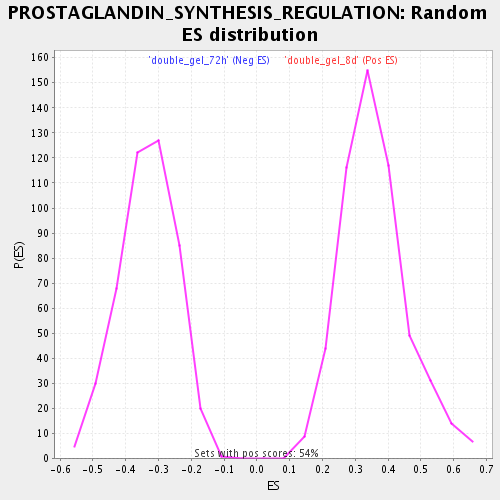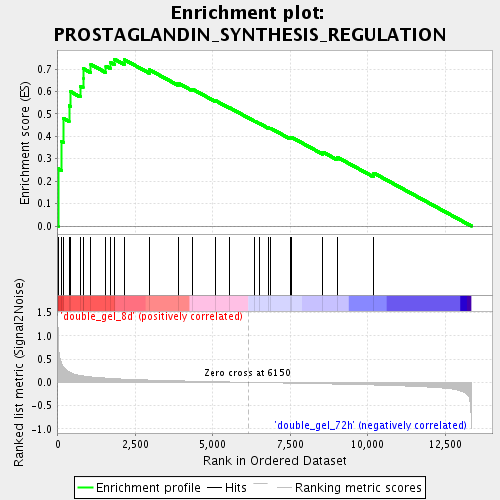
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | PROSTAGLANDIN_SYNTHESIS_REGULATION |
| Enrichment Score (ES) | 0.74401015 |
| Normalized Enrichment Score (NES) | 2.0853746 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0027639663 |
| FWER p-Value | 0.049 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 10 | 0.815 | 0.2559 | Yes |
| 2 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 113 | 0.410 | 0.3774 | Yes |
| 3 | ANXA8 | ANXA8 Entrez, Source | annexin A8 | 177 | 0.340 | 0.4796 | Yes |
| 4 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 361 | 0.225 | 0.5368 | Yes |
| 5 | S100A6 | S100A6 Entrez, Source | S100 calcium binding protein A6 | 402 | 0.209 | 0.5996 | Yes |
| 6 | ANXA4 | ANXA4 Entrez, Source | annexin A4 | 722 | 0.148 | 0.6223 | Yes |
| 7 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 812 | 0.139 | 0.6592 | Yes |
| 8 | PTGER1 | PTGER1 Entrez, Source | prostaglandin E receptor 1 (subtype EP1), 42kDa | 818 | 0.137 | 0.7021 | Yes |
| 9 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 1062 | 0.117 | 0.7205 | Yes |
| 10 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 1550 | 0.092 | 0.7128 | Yes |
| 11 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 1691 | 0.085 | 0.7290 | Yes |
| 12 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 1825 | 0.079 | 0.7440 | Yes |
| 13 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 2143 | 0.069 | 0.7418 | No |
| 14 | ANXA6 | ANXA6 Entrez, Source | annexin A6 | 2949 | 0.050 | 0.6971 | No |
| 15 | PTGFR | PTGFR Entrez, Source | prostaglandin F receptor (FP) | 3885 | 0.032 | 0.6369 | No |
| 16 | HSD11B2 | HSD11B2 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 2 | 4347 | 0.025 | 0.6101 | No |
| 17 | SCGB1A1 | SCGB1A1 Entrez, Source | secretoglobin, family 1A, member 1 (uteroglobin) | 5076 | 0.014 | 0.5599 | No |
| 18 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 5536 | 0.008 | 0.5279 | No |
| 19 | PRL | PRL Entrez, Source | prolactin | 6356 | -0.002 | 0.4671 | No |
| 20 | EDNRA | EDNRA Entrez, Source | endothelin receptor type A | 6497 | -0.004 | 0.4580 | No |
| 21 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6790 | -0.008 | 0.4386 | No |
| 22 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 6866 | -0.009 | 0.4358 | No |
| 23 | CYP11A1 | CYP11A1 Entrez, Source | cytochrome P450, family 11, subfamily A, polypeptide 1 | 7492 | -0.017 | 0.3941 | No |
| 24 | EDNRB | EDNRB Entrez, Source | endothelin receptor type B | 7528 | -0.017 | 0.3968 | No |
| 25 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 8552 | -0.031 | 0.3299 | No |
| 26 | PTGIS | PTGIS Entrez, Source | prostaglandin I2 (prostacyclin) synthase | 9012 | -0.038 | 0.3073 | No |
| 27 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 10192 | -0.057 | 0.2366 | No |