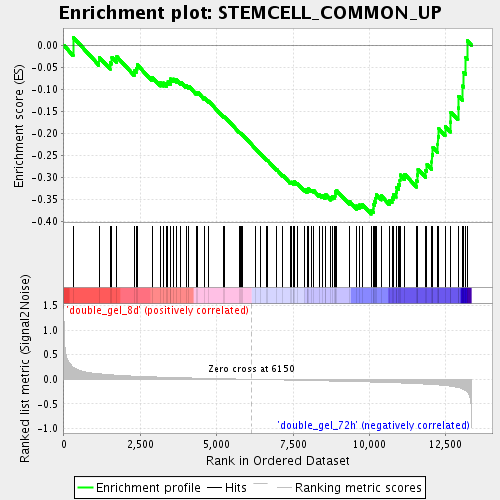
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
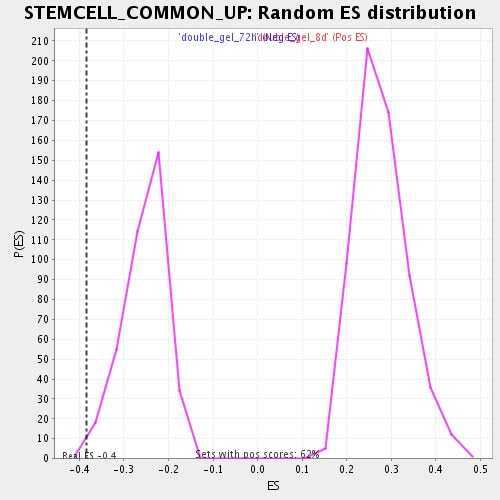
| GeneSet | STEMCELL_COMMON_UP |
| Enrichment Score (ES) | -0.3832868 |
| Normalized Enrichment Score (NES) | -1.5108733 |
| Nominal p-value | 0.007978723 |
| FDR q-value | 0.3188124 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC38A2 | SLC38A2 Entrez, Source | solute carrier family 38, member 2 | 302 | 0.248 | 0.0188 | No |
| 2 | KLHL7 | KLHL7 Entrez, Source | kelch-like 7 (Drosophila) | 1148 | 0.112 | -0.0263 | No |
| 3 | PANX3 | PANX3 Entrez, Source | pannexin 3 | 1514 | 0.093 | -0.0382 | No |
| 4 | LAPTM4A | LAPTM4A Entrez, Source | lysosomal-associated protein transmembrane 4 alpha | 1569 | 0.091 | -0.0271 | No |
| 5 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 1731 | 0.083 | -0.0253 | No |
| 6 | PEX7 | PEX7 Entrez, Source | peroxisomal biogenesis factor 7 | 2300 | 0.064 | -0.0574 | No |
| 7 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 2366 | 0.063 | -0.0518 | No |
| 8 | MPDU1 | MPDU1 Entrez, Source | mannose-P-dolichol utilization defect 1 | 2393 | 0.062 | -0.0434 | No |
| 9 | PIGX | PIGX Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class X | 2887 | 0.051 | -0.0720 | No |
| 10 | SUCLG2 | SUCLG2 Entrez, Source | succinate-CoA ligase, GDP-forming, beta subunit | 3155 | 0.045 | -0.0846 | No |
| 11 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 3258 | 0.044 | -0.0850 | No |
| 12 | PPP1R2 | PPP1R2 Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 2 | 3368 | 0.041 | -0.0864 | No |
| 13 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 3405 | 0.040 | -0.0824 | No |
| 14 | UPP1 | UPP1 Entrez, Source | uridine phosphorylase 1 | 3489 | 0.039 | -0.0821 | No |
| 15 | CTTN | CTTN Entrez, Source | cortactin | 3493 | 0.039 | -0.0759 | No |
| 16 | RNF4 | RNF4 Entrez, Source | ring finger protein 4 | 3580 | 0.037 | -0.0761 | No |
| 17 | YWHAB | YWHAB Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide | 3679 | 0.036 | -0.0776 | No |
| 18 | MTERFD1 | MTERFD1 Entrez, Source | MTERF domain containing 1 | 3825 | 0.033 | -0.0830 | No |
| 19 | GSTA4 | GSTA4 Entrez, Source | glutathione S-transferase A4 | 3997 | 0.030 | -0.0909 | No |
| 20 | EIF4EBP1 | EIF4EBP1 Entrez, Source | eukaryotic translation initiation factor 4E binding protein 1 | 4091 | 0.029 | -0.0931 | No |
| 21 | TPP2 | TPP2 Entrez, Source | tripeptidyl peptidase II | 4336 | 0.025 | -0.1073 | No |
| 22 | TBC1D15 | TBC1D15 Entrez, Source | TBC1 domain family, member 15 | 4380 | 0.024 | -0.1065 | No |
| 23 | KCNAB3 | KCNAB3 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, beta member 3 | 4593 | 0.021 | -0.1190 | No |
| 24 | TXNL1 | TXNL1 Entrez, Source | thioredoxin-like 1 | 4730 | 0.019 | -0.1260 | No |
| 25 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 5229 | 0.012 | -0.1616 | No |
| 26 | FHL1 | FHL1 Entrez, Source | four and a half LIM domains 1 | 5266 | 0.012 | -0.1624 | No |
| 27 | MARCH7 | MARCH7 Entrez, Source | membrane-associated ring finger (C3HC4) 7 | 5740 | 0.005 | -0.1972 | No |
| 28 | PSMD12 | PSMD12 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 12 | 5781 | 0.005 | -0.1995 | No |
| 29 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 5809 | 0.004 | -0.2008 | No |
| 30 | PLS3 | PLS3 Entrez, Source | plastin 3 (T isoform) | 5853 | 0.004 | -0.2034 | No |
| 31 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 6253 | -0.001 | -0.2333 | No |
| 32 | STXBP3 | STXBP3 Entrez, Source | syntaxin binding protein 3 | 6429 | -0.004 | -0.2459 | No |
| 33 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 6644 | -0.006 | -0.2610 | No |
| 34 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 6658 | -0.007 | -0.2609 | No |
| 35 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 6967 | -0.010 | -0.2824 | No |
| 36 | STATIP1 | STATIP1 Entrez, Source | signal transducer and activator of transcription 3 interacting protein 1 | 7167 | -0.013 | -0.2953 | No |
| 37 | TXNDC9 | TXNDC9 Entrez, Source | thioredoxin domain containing 9 | 7402 | -0.015 | -0.3104 | No |
| 38 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 7437 | -0.016 | -0.3103 | No |
| 39 | MRPL34 | MRPL34 Entrez, Source | mitochondrial ribosomal protein L34 | 7509 | -0.017 | -0.3129 | No |
| 40 | ZMAT3 | ZMAT3 Entrez, Source | zinc finger, matrin type 3 | 7516 | -0.017 | -0.3105 | No |
| 41 | MRPL17 | MRPL17 Entrez, Source | mitochondrial ribosomal protein L17 | 7537 | -0.017 | -0.3091 | No |
| 42 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 7629 | -0.019 | -0.3129 | No |
| 43 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 7870 | -0.022 | -0.3274 | No |
| 44 | RAD23B | RAD23B Entrez, Source | RAD23 homolog B (S. cerevisiae) | 7961 | -0.023 | -0.3304 | No |
| 45 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 7976 | -0.023 | -0.3275 | No |
| 46 | NUP35 | NUP35 Entrez, Source | nucleoporin 35kDa | 8003 | -0.024 | -0.3256 | No |
| 47 | USP10 | USP10 Entrez, Source | ubiquitin specific peptidase 10 | 8103 | -0.025 | -0.3289 | No |
| 48 | RYK | RYK Entrez, Source | RYK receptor-like tyrosine kinase | 8165 | -0.026 | -0.3291 | No |
| 49 | TBRG4 | TBRG4 Entrez, Source | transforming growth factor beta regulator 4 | 8364 | -0.029 | -0.3392 | No |
| 50 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 8447 | -0.030 | -0.3404 | No |
| 51 | ARCN1 | ARCN1 Entrez, Source | archain 1 | 8549 | -0.031 | -0.3428 | No |
| 52 | ZCCHC10 | ZCCHC10 Entrez, Source | zinc finger, CCHC domain containing 10 | 8571 | -0.032 | -0.3390 | No |
| 53 | PLA2G6 | PLA2G6 Entrez, Source | phospholipase A2, group VI (cytosolic, calcium-independent) | 8737 | -0.034 | -0.3458 | No |
| 54 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 8780 | -0.035 | -0.3432 | No |
| 55 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 8847 | -0.035 | -0.3423 | No |
| 56 | LAPTM4B | LAPTM4B Entrez, Source | lysosomal associated protein transmembrane 4 beta | 8874 | -0.036 | -0.3382 | No |
| 57 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 8876 | -0.036 | -0.3323 | No |
| 58 | SLC4A7 | SLC4A7 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 7 | 8929 | -0.037 | -0.3301 | No |
| 59 | RPUSD4 | RPUSD4 Entrez, Source | RNA pseudouridylate synthase domain containing 4 | 9355 | -0.043 | -0.3551 | No |
| 60 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 9586 | -0.047 | -0.3646 | No |
| 61 | TARSL1 | TARSL1 Entrez, Source | threonyl-tRNA synthetase-like 1 | 9663 | -0.048 | -0.3623 | No |
| 62 | IMPAD1 | IMPAD1 Entrez, Source | inositol monophosphatase domain containing 1 | 9755 | -0.049 | -0.3609 | No |
| 63 | FKBP9 | FKBP9 Entrez, Source | FK506 binding protein 9, 63 kDa | 10052 | -0.054 | -0.3742 | Yes |
| 64 | SEC23IP | SEC23IP Entrez, Source | SEC23 interacting protein | 10132 | -0.056 | -0.3708 | Yes |
| 65 | COPS4 | COPS4 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 4 (Arabidopsis) | 10134 | -0.056 | -0.3616 | Yes |
| 66 | FBXO8 | FBXO8 Entrez, Source | F-box protein 8 | 10173 | -0.056 | -0.3550 | Yes |
| 67 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 10199 | -0.057 | -0.3475 | Yes |
| 68 | ARIH1 | ARIH1 Entrez, Source | ariadne homolog, ubiquitin-conjugating enzyme E2 binding protein, 1 (Drosophila) | 10214 | -0.057 | -0.3390 | Yes |
| 69 | RABGGTB | RABGGTB Entrez, Source | Rab geranylgeranyltransferase, beta subunit | 10378 | -0.060 | -0.3413 | Yes |
| 70 | GRWD1 | GRWD1 Entrez, Source | glutamate-rich WD repeat containing 1 | 10659 | -0.065 | -0.3515 | Yes |
| 71 | PRPSAP1 | PRPSAP1 Entrez, Source | phosphoribosyl pyrophosphate synthetase-associated protein 1 | 10740 | -0.067 | -0.3463 | Yes |
| 72 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 10785 | -0.068 | -0.3383 | Yes |
| 73 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 10870 | -0.070 | -0.3329 | Yes |
| 74 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 10886 | -0.070 | -0.3222 | Yes |
| 75 | MKI67IP | MKI67IP Entrez, Source | MKI67 (FHA domain) interacting nucleolar phosphoprotein | 10951 | -0.072 | -0.3150 | Yes |
| 76 | RCL1 | RCL1 Entrez, Source | RNA terminal phosphate cyclase-like 1 | 10995 | -0.073 | -0.3060 | Yes |
| 77 | CIRH1A | CIRH1A Entrez, Source | cirrhosis, autosomal recessive 1A (cirhin) | 11001 | -0.073 | -0.2941 | Yes |
| 78 | LSG1 | LSG1 Entrez, Source | large subunit GTPase 1 homolog (S. cerevisiae) | 11150 | -0.077 | -0.2924 | Yes |
| 79 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 11543 | -0.088 | -0.3073 | Yes |
| 80 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 11564 | -0.088 | -0.2941 | Yes |
| 81 | CRSP3 | CRSP3 Entrez, Source | cofactor required for Sp1 transcriptional activation, subunit 3, 130kDa | 11586 | -0.089 | -0.2808 | Yes |
| 82 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 11841 | -0.097 | -0.2837 | Yes |
| 83 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 11880 | -0.098 | -0.2701 | Yes |
| 84 | MAP4K3 | MAP4K3 Entrez, Source | mitogen-activated protein kinase kinase kinase kinase 3 | 12041 | -0.104 | -0.2647 | Yes |
| 85 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 12046 | -0.105 | -0.2475 | Yes |
| 86 | RPL22 | RPL22 Entrez, Source | ribosomal protein L22 | 12075 | -0.106 | -0.2319 | Yes |
| 87 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 12235 | -0.113 | -0.2251 | Yes |
| 88 | PDCD2 | PDCD2 Entrez, Source | programmed cell death 2 | 12242 | -0.113 | -0.2066 | Yes |
| 89 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 12268 | -0.115 | -0.1893 | Yes |
| 90 | RFFL | RFFL Entrez, Source | ring finger and FYVE-like domain containing 1 | 12475 | -0.126 | -0.1838 | Yes |
| 91 | RNF138 | RNF138 Entrez, Source | ring finger protein 138 | 12649 | -0.138 | -0.1738 | Yes |
| 92 | FKBP11 | FKBP11 Entrez, Source | FK506 binding protein 11, 19 kDa | 12662 | -0.138 | -0.1516 | Yes |
| 93 | XTP3TPA | XTP3TPA Entrez, Source | - | 12898 | -0.165 | -0.1418 | Yes |
| 94 | GNL2 | GNL2 Entrez, Source | guanine nucleotide binding protein-like 2 (nucleolar) | 12920 | -0.167 | -0.1154 | Yes |
| 95 | DPH5 | DPH5 Entrez, Source | DPH5 homolog (S. cerevisiae) | 13045 | -0.196 | -0.0919 | Yes |
| 96 | GCAT | GCAT Entrez, Source | glycine C-acetyltransferase (2-amino-3-ketobutyrate coenzyme A ligase) | 13084 | -0.205 | -0.0605 | Yes |
| 97 | DICER1 | DICER1 Entrez, Source | Dicer1, Dcr-1 homolog (Drosophila) | 13152 | -0.225 | -0.0279 | Yes |
| 98 | PPIC | PPIC Entrez, Source | peptidylprolyl isomerase C (cyclophilin C) | 13197 | -0.252 | 0.0109 | Yes |