Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | PURINE_METABOLISM |
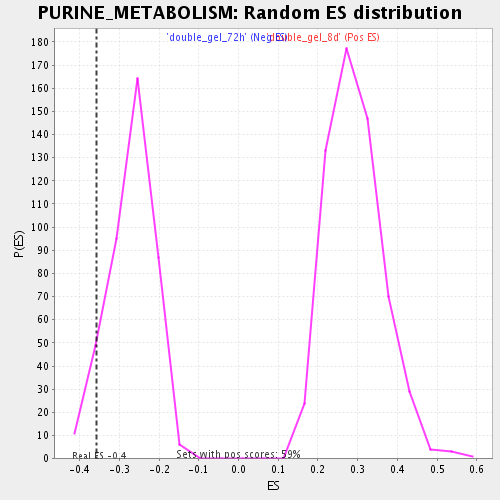
| Enrichment Score (ES) | -0.35851946 |
| Normalized Enrichment Score (NES) | -1.3256985 |
| Nominal p-value | 0.07038835 |
| FDR q-value | 0.54299027 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 180 | 0.337 | 0.0519 | No |
| 2 | GDA | GDA Entrez, Source | guanine deaminase | 342 | 0.233 | 0.0849 | No |
| 3 | AMPD1 | AMPD1 Entrez, Source | adenosine monophosphate deaminase 1 (isoform M) | 667 | 0.155 | 0.0907 | No |
| 4 | GUCY1B3 | GUCY1B3 Entrez, Source | guanylate cyclase 1, soluble, beta 3 | 801 | 0.140 | 0.1078 | No |
| 5 | PDE4C | PDE4C Entrez, Source | phosphodiesterase 4C, cAMP-specific (phosphodiesterase E1 dunce homolog, Drosophila) | 941 | 0.126 | 0.1217 | No |
| 6 | GUK1 | GUK1 Entrez, Source | guanylate kinase 1 | 1568 | 0.091 | 0.0921 | No |
| 7 | NUDT2 | NUDT2 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 2 | 2160 | 0.068 | 0.0608 | No |
| 8 | POLR2G | POLR2G Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide G | 2604 | 0.057 | 0.0386 | No |
| 9 | GUCY1A3 | GUCY1A3 Entrez, Source | guanylate cyclase 1, soluble, alpha 3 | 3107 | 0.046 | 0.0098 | No |
| 10 | POLG | POLG Entrez, Source | polymerase (DNA directed), gamma | 3170 | 0.045 | 0.0139 | No |
| 11 | FHIT | FHIT Entrez, Source | fragile histidine triad gene | 3384 | 0.041 | 0.0057 | No |
| 12 | ENTPD2 | ENTPD2 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 2 | 4219 | 0.027 | -0.0519 | No |
| 13 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 4788 | 0.018 | -0.0912 | No |
| 14 | AK1 | AK1 Entrez, Source | adenylate kinase 1 | 4876 | 0.017 | -0.0944 | No |
| 15 | ADCY2 | ADCY2 Entrez, Source | adenylate cyclase 2 (brain) | 4922 | 0.016 | -0.0946 | No |
| 16 | GUCY2C | GUCY2C Entrez, Source | guanylate cyclase 2C (heat stable enterotoxin receptor) | 5109 | 0.014 | -0.1060 | No |
| 17 | PDE4A | PDE4A Entrez, Source | phosphodiesterase 4A, cAMP-specific (phosphodiesterase E2 dunce homolog, Drosophila) | 5171 | 0.013 | -0.1080 | No |
| 18 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 5742 | 0.005 | -0.1500 | No |
| 19 | POLR2F | POLR2F Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide F | 5958 | 0.003 | -0.1657 | No |
| 20 | ADCY6 | ADCY6 Entrez, Source | adenylate cyclase 6 | 6028 | 0.002 | -0.1705 | No |
| 21 | GUCY2D | GUCY2D Entrez, Source | guanylate cyclase 2D, membrane (retina-specific) | 6484 | -0.004 | -0.2040 | No |
| 22 | ADCY5 | ADCY5 Entrez, Source | adenylate cyclase 5 | 6683 | -0.007 | -0.2176 | No |
| 23 | ATP5J | ATP5J Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 | 6818 | -0.008 | -0.2261 | No |
| 24 | PDE8A | PDE8A Entrez, Source | phosphodiesterase 8A | 6843 | -0.009 | -0.2262 | No |
| 25 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 6946 | -0.010 | -0.2319 | No |
| 26 | ATP5B | ATP5B Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 7012 | -0.011 | -0.2347 | No |
| 27 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 7018 | -0.011 | -0.2330 | No |
| 28 | GUCY1B2 | GUCY1B2 Entrez, Source | guanylate cyclase 1, soluble, beta 2 | 7329 | -0.015 | -0.2535 | No |
| 29 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 7449 | -0.016 | -0.2594 | No |
| 30 | ATP5I | ATP5I Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit E | 7717 | -0.020 | -0.2757 | No |
| 31 | NPR1 | NPR1 Entrez, Source | natriuretic peptide receptor A/guanylate cyclase A (atrionatriuretic peptide receptor A) | 7769 | -0.020 | -0.2756 | No |
| 32 | ADK | ADK Entrez, Source | adenosine kinase | 7944 | -0.023 | -0.2843 | No |
| 33 | ADA | ADA Entrez, Source | adenosine deaminase | 8062 | -0.024 | -0.2884 | No |
| 34 | ADCY8 | ADCY8 Entrez, Source | adenylate cyclase 8 (brain) | 8194 | -0.026 | -0.2931 | No |
| 35 | NPR2 | NPR2 Entrez, Source | natriuretic peptide receptor B/guanylate cyclase B (atrionatriuretic peptide receptor B) | 8412 | -0.030 | -0.3037 | No |
| 36 | ADCY3 | ADCY3 Entrez, Source | adenylate cyclase 3 | 8531 | -0.031 | -0.3065 | No |
| 37 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 8663 | -0.033 | -0.3100 | No |
| 38 | ATIC | ATIC Entrez, Source | 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase | 8726 | -0.034 | -0.3081 | No |
| 39 | ATP5H | ATP5H Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit d | 8736 | -0.034 | -0.3021 | No |
| 40 | POLR2C | POLR2C Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide C, 33kDa | 8936 | -0.037 | -0.3100 | No |
| 41 | NME2 | NME2 Entrez, Source | non-metastatic cells 2, protein (NM23B) expressed in | 9131 | -0.039 | -0.3170 | No |
| 42 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 9276 | -0.041 | -0.3198 | No |
| 43 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 9290 | -0.042 | -0.3127 | No |
| 44 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 9537 | -0.046 | -0.3224 | No |
| 45 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 9770 | -0.050 | -0.3302 | No |
| 46 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 9884 | -0.052 | -0.3287 | No |
| 47 | ATP5C1 | ATP5C1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, gamma polypeptide 1 | 9920 | -0.052 | -0.3212 | No |
| 48 | GMPS | GMPS Entrez, Source | guanine monphosphate synthetase | 10172 | -0.056 | -0.3292 | No |
| 49 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10269 | -0.058 | -0.3252 | No |
| 50 | ATP5G1 | ATP5G1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C1 (subunit 9) | 10467 | -0.061 | -0.3281 | No |
| 51 | GUCY1A2 | GUCY1A2 Entrez, Source | guanylate cyclase 1, soluble, alpha 2 | 10872 | -0.070 | -0.3449 | Yes |
| 52 | CANT1 | CANT1 Entrez, Source | calcium activated nucleotidase 1 | 10903 | -0.071 | -0.3334 | Yes |
| 53 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10927 | -0.072 | -0.3212 | Yes |
| 54 | PAICS | PAICS Entrez, Source | phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | 10929 | -0.072 | -0.3074 | Yes |
| 55 | ENTPD1 | ENTPD1 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 1 | 10967 | -0.073 | -0.2961 | Yes |
| 56 | NT5E | NT5E Entrez, Source | 5'-nucleotidase, ecto (CD73) | 11299 | -0.081 | -0.3052 | Yes |
| 57 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 11362 | -0.083 | -0.2938 | Yes |
| 58 | PDE5A | PDE5A Entrez, Source | phosphodiesterase 5A, cGMP-specific | 11495 | -0.086 | -0.2870 | Yes |
| 59 | GUCY2F | GUCY2F Entrez, Source | guanylate cyclase 2F, retinal | 11508 | -0.087 | -0.2711 | Yes |
| 60 | POLB | POLB Entrez, Source | polymerase (DNA directed), beta | 11626 | -0.090 | -0.2625 | Yes |
| 61 | PDE9A | PDE9A Entrez, Source | phosphodiesterase 9A | 11700 | -0.092 | -0.2501 | Yes |
| 62 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 11914 | -0.100 | -0.2469 | Yes |
| 63 | ATP5G2 | ATP5G2 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C2 (subunit 9) | 11934 | -0.100 | -0.2288 | Yes |
| 64 | ADCY4 | ADCY4 Entrez, Source | adenylate cyclase 4 | 12124 | -0.108 | -0.2220 | Yes |
| 65 | PDE1A | PDE1A Entrez, Source | phosphodiesterase 1A, calmodulin-dependent | 12401 | -0.121 | -0.2193 | Yes |
| 66 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 12451 | -0.124 | -0.1990 | Yes |
| 67 | POLR1B | POLR1B Entrez, Source | polymerase (RNA) I polypeptide B, 128kDa | 12576 | -0.133 | -0.1825 | Yes |
| 68 | DCK | DCK Entrez, Source | deoxycytidine kinase | 12784 | -0.151 | -0.1688 | Yes |
| 69 | ENPP3 | ENPP3 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 3 | 12806 | -0.153 | -0.1407 | Yes |
| 70 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 12816 | -0.154 | -0.1114 | Yes |
| 71 | PRPS2 | PRPS2 Entrez, Source | phosphoribosyl pyrophosphate synthetase 2 | 12857 | -0.160 | -0.0834 | Yes |
| 72 | PDE7B | PDE7B Entrez, Source | phosphodiesterase 7B | 12940 | -0.171 | -0.0564 | Yes |
| 73 | PKLR | PKLR Entrez, Source | pyruvate kinase, liver and RBC | 13313 | -0.446 | 0.0022 | Yes |