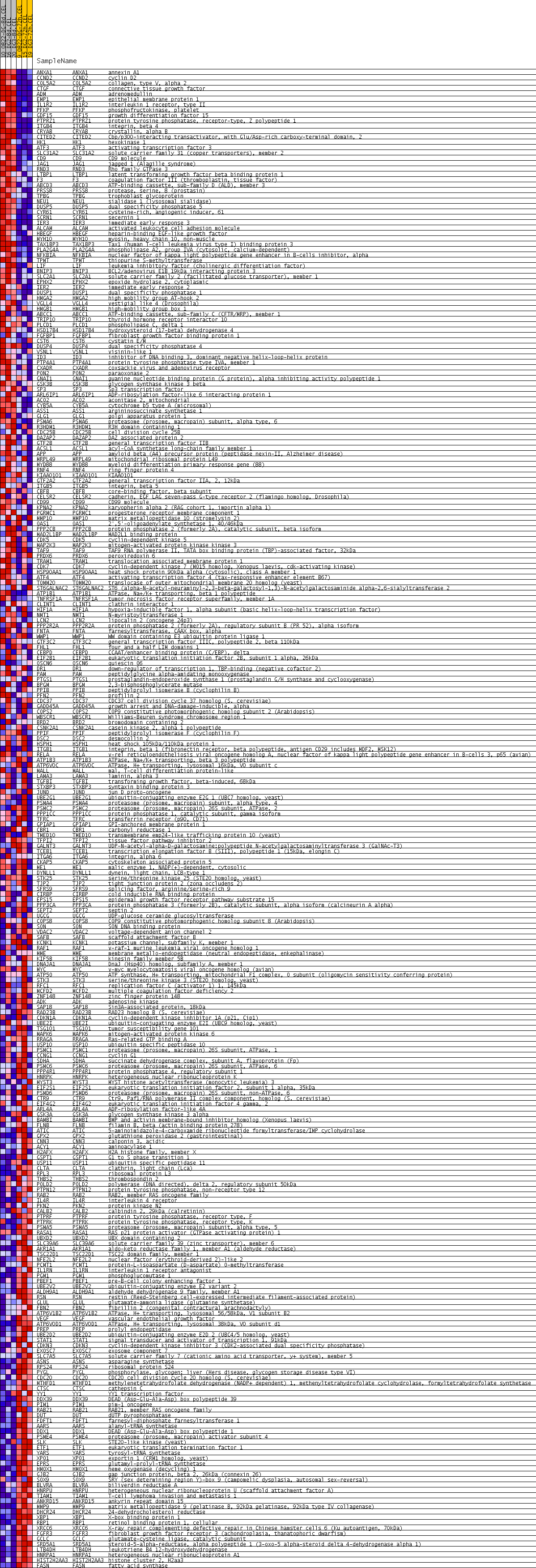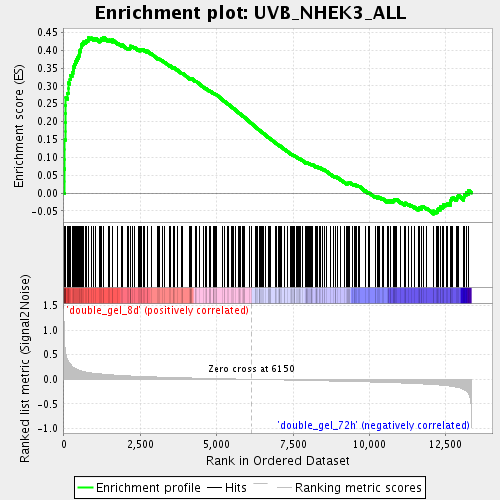
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | UVB_NHEK3_ALL |
| Enrichment Score (ES) | 0.4361821 |
| Normalized Enrichment Score (NES) | 1.801865 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.03201514 |
| FWER p-Value | 0.969 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 10 | 0.815 | 0.0334 | Yes |
| 2 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 11 | 0.815 | 0.0675 | Yes |
| 3 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 28 | 0.668 | 0.0943 | Yes |
| 4 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 32 | 0.660 | 0.1217 | Yes |
| 5 | ADM | ADM Entrez, Source | adrenomedullin | 34 | 0.649 | 0.1488 | Yes |
| 6 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 44 | 0.606 | 0.1735 | Yes |
| 7 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 46 | 0.599 | 0.1985 | Yes |
| 8 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 52 | 0.576 | 0.2222 | Yes |
| 9 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 58 | 0.559 | 0.2453 | Yes |
| 10 | PTPRZ1 | PTPRZ1 Entrez, Source | protein tyrosine phosphatase, receptor-type, Z polypeptide 1 | 65 | 0.523 | 0.2667 | Yes |
| 11 | ITGB4 | ITGB4 Entrez, Source | integrin, beta 4 | 121 | 0.401 | 0.2793 | Yes |
| 12 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 141 | 0.371 | 0.2934 | Yes |
| 13 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 145 | 0.367 | 0.3085 | Yes |
| 14 | HK1 | HK1 Entrez, Source | hexokinase 1 | 197 | 0.323 | 0.3181 | Yes |
| 15 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 218 | 0.309 | 0.3296 | Yes |
| 16 | SLC31A2 | SLC31A2 Entrez, Source | solute carrier family 31 (copper transporters), member 2 | 268 | 0.267 | 0.3370 | Yes |
| 17 | CD9 | CD9 Entrez, Source | CD9 molecule | 299 | 0.250 | 0.3452 | Yes |
| 18 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 323 | 0.239 | 0.3534 | Yes |
| 19 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 350 | 0.230 | 0.3611 | Yes |
| 20 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 372 | 0.219 | 0.3686 | Yes |
| 21 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 418 | 0.205 | 0.3738 | Yes |
| 22 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 451 | 0.195 | 0.3795 | Yes |
| 23 | PRSS8 | PRSS8 Entrez, Source | protease, serine, 8 (prostasin) | 490 | 0.183 | 0.3843 | Yes |
| 24 | TPBG | TPBG Entrez, Source | trophoblast glycoprotein | 503 | 0.182 | 0.3909 | Yes |
| 25 | NEU1 | NEU1 Entrez, Source | sialidase 1 (lysosomal sialidase) | 510 | 0.181 | 0.3981 | Yes |
| 26 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 553 | 0.172 | 0.4020 | Yes |
| 27 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 559 | 0.171 | 0.4088 | Yes |
| 28 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 568 | 0.170 | 0.4153 | Yes |
| 29 | IER3 | IER3 Entrez, Source | immediate early response 3 | 598 | 0.165 | 0.4200 | Yes |
| 30 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 646 | 0.158 | 0.4231 | Yes |
| 31 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 717 | 0.149 | 0.4239 | Yes |
| 32 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 744 | 0.145 | 0.4280 | Yes |
| 33 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 791 | 0.140 | 0.4304 | Yes |
| 34 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 812 | 0.139 | 0.4347 | Yes |
| 35 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 888 | 0.130 | 0.4344 | Yes |
| 36 | TPMT | TPMT Entrez, Source | thiopurine S-methyltransferase | 980 | 0.123 | 0.4326 | Yes |
| 37 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 1037 | 0.118 | 0.4332 | Yes |
| 38 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 1177 | 0.111 | 0.4272 | Yes |
| 39 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 1212 | 0.109 | 0.4292 | Yes |
| 40 | EPHX2 | EPHX2 Entrez, Source | epoxide hydrolase 2, cytoplasmic | 1222 | 0.108 | 0.4330 | Yes |
| 41 | IER2 | IER2 Entrez, Source | immediate early response 2 | 1295 | 0.104 | 0.4319 | Yes |
| 42 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1297 | 0.104 | 0.4362 | Yes |
| 43 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 1465 | 0.096 | 0.4274 | No |
| 44 | VGLL4 | VGLL4 Entrez, Source | vestigial like 4 (Drosophila) | 1500 | 0.094 | 0.4287 | No |
| 45 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 1582 | 0.090 | 0.4263 | No |
| 46 | ABCC1 | ABCC1 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 1 | 1583 | 0.090 | 0.4301 | No |
| 47 | TRIP10 | TRIP10 Entrez, Source | thyroid hormone receptor interactor 10 | 1763 | 0.082 | 0.4198 | No |
| 48 | PLCD1 | PLCD1 Entrez, Source | phospholipase C, delta 1 | 1899 | 0.077 | 0.4127 | No |
| 49 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 1910 | 0.076 | 0.4151 | No |
| 50 | FGFBP1 | FGFBP1 Entrez, Source | fibroblast growth factor binding protein 1 | 2072 | 0.071 | 0.4057 | No |
| 51 | CST6 | CST6 Entrez, Source | cystatin E/M | 2108 | 0.069 | 0.4060 | No |
| 52 | DUSP4 | DUSP4 Entrez, Source | dual specificity phosphatase 4 | 2162 | 0.068 | 0.4048 | No |
| 53 | VSNL1 | VSNL1 Entrez, Source | visinin-like 1 | 2169 | 0.068 | 0.4072 | No |
| 54 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 2181 | 0.068 | 0.4092 | No |
| 55 | PTP4A1 | PTP4A1 Entrez, Source | protein tyrosine phosphatase type IVA, member 1 | 2184 | 0.068 | 0.4118 | No |
| 56 | CXADR | CXADR Entrez, Source | coxsackie virus and adenovirus receptor | 2231 | 0.066 | 0.4111 | No |
| 57 | PON2 | PON2 Entrez, Source | paraoxonase 2 | 2308 | 0.064 | 0.4080 | No |
| 58 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 2424 | 0.061 | 0.4017 | No |
| 59 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 2479 | 0.060 | 0.4001 | No |
| 60 | SP3 | SP3 Entrez, Source | Sp3 transcription factor | 2512 | 0.059 | 0.4002 | No |
| 61 | ARL6IP1 | ARL6IP1 Entrez, Source | ADP-ribosylation factor-like 6 interacting protein 1 | 2527 | 0.059 | 0.4016 | No |
| 62 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 2544 | 0.059 | 0.4028 | No |
| 63 | CYB5A | CYB5A Entrez, Source | cytochrome b5 type A (microsomal) | 2595 | 0.058 | 0.4014 | No |
| 64 | ASS1 | ASS1 Entrez, Source | argininosuccinate synthetase 1 | 2650 | 0.056 | 0.3996 | No |
| 65 | GLG1 | GLG1 Entrez, Source | golgi apparatus protein 1 | 2729 | 0.054 | 0.3959 | No |
| 66 | PSMA6 | PSMA6 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 6 | 2734 | 0.054 | 0.3979 | No |
| 67 | R3HDM1 | R3HDM1 Entrez, Source | R3H domain containing 1 | 2857 | 0.052 | 0.3907 | No |
| 68 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 3066 | 0.047 | 0.3768 | No |
| 69 | DAZAP2 | DAZAP2 Entrez, Source | DAZ associated protein 2 | 3109 | 0.046 | 0.3755 | No |
| 70 | GTF2B | GTF2B Entrez, Source | general transcription factor IIB | 3140 | 0.046 | 0.3751 | No |
| 71 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 3216 | 0.044 | 0.3712 | No |
| 72 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 3298 | 0.043 | 0.3668 | No |
| 73 | MRPL49 | MRPL49 Entrez, Source | mitochondrial ribosomal protein L49 | 3463 | 0.039 | 0.3559 | No |
| 74 | MYD88 | MYD88 Entrez, Source | myeloid differentiation primary response gene (88) | 3471 | 0.039 | 0.3570 | No |
| 75 | RNF4 | RNF4 Entrez, Source | ring finger protein 4 | 3580 | 0.037 | 0.3503 | No |
| 76 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 3593 | 0.037 | 0.3509 | No |
| 77 | GTF2A2 | GTF2A2 Entrez, Source | general transcription factor IIA, 2, 12kDa | 3615 | 0.037 | 0.3508 | No |
| 78 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 3714 | 0.035 | 0.3448 | No |
| 79 | CBFB | CBFB Entrez, Source | core-binding factor, beta subunit | 3853 | 0.032 | 0.3356 | No |
| 80 | CELSR2 | CELSR2 Entrez, Source | cadherin, EGF LAG seven-pass G-type receptor 2 (flamingo homolog, Drosophila) | 3880 | 0.032 | 0.3349 | No |
| 81 | CD99 | CD99 Entrez, Source | CD99 molecule | 3891 | 0.032 | 0.3355 | No |
| 82 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 4109 | 0.029 | 0.3201 | No |
| 83 | PGRMC1 | PGRMC1 Entrez, Source | progesterone receptor membrane component 1 | 4136 | 0.028 | 0.3193 | No |
| 84 | MMP10 | MMP10 Entrez, Source | matrix metallopeptidase 10 (stromelysin 2) | 4140 | 0.028 | 0.3202 | No |
| 85 | OAS1 | OAS1 Entrez, Source | 2',5'-oligoadenylate synthetase 1, 40/46kDa | 4150 | 0.028 | 0.3207 | No |
| 86 | PPP2CB | PPP2CB Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, beta isoform | 4166 | 0.028 | 0.3207 | No |
| 87 | MAD2L1BP | MAD2L1BP Entrez, Source | MAD2L1 binding protein | 4190 | 0.027 | 0.3201 | No |
| 88 | CDK5 | CDK5 Entrez, Source | cyclin-dependent kinase 5 | 4304 | 0.025 | 0.3125 | No |
| 89 | MAP2K3 | MAP2K3 Entrez, Source | mitogen-activated protein kinase kinase 3 | 4321 | 0.025 | 0.3124 | No |
| 90 | TAF9 | TAF9 Entrez, Source | TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 32kDa | 4322 | 0.025 | 0.3134 | No |
| 91 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 4430 | 0.023 | 0.3062 | No |
| 92 | TRAM1 | TRAM1 Entrez, Source | translocation associated membrane protein 1 | 4581 | 0.021 | 0.2956 | No |
| 93 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 4639 | 0.020 | 0.2921 | No |
| 94 | HSP90AA1 | HSP90AA1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class A member 1 | 4655 | 0.020 | 0.2918 | No |
| 95 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 4662 | 0.020 | 0.2922 | No |
| 96 | TOMM20 | TOMM20 Entrez, Source | translocase of outer mitochondrial membrane 20 homolog (yeast) | 4757 | 0.019 | 0.2858 | No |
| 97 | ST6GALNAC2 | ST6GALNAC2 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 2 | 4765 | 0.019 | 0.2860 | No |
| 98 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 4788 | 0.018 | 0.2851 | No |
| 99 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 4794 | 0.018 | 0.2855 | No |
| 100 | CLINT1 | CLINT1 Entrez, Source | clathrin interactor 1 | 4885 | 0.017 | 0.2793 | No |
| 101 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 4892 | 0.017 | 0.2796 | No |
| 102 | NMT1 | NMT1 Entrez, Source | N-myristoyltransferase 1 | 4905 | 0.017 | 0.2793 | No |
| 103 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 4934 | 0.016 | 0.2779 | No |
| 104 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 4968 | 0.016 | 0.2760 | No |
| 105 | FNTA | FNTA Entrez, Source | farnesyltransferase, CAAX box, alpha | 4985 | 0.015 | 0.2754 | No |
| 106 | WWP1 | WWP1 Entrez, Source | WW domain containing E3 ubiquitin protein ligase 1 | 4986 | 0.015 | 0.2761 | No |
| 107 | GTF3C2 | GTF3C2 Entrez, Source | general transcription factor IIIC, polypeptide 2, beta 110kDa | 5178 | 0.013 | 0.2620 | No |
| 108 | FHL1 | FHL1 Entrez, Source | four and a half LIM domains 1 | 5266 | 0.012 | 0.2558 | No |
| 109 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 5268 | 0.012 | 0.2562 | No |
| 110 | EIF2B1 | EIF2B1 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 1 alpha, 26kDa | 5340 | 0.011 | 0.2512 | No |
| 111 | QSCN6 | QSCN6 Entrez, Source | quiescin Q6 | 5385 | 0.010 | 0.2483 | No |
| 112 | DR1 | DR1 Entrez, Source | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | 5479 | 0.009 | 0.2415 | No |
| 113 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 5501 | 0.008 | 0.2403 | No |
| 114 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 5536 | 0.008 | 0.2380 | No |
| 115 | BPGM | BPGM Entrez, Source | 2,3-bisphosphoglycerate mutase | 5600 | 0.007 | 0.2335 | No |
| 116 | PPIB | PPIB Entrez, Source | peptidylprolyl isomerase B (cyclophilin B) | 5713 | 0.006 | 0.2251 | No |
| 117 | PFN2 | PFN2 Entrez, Source | profilin 2 | 5738 | 0.005 | 0.2235 | No |
| 118 | CDC37 | CDC37 Entrez, Source | CDC37 cell division cycle 37 homolog (S. cerevisiae) | 5747 | 0.005 | 0.2231 | No |
| 119 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 5788 | 0.005 | 0.2202 | No |
| 120 | COPS2 | COPS2 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 2 (Arabidopsis) | 5856 | 0.004 | 0.2153 | No |
| 121 | WBSCR1 | WBSCR1 Entrez, Source | Williams-Beuren syndrome chromosome region 1 | 5863 | 0.004 | 0.2150 | No |
| 122 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 5876 | 0.004 | 0.2142 | No |
| 123 | CSNK2A1 | CSNK2A1 Entrez, Source | casein kinase 2, alpha 1 polypeptide | 5898 | 0.003 | 0.2127 | No |
| 124 | PPIF | PPIF Entrez, Source | peptidylprolyl isomerase F (cyclophilin F) | 5913 | 0.003 | 0.2118 | No |
| 125 | DSC2 | DSC2 Entrez, Source | desmocollin 2 | 6067 | 0.001 | 0.2001 | No |
| 126 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 6127 | 0.000 | 0.1956 | No |
| 127 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 6253 | -0.001 | 0.1861 | No |
| 128 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 6260 | -0.001 | 0.1857 | No |
| 129 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 6301 | -0.002 | 0.1827 | No |
| 130 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 6333 | -0.002 | 0.1805 | No |
| 131 | MALL | MALL Entrez, Source | mal, T-cell differentiation protein-like | 6400 | -0.003 | 0.1755 | No |
| 132 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 6404 | -0.003 | 0.1754 | No |
| 133 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 6410 | -0.003 | 0.1752 | No |
| 134 | STXBP3 | STXBP3 Entrez, Source | syntaxin binding protein 3 | 6429 | -0.004 | 0.1740 | No |
| 135 | JUND | JUND Entrez, Source | jun D proto-oncogene | 6457 | -0.004 | 0.1721 | No |
| 136 | UBE2G1 | UBE2G1 Entrez, Source | ubiquitin-conjugating enzyme E2G 1 (UBC7 homolog, yeast) | 6486 | -0.004 | 0.1701 | No |
| 137 | PSMA4 | PSMA4 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 4 | 6523 | -0.005 | 0.1675 | No |
| 138 | PSMC2 | PSMC2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 2 | 6594 | -0.006 | 0.1624 | No |
| 139 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 6696 | -0.007 | 0.1550 | No |
| 140 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 6700 | -0.007 | 0.1550 | No |
| 141 | GPIAP1 | GPIAP1 Entrez, Source | GPI-anchored membrane protein 1 | 6728 | -0.007 | 0.1533 | No |
| 142 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 6731 | -0.007 | 0.1535 | No |
| 143 | TMED10 | TMED10 Entrez, Source | transmembrane emp24-like trafficking protein 10 (yeast) | 6770 | -0.008 | 0.1509 | No |
| 144 | TFPI2 | TFPI2 Entrez, Source | tissue factor pathway inhibitor 2 | 6771 | -0.008 | 0.1512 | No |
| 145 | GALNT3 | GALNT3 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3) | 6927 | -0.010 | 0.1397 | No |
| 146 | TCEB1 | TCEB1 Entrez, Source | transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) | 6931 | -0.010 | 0.1399 | No |
| 147 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 6967 | -0.010 | 0.1377 | No |
| 148 | CKAP5 | CKAP5 Entrez, Source | cytoskeleton associated protein 5 | 7020 | -0.011 | 0.1342 | No |
| 149 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 7060 | -0.011 | 0.1317 | No |
| 150 | DYNLL1 | DYNLL1 Entrez, Source | dynein, light chain, LC8-type 1 | 7062 | -0.011 | 0.1321 | No |
| 151 | STK25 | STK25 Entrez, Source | serine/threonine kinase 25 (STE20 homolog, yeast) | 7067 | -0.011 | 0.1322 | No |
| 152 | TJP2 | TJP2 Entrez, Source | tight junction protein 2 (zona occludens 2) | 7094 | -0.012 | 0.1307 | No |
| 153 | SFRS9 | SFRS9 Entrez, Source | splicing factor, arginine/serine-rich 9 | 7120 | -0.012 | 0.1293 | No |
| 154 | CIRBP | CIRBP Entrez, Source | cold inducible RNA binding protein | 7213 | -0.013 | 0.1228 | No |
| 155 | EPS15 | EPS15 Entrez, Source | epidermal growth factor receptor pathway substrate 15 | 7217 | -0.013 | 0.1232 | No |
| 156 | PPP3CA | PPP3CA Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, alpha isoform (calcineurin A alpha) | 7318 | -0.014 | 0.1161 | No |
| 157 | SEPT2 | SEPT2 Entrez, Source | septin 2 | 7427 | -0.016 | 0.1085 | No |
| 158 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 7435 | -0.016 | 0.1086 | No |
| 159 | COPS8 | COPS8 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 8 (Arabidopsis) | 7463 | -0.016 | 0.1073 | No |
| 160 | SON | SON Entrez, Source | SON DNA binding protein | 7481 | -0.016 | 0.1066 | No |
| 161 | VDAC2 | VDAC2 Entrez, Source | voltage-dependent anion channel 2 | 7498 | -0.017 | 0.1061 | No |
| 162 | SAFB | SAFB Entrez, Source | scaffold attachment factor B | 7539 | -0.017 | 0.1038 | No |
| 163 | KCNK1 | KCNK1 Entrez, Source | potassium channel, subfamily K, member 1 | 7542 | -0.017 | 0.1043 | No |
| 164 | RAF1 | RAF1 Entrez, Source | v-raf-1 murine leukemia viral oncogene homolog 1 | 7552 | -0.017 | 0.1044 | No |
| 165 | MME | MME Entrez, Source | membrane metallo-endopeptidase (neutral endopeptidase, enkephalinase) | 7620 | -0.018 | 0.1000 | No |
| 166 | KIF5B | KIF5B Entrez, Source | kinesin family member 5B | 7649 | -0.019 | 0.0987 | No |
| 167 | DNAJA1 | DNAJA1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 1 | 7683 | -0.019 | 0.0969 | No |
| 168 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 7715 | -0.020 | 0.0954 | No |
| 169 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 7722 | -0.020 | 0.0957 | No |
| 170 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 7752 | -0.020 | 0.0944 | No |
| 171 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 7792 | -0.021 | 0.0922 | No |
| 172 | MCFD2 | MCFD2 Entrez, Source | multiple coagulation factor deficiency 2 | 7916 | -0.022 | 0.0838 | No |
| 173 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 7943 | -0.023 | 0.0827 | No |
| 174 | ADK | ADK Entrez, Source | adenosine kinase | 7944 | -0.023 | 0.0837 | No |
| 175 | SAP18 | SAP18 Entrez, Source | Sin3A-associated protein, 18kDa | 7954 | -0.023 | 0.0839 | No |
| 176 | RAD23B | RAD23B Entrez, Source | RAD23 homolog B (S. cerevisiae) | 7961 | -0.023 | 0.0844 | No |
| 177 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 7976 | -0.023 | 0.0843 | No |
| 178 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 8006 | -0.024 | 0.0831 | No |
| 179 | TSG101 | TSG101 Entrez, Source | tumor susceptibility gene 101 | 8034 | -0.024 | 0.0820 | No |
| 180 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 8081 | -0.025 | 0.0796 | No |
| 181 | RRAGA | RRAGA Entrez, Source | Ras-related GTP binding A | 8100 | -0.025 | 0.0792 | No |
| 182 | USP10 | USP10 Entrez, Source | ubiquitin specific peptidase 10 | 8103 | -0.025 | 0.0801 | No |
| 183 | PSMC1 | PSMC1 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 1 | 8118 | -0.025 | 0.0801 | No |
| 184 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 8130 | -0.026 | 0.0803 | No |
| 185 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 8238 | -0.027 | 0.0733 | No |
| 186 | PSMC6 | PSMC6 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 6 | 8252 | -0.027 | 0.0734 | No |
| 187 | PPP4R1 | PPP4R1 Entrez, Source | protein phosphatase 4, regulatory subunit 1 | 8256 | -0.027 | 0.0743 | No |
| 188 | HNRPK | HNRPK Entrez, Source | heterogeneous nuclear ribonucleoprotein K | 8260 | -0.027 | 0.0752 | No |
| 189 | MYST3 | MYST3 Entrez, Source | MYST histone acetyltransferase (monocytic leukemia) 3 | 8296 | -0.028 | 0.0737 | No |
| 190 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 8354 | -0.029 | 0.0706 | No |
| 191 | PSMD6 | PSMD6 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 6 | 8376 | -0.029 | 0.0702 | No |
| 192 | CTR9 | CTR9 Entrez, Source | Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 8385 | -0.029 | 0.0708 | No |
| 193 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 8447 | -0.030 | 0.0673 | No |
| 194 | ARL4A | ARL4A Entrez, Source | ADP-ribosylation factor-like 4A | 8475 | -0.031 | 0.0666 | No |
| 195 | GSK3A | GSK3A Entrez, Source | glycogen synthase kinase 3 alpha | 8514 | -0.031 | 0.0649 | No |
| 196 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 8540 | -0.031 | 0.0643 | No |
| 197 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 8603 | -0.032 | 0.0609 | No |
| 198 | ATIC | ATIC Entrez, Source | 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase | 8726 | -0.034 | 0.0530 | No |
| 199 | GPX2 | GPX2 Entrez, Source | glutathione peroxidase 2 (gastrointestinal) | 8829 | -0.035 | 0.0467 | No |
| 200 | CNN3 | CNN3 Entrez, Source | calponin 3, acidic | 8886 | -0.036 | 0.0439 | No |
| 201 | ACY1 | ACY1 Entrez, Source | aminoacylase 1 | 8889 | -0.036 | 0.0453 | No |
| 202 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 8894 | -0.036 | 0.0465 | No |
| 203 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 8969 | -0.037 | 0.0424 | No |
| 204 | USP11 | USP11 Entrez, Source | ubiquitin specific peptidase 11 | 9055 | -0.038 | 0.0375 | No |
| 205 | CLTA | CLTA Entrez, Source | clathrin, light chain (Lca) | 9172 | -0.040 | 0.0303 | No |
| 206 | RPL3 | RPL3 Entrez, Source | ribosomal protein L3 | 9257 | -0.041 | 0.0256 | No |
| 207 | THBS2 | THBS2 Entrez, Source | thrombospondin 2 | 9270 | -0.041 | 0.0264 | No |
| 208 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 9276 | -0.041 | 0.0277 | No |
| 209 | PTPN12 | PTPN12 Entrez, Source | protein tyrosine phosphatase, non-receptor type 12 | 9303 | -0.042 | 0.0275 | No |
| 210 | RAB2 | RAB2 Entrez, Source | RAB2, member RAS oncogene family | 9314 | -0.042 | 0.0285 | No |
| 211 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 9332 | -0.042 | 0.0289 | No |
| 212 | PKN2 | PKN2 Entrez, Source | protein kinase N2 | 9361 | -0.043 | 0.0286 | No |
| 213 | CALB2 | CALB2 Entrez, Source | calbindin 2, 29kDa (calretinin) | 9436 | -0.044 | 0.0247 | No |
| 214 | PTPRF | PTPRF Entrez, Source | protein tyrosine phosphatase, receptor type, F | 9495 | -0.045 | 0.0222 | No |
| 215 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 9501 | -0.045 | 0.0237 | No |
| 216 | PSMA5 | PSMA5 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 5 | 9533 | -0.046 | 0.0232 | No |
| 217 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 9586 | -0.047 | 0.0212 | No |
| 218 | UBXD2 | UBXD2 Entrez, Source | UBX domain containing 2 | 9639 | -0.048 | 0.0192 | No |
| 219 | SLC39A6 | SLC39A6 Entrez, Source | solute carrier family 39 (zinc transporter), member 6 | 9664 | -0.048 | 0.0194 | No |
| 220 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 9857 | -0.051 | 0.0068 | No |
| 221 | TSC22D1 | TSC22D1 Entrez, Source | TSC22 domain family, member 1 | 9967 | -0.053 | 0.0007 | No |
| 222 | NFE2L2 | NFE2L2 Entrez, Source | nuclear factor (erythroid-derived 2)-like 2 | 10005 | -0.054 | 0.0001 | No |
| 223 | PCMT1 | PCMT1 Entrez, Source | protein-L-isoaspartate (D-aspartate) O-methyltransferase | 10182 | -0.056 | -0.0110 | No |
| 224 | IL1RN | IL1RN Entrez, Source | interleukin 1 receptor antagonist | 10196 | -0.057 | -0.0096 | No |
| 225 | PGM1 | PGM1 Entrez, Source | phosphoglucomutase 1 | 10277 | -0.058 | -0.0133 | No |
| 226 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 10279 | -0.058 | -0.0109 | No |
| 227 | UBE2V2 | UBE2V2 Entrez, Source | ubiquitin-conjugating enzyme E2 variant 2 | 10326 | -0.059 | -0.0120 | No |
| 228 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 10414 | -0.060 | -0.0161 | No |
| 229 | RSN | RSN Entrez, Source | restin (Reed-Steinberg cell-expressed intermediate filament-associated protein) | 10454 | -0.061 | -0.0165 | No |
| 230 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 10574 | -0.064 | -0.0230 | No |
| 231 | FBN2 | FBN2 Entrez, Source | fibrillin 2 (congenital contractural arachnodactyly) | 10607 | -0.064 | -0.0227 | No |
| 232 | ATP6V1B2 | ATP6V1B2 Entrez, Source | ATPase, H+ transporting, lysosomal 56/58kDa, V1 subunit B2 | 10631 | -0.065 | -0.0218 | No |
| 233 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 10686 | -0.066 | -0.0231 | No |
| 234 | ATP6V0D1 | ATP6V0D1 Entrez, Source | ATPase, H+ transporting, lysosomal 38kDa, V0 subunit d1 | 10697 | -0.066 | -0.0211 | No |
| 235 | PREP | PREP Entrez, Source | prolyl endopeptidase | 10771 | -0.068 | -0.0239 | No |
| 236 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 10785 | -0.068 | -0.0220 | No |
| 237 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 10803 | -0.068 | -0.0205 | No |
| 238 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 10809 | -0.069 | -0.0180 | No |
| 239 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 10861 | -0.070 | -0.0190 | No |
| 240 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 10889 | -0.071 | -0.0181 | No |
| 241 | ASNS | ASNS Entrez, Source | asparagine synthetase | 11024 | -0.074 | -0.0252 | No |
| 242 | RPS24 | RPS24 Entrez, Source | ribosomal protein S24 | 11160 | -0.077 | -0.0323 | No |
| 243 | PYGL | PYGL Entrez, Source | phosphorylase, glycogen; liver (Hers disease, glycogen storage disease type VI) | 11176 | -0.078 | -0.0302 | No |
| 244 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 11187 | -0.078 | -0.0277 | No |
| 245 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 11282 | -0.081 | -0.0316 | No |
| 246 | CTSC | CTSC Entrez, Source | cathepsin C | 11373 | -0.083 | -0.0349 | No |
| 247 | YY1 | YY1 Entrez, Source | YY1 transcription factor | 11474 | -0.086 | -0.0390 | No |
| 248 | DDX39 | DDX39 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 | 11591 | -0.089 | -0.0442 | No |
| 249 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 11624 | -0.090 | -0.0429 | No |
| 250 | RAB21 | RAB21 Entrez, Source | RAB21, member RAS oncogene family | 11632 | -0.090 | -0.0397 | No |
| 251 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 11707 | -0.092 | -0.0415 | No |
| 252 | FDFT1 | FDFT1 Entrez, Source | farnesyl-diphosphate farnesyltransferase 1 | 11716 | -0.093 | -0.0382 | No |
| 253 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 11763 | -0.094 | -0.0378 | No |
| 254 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 11880 | -0.098 | -0.0425 | No |
| 255 | PSME4 | PSME4 Entrez, Source | proteasome (prosome, macropain) activator subunit 4 | 12096 | -0.107 | -0.0545 | No |
| 256 | SLK | SLK Entrez, Source | STE20-like kinase (yeast) | 12099 | -0.107 | -0.0502 | No |
| 257 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 12179 | -0.110 | -0.0516 | No |
| 258 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 12233 | -0.113 | -0.0509 | No |
| 259 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 12235 | -0.113 | -0.0463 | No |
| 260 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 12268 | -0.115 | -0.0439 | No |
| 261 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 12319 | -0.117 | -0.0429 | No |
| 262 | GJB2 | GJB2 Entrez, Source | gap junction protein, beta 2, 26kDa (connexin 26) | 12323 | -0.117 | -0.0382 | No |
| 263 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 12395 | -0.121 | -0.0385 | No |
| 264 | BLVRA | BLVRA Entrez, Source | biliverdin reductase A | 12414 | -0.122 | -0.0348 | No |
| 265 | HNRPU | HNRPU Entrez, Source | heterogeneous nuclear ribonucleoprotein U (scaffold attachment factor A) | 12429 | -0.123 | -0.0307 | No |
| 266 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 12504 | -0.128 | -0.0310 | No |
| 267 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 12561 | -0.132 | -0.0298 | No |
| 268 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12642 | -0.137 | -0.0302 | No |
| 269 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 12656 | -0.138 | -0.0254 | No |
| 270 | XBP1 | XBP1 Entrez, Source | X-box binding protein 1 | 12661 | -0.138 | -0.0199 | No |
| 271 | RBP1 | RBP1 Entrez, Source | retinol binding protein 1, cellular | 12674 | -0.139 | -0.0150 | No |
| 272 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12731 | -0.144 | -0.0132 | No |
| 273 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 12848 | -0.158 | -0.0155 | No |
| 274 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 12866 | -0.161 | -0.0101 | No |
| 275 | SRD5A1 | SRD5A1 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 1 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 1) | 12915 | -0.167 | -0.0067 | No |
| 276 | LTB4DH | LTB4DH Entrez, Source | leukotriene B4 12-hydroxydehydrogenase | 13092 | -0.207 | -0.0115 | No |
| 277 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 13096 | -0.208 | -0.0030 | No |
| 278 | HIST2H2AA3 | HIST2H2AA3 Entrez, Source | histone cluster 2, H2aa3 | 13163 | -0.231 | 0.0016 | No |
| 279 | FASN | FASN Entrez, Source | fatty acid synthase | 13241 | -0.288 | 0.0077 | No |