBBP6 | RBBP6 Entrez, Source | retinoblastoma binding protein 6 | 8839 | -0.035 | -0.3081 | No |
| 74 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 8969 | -0.037 | -0.3141 | No |
| 75 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 9054 | -0.038 | -0.3166 | No |
| 76 | CCT4 | CCT4 Entrez, Source | chaperonin containing TCP1, subunit 4 (delta) | 9088 | -0.039 | -0.3152 | No |
| 77 | COPB2 | COPB2 Entrez, Source | coatomer protein complex, subunit beta 2 (beta prime) | 9125 | -0.039 | -0.3139 | No |
| 78 | PGK1 | PGK1 Entrez, Source | phosphoglycerate kinase 1 | 9144 | -0.039 | -0.3113 | No |
| 79 | RAE1 | RAE1 Entrez, Source | RAE1 RNA export 1 homolog (S. pombe) | 9329 | -0.042 | -0.3210 | No |
| 80 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 9335 | -0.042 | -0.3171 | No |
| 81 | SSBP1 | SSBP1 Entrez, Source | single-stranded DNA binding protein 1 | 9358 | -0.043 | -0.3144 | No |
| 82 | TOMM40 | TOMM40 Entrez, Source | translocase of outer mitochondrial membrane 40 homolog (yeast) | 9457 | -0.044 | -0.3173 | No |
| 83 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 9466 | -0.044 | -0.3134 | No |
| 84 | CCT5 | CCT5 Entrez, Source | chaperonin containing TCP1, subunit 5 (epsilon) | 9486 | -0.045 | -0.3103 | No |
| 85 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 9509 | -0.045 | -0.3074 | No |
| 86 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 9628 | -0.047 | -0.3115 | No |
| 87 | SQLE | SQLE Entrez, Source | squalene epoxidase | 9644 | -0.048 | -0.3078 | No |
| 88 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 9770 | -0.050 | -0.3122 | No |
| 89 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 9915 | -0.052 | -0.3178 | No |
| 90 | NARS | NARS Entrez, Source | asparaginyl-tRNA synthetase | 10017 | -0.054 | -0.3200 | No |
| 91 | PITPNB | PITPNB Entrez, Source | phosphatidylinositol transfer protein, beta | 10077 | -0.055 | -0.3189 | No |
| 92 | BYSL | BYSL Entrez, Source | bystin-like | 10106 | -0.055 | -0.3154 | No |
| 93 | PTDSS1 | PTDSS1 Entrez, Source | phosphatidylserine synthase 1 | 10150 | -0.056 | -0.3130 | No |
| 94 | KPNA3 | KPNA3 Entrez, Source | karyopherin alpha 3 (importin alpha 4) | 10274 | -0.058 | -0.3164 | No |
| 95 | RABGGTB | RABGGTB Entrez, Source | Rab geranylgeranyltransferase, beta subunit | 10378 | -0.060 | -0.3182 | No |
| 96 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 10415 | -0.061 | -0.3148 | No |
| 97 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 10434 | -0.061 | -0.3099 | No |
| 98 | NXF1 | NXF1 Entrez, Source | nuclear RNA export factor 1 | 10534 | -0.063 | -0.3110 | No |
| 99 | RPS26 | RPS26 Entrez, Source | ribosomal protein S26 | 10735 | -0.067 | -0.3194 | No |
| 100 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 10889 | -0.071 | -0.3238 | Yes |
| 101 | EIF3S9 | EIF3S9 Entrez, Source | eukaryotic translation initiation factor 3, subunit 9 eta, 116kDa | 10891 | -0.071 | -0.3167 | Yes |
| 102 | ASNS | ASNS Entrez, Source | asparagine synthetase | 11024 | -0.074 | -0.3192 | Yes |
| 103 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 11025 | -0.074 | -0.3117 | Yes |
| 104 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 11103 | -0.076 | -0.3099 | Yes |
| 105 | GLO1 | GLO1 Entrez, Source | glyoxalase I | 11333 | -0.082 | -0.3189 | Yes |
| 106 | CTSC | CTSC Entrez, Source | cathepsin C | 11373 | -0.083 | -0.3134 | Yes |
| 107 | TARS | TARS Entrez, Source | threonyl-tRNA synthetase | 11414 | -0.084 | -0.3078 | Yes |
| 108 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 11432 | -0.085 | -0.3005 | Yes |
| 109 | C1QBP | C1QBP Entrez, Source | complement component 1, q subcomponent binding protein | 11506 | -0.086 | -0.2973 | Yes |
| 110 | DAP3 | DAP3 Entrez, Source | death associated protein 3 | 11558 | -0.088 | -0.2922 | Yes |
| 111 | EBNA1BP2 | EBNA1BP2 Entrez, Source | EBNA1 binding protein 2 | 11605 | -0.089 | -0.2866 | Yes |
| 112 | SF3A3 | SF3A3 Entrez, Source | splicing factor 3a, subunit 3, 60kDa | 11660 | -0.090 | -0.2815 | Yes |
| 113 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 11792 | -0.095 | -0.2818 | Yes |
| 114 | NOL5A | NOL5A Entrez, Source | nucleolar protein 5A (56kDa with KKE/D repeat) | 11821 | -0.096 | -0.2741 | Yes |
| 115 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 11909 | -0.100 | -0.2706 | Yes |
| 116 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 11914 | -0.100 | -0.2608 | Yes |
| 117 | EBP | EBP Entrez, Source | emopamil binding protein (sterol isomerase) | 11949 | -0.101 | -0.2532 | Yes |
| 118 | GTF3C1 | GTF3C1 Entrez, Source | general transcription factor IIIC, polypeptide 1, alpha 220kDa | 11971 | -0.102 | -0.2444 | Yes |
| 119 | BCS1L | BCS1L Entrez, Source | BCS1-like (yeast) | 12138 | -0.109 | -0.2460 | Yes |
| 120 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 12159 | -0.110 | -0.2363 | Yes |
| 121 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 12179 | -0.110 | -0.2265 | Yes |
| 122 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 12195 | -0.111 | -0.2164 | Yes |
| 123 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 12233 | -0.113 | -0.2077 | Yes |
| 124 | FKBP4 | FKBP4 Entrez, Source | FK506 binding protein 4, 59kDa | 12376 | -0.120 | -0.2063 | Yes |
| 125 | PA2G4 | PA2G4 Entrez, Source | proliferation-associated 2G4, 38kDa | 12425 | -0.123 | -0.1975 | Yes |
| 126 | AATF | AATF Entrez, Source | apoptosis antagonizing transcription factor | 12443 | -0.123 | -0.1863 | Yes |
| 127 | XPO6 | XPO6 Entrez, Source | exportin 6 | 12613 | -0.135 | -0.1853 | Yes |
| 128 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 12656 | -0.138 | -0.1745 | Yes |
| 129 | XBP1 | XBP1 Entrez, Source | X-box binding protein 1 | 12661 | -0.138 | -0.1608 | Yes |
| 130 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 12760 | -0.148 | -0.1532 | Yes |
| 131 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 12879 | -0.162 | -0.1457 | Yes |
| 132 | HYOU1 | HYOU1 Entrez, Source | hypoxia up-regulated 1 | 12907 | -0.166 | -0.1309 | Yes |
| 133 | SRM | SRM Entrez, Source | spermidine synthase | 12923 | -0.168 | -0.1150 | Yes |
| 134 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 12976 | -0.177 | -0.1009 | Yes |
| 135 | LIMK2 | LIMK2 Entrez, Source | LIM domain kinase 2 | 13008 | -0.187 | -0.0843 | Yes |
| 136 | SLC3A2 | SLC3A2 Entrez, Source | solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2 | 13019 | -0.189 | -0.0659 | Yes |
| 137 | SLC29A1 | SLC29A1 Entrez, Source | solute carrier family 29 (nucleoside transporters), member 1 | 13027 | -0.192 | -0.0469 | Yes |
| 138 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 13029 | -0.193 | -0.0274 | Yes |
| 139 | SREBF1 | SREBF1 Entrez, Source | sterol regulatory element binding transcription factor 1 | 13110 | -0.214 | -0.0117 | Yes |
| 140 | FASN | FASN Entrez, Source | fatty acid synthase | 13241 | -0.288 | 0.0076 | Yes |