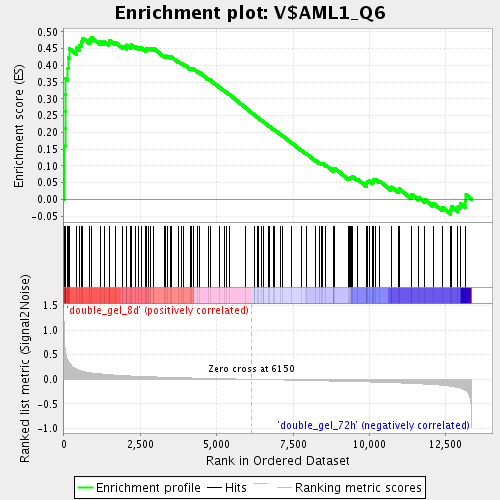
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$AML1_Q6 |
| Enrichment Score (ES) | 0.48394975 |
| Normalized Enrichment Score (NES) | 1.7757794 |
| Nominal p-value | 0.0016393443 |
| FDR q-value | 0.035893057 |
| FWER p-Value | 0.993 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VIM | VIM Entrez, Source | vimentin | 6 | 0.953 | 0.0845 | Yes |
| 2 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 8 | 0.860 | 0.1611 | Yes |
| 3 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 44 | 0.606 | 0.2125 | Yes |
| 4 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 50 | 0.578 | 0.2636 | Yes |
| 5 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 55 | 0.564 | 0.3136 | Yes |
| 6 | ANKRD1 | ANKRD1 Entrez, Source | ankyrin repeat domain 1 (cardiac muscle) | 63 | 0.532 | 0.3604 | Yes |
| 7 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 130 | 0.390 | 0.3902 | Yes |
| 8 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 140 | 0.372 | 0.4227 | Yes |
| 9 | ANXA8 | ANXA8 Entrez, Source | annexin A8 | 177 | 0.340 | 0.4503 | Yes |
| 10 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 401 | 0.209 | 0.4521 | Yes |
| 11 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.4594 | Yes |
| 12 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 573 | 0.169 | 0.4703 | Yes |
| 13 | PXN | PXN Entrez, Source | paxillin | 617 | 0.163 | 0.4815 | Yes |
| 14 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 849 | 0.134 | 0.4760 | Yes |
| 15 | CD69 | CD69 Entrez, Source | CD69 molecule | 898 | 0.129 | 0.4839 | Yes |
| 16 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1192 | 0.110 | 0.4716 | No |
| 17 | MMP14 | MMP14 Entrez, Source | matrix metallopeptidase 14 (membrane-inserted) | 1312 | 0.104 | 0.4718 | No |
| 18 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 1475 | 0.095 | 0.4681 | No |
| 19 | NUCB2 | NUCB2 Entrez, Source | nucleobindin 2 | 1503 | 0.094 | 0.4744 | No |
| 20 | PDE6H | PDE6H Entrez, Source | phosphodiesterase 6H, cGMP-specific, cone, gamma | 1673 | 0.086 | 0.4693 | No |
| 21 | SCN8A | SCN8A Entrez, Source | sodium channel, voltage gated, type VIII, alpha | 1920 | 0.076 | 0.4574 | No |
| 22 | PACS1 | PACS1 Entrez, Source | phosphofurin acidic cluster sorting protein 1 | 2036 | 0.072 | 0.4551 | No |
| 23 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 2045 | 0.071 | 0.4609 | No |
| 24 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 2181 | 0.068 | 0.4567 | No |
| 25 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 2200 | 0.067 | 0.4614 | No |
| 26 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 2354 | 0.063 | 0.4554 | No |
| 27 | DIABLO | DIABLO Entrez, Source | diablo homolog (Drosophila) | 2448 | 0.061 | 0.4538 | No |
| 28 | GRASP | GRASP Entrez, Source | GRP1 (general receptor for phosphoinositides 1)-associated scaffold protein | 2540 | 0.059 | 0.4522 | No |
| 29 | MYOD1 | MYOD1 Entrez, Source | myogenic differentiation 1 | 2669 | 0.056 | 0.4475 | No |
| 30 | DAB1 | DAB1 Entrez, Source | disabled homolog 1 (Drosophila) | 2694 | 0.055 | 0.4506 | No |
| 31 | RIN1 | RIN1 Entrez, Source | Ras and Rab interactor 1 | 2778 | 0.053 | 0.4491 | No |
| 32 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 2819 | 0.052 | 0.4507 | No |
| 33 | GPATC2 | GPATC2 Entrez, Source | G patch domain containing 2 | 2920 | 0.050 | 0.4476 | No |
| 34 | IFNG | IFNG Entrez, Source | interferon, gamma | 2940 | 0.050 | 0.4507 | No |
| 35 | NGB | NGB Entrez, Source | neuroglobin | 3301 | 0.043 | 0.4272 | No |
| 36 | SNX1 | SNX1 Entrez, Source | sorting nexin 1 | 3339 | 0.042 | 0.4282 | No |
| 37 | MAMDC1 | MAMDC1 Entrez, Source | MAM domain containing 1 | 3395 | 0.041 | 0.4276 | No |
| 38 | SLC6A9 | SLC6A9 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, glycine), member 9 | 3495 | 0.039 | 0.4236 | No |
| 39 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 3509 | 0.038 | 0.4260 | No |
| 40 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 3736 | 0.035 | 0.4120 | No |
| 41 | PTK9 | PTK9 Entrez, Source | PTK9 protein tyrosine kinase 9 | 3860 | 0.032 | 0.4056 | No |
| 42 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 3925 | 0.031 | 0.4036 | No |
| 43 | SCN2B | SCN2B Entrez, Source | sodium channel, voltage-gated, type II, beta | 4134 | 0.028 | 0.3904 | No |
| 44 | TNNT2 | TNNT2 Entrez, Source | troponin T type 2 (cardiac) | 4182 | 0.027 | 0.3893 | No |
| 45 | PSMA1 | PSMA1 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 1 | 4225 | 0.027 | 0.3885 | No |
| 46 | CHRNA10 | CHRNA10 Entrez, Source | cholinergic receptor, nicotinic, alpha 10 | 4232 | 0.026 | 0.3904 | No |
| 47 | PRICKLE1 | PRICKLE1 Entrez, Source | prickle homolog 1 (Drosophila) | 4383 | 0.024 | 0.3812 | No |
| 48 | BLOC1S2 | BLOC1S2 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 2 | 4447 | 0.023 | 0.3785 | No |
| 49 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 4727 | 0.019 | 0.3591 | No |
| 50 | DNM3 | DNM3 Entrez, Source | dynamin 3 | 4784 | 0.018 | 0.3565 | No |
| 51 | SCN3B | SCN3B Entrez, Source | sodium channel, voltage-gated, type III, beta | 5097 | 0.014 | 0.3342 | No |
| 52 | IL23A | IL23A Entrez, Source | interleukin 23, alpha subunit p19 | 5247 | 0.012 | 0.3240 | No |
| 53 | SCNN1A | SCNN1A Entrez, Source | sodium channel, nonvoltage-gated 1 alpha | 5333 | 0.011 | 0.3185 | No |
| 54 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 5416 | 0.010 | 0.3131 | No |
| 55 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 5926 | 0.003 | 0.2749 | No |
| 56 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 6247 | -0.001 | 0.2509 | No |
| 57 | SLC37A4 | SLC37A4 Entrez, Source | solute carrier family 37 (glycerol-6-phosphate transporter), member 4 | 6249 | -0.001 | 0.2509 | No |
| 58 | CRTAC1 | CRTAC1 Entrez, Source | cartilage acidic protein 1 | 6321 | -0.002 | 0.2457 | No |
| 59 | CD6 | CD6 Entrez, Source | CD6 molecule | 6361 | -0.003 | 0.2430 | No |
| 60 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 6475 | -0.004 | 0.2349 | No |
| 61 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 6529 | -0.005 | 0.2313 | No |
| 62 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 6534 | -0.005 | 0.2314 | No |
| 63 | TNFSF4 | TNFSF4 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 4 (tax-transcriptionally activated glycoprotein 1, 34kDa) | 6705 | -0.007 | 0.2192 | No |
| 64 | CUGBP1 | CUGBP1 Entrez, Source | CUG triplet repeat, RNA binding protein 1 | 6720 | -0.007 | 0.2188 | No |
| 65 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 6850 | -0.009 | 0.2098 | No |
| 66 | BTG4 | BTG4 Entrez, Source | B-cell translocation gene 4 | 6902 | -0.009 | 0.2068 | No |
| 67 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 7071 | -0.012 | 0.1951 | No |
| 68 | SLC12A6 | SLC12A6 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 6 | 7142 | -0.012 | 0.1909 | No |
| 69 | KCNJ1 | KCNJ1 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 1 | 7444 | -0.016 | 0.1696 | No |
| 70 | TBX5 | TBX5 Entrez, Source | T-box 5 | 7774 | -0.020 | 0.1466 | No |
| 71 | SEMA4A | SEMA4A Entrez, Source | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A | 7923 | -0.022 | 0.1374 | No |
| 72 | SLC15A3 | SLC15A3 Entrez, Source | solute carrier family 15, member 3 | 8234 | -0.027 | 0.1163 | No |
| 73 | TACC2 | TACC2 Entrez, Source | transforming, acidic coiled-coil containing protein 2 | 8375 | -0.029 | 0.1083 | No |
| 74 | DNAJC14 | DNAJC14 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 14 | 8436 | -0.030 | 0.1064 | No |
| 75 | INPPL1 | INPPL1 Entrez, Source | inositol polyphosphate phosphatase-like 1 | 8457 | -0.030 | 0.1076 | No |
| 76 | MATN1 | MATN1 Entrez, Source | matrilin 1, cartilage matrix protein | 8575 | -0.032 | 0.1016 | No |
| 77 | ARF3 | ARF3 Entrez, Source | ADP-ribosylation factor 3 | 8812 | -0.035 | 0.0869 | No |
| 78 | XCL1 | XCL1 Entrez, Source | chemokine (C motif) ligand 1 | 8819 | -0.035 | 0.0896 | No |
| 79 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 8836 | -0.035 | 0.0915 | No |
| 80 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 8862 | -0.036 | 0.0928 | No |
| 81 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 9316 | -0.042 | 0.0623 | No |
| 82 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 9350 | -0.043 | 0.0636 | No |
| 83 | AGTRL1 | AGTRL1 Entrez, Source | angiotensin II receptor-like 1 | 9375 | -0.043 | 0.0656 | No |
| 84 | FGF4 | FGF4 Entrez, Source | fibroblast growth factor 4 (heparin secretory transforming protein 1, Kaposi sarcoma oncogene) | 9420 | -0.044 | 0.0662 | No |
| 85 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 9444 | -0.044 | 0.0684 | No |
| 86 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 9611 | -0.047 | 0.0601 | No |
| 87 | AKAP3 | AKAP3 Entrez, Source | A kinase (PRKA) anchor protein 3 | 9897 | -0.052 | 0.0431 | No |
| 88 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 9899 | -0.052 | 0.0477 | No |
| 89 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 9915 | -0.052 | 0.0512 | No |
| 90 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 9943 | -0.053 | 0.0538 | No |
| 91 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 9985 | -0.053 | 0.0555 | No |
| 92 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 10100 | -0.055 | 0.0518 | No |
| 93 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 10129 | -0.056 | 0.0546 | No |
| 94 | PTPN7 | PTPN7 Entrez, Source | protein tyrosine phosphatase, non-receptor type 7 | 10142 | -0.056 | 0.0587 | No |
| 95 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 10190 | -0.057 | 0.0602 | No |
| 96 | ARNT | ARNT Entrez, Source | aryl hydrocarbon receptor nuclear translocator | 10332 | -0.059 | 0.0548 | No |
| 97 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 10706 | -0.066 | 0.0325 | No |
| 98 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 10727 | -0.067 | 0.0369 | No |
| 99 | STAT2 | STAT2 Entrez, Source | signal transducer and activator of transcription 2, 113kDa | 10946 | -0.072 | 0.0269 | No |
| 100 | ENTPD1 | ENTPD1 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 1 | 10967 | -0.073 | 0.0318 | No |
| 101 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 11375 | -0.083 | 0.0085 | No |
| 102 | FGD4 | FGD4 Entrez, Source | FYVE, RhoGEF and PH domain containing 4 | 11381 | -0.084 | 0.0156 | No |
| 103 | ABCD4 | ABCD4 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 4 | 11616 | -0.089 | 0.0059 | No |
| 104 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 11816 | -0.096 | -0.0006 | No |
| 105 | GTF2A1 | GTF2A1 Entrez, Source | general transcription factor IIA, 1, 19/37kDa | 12080 | -0.106 | -0.0110 | No |
| 106 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 12383 | -0.120 | -0.0231 | No |
| 107 | BMF | BMF Entrez, Source | Bcl2 modifying factor | 12660 | -0.138 | -0.0317 | No |
| 108 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 12682 | -0.140 | -0.0208 | No |
| 109 | PDZK1 | PDZK1 Entrez, Source | PDZ domain containing 1 | 12895 | -0.164 | -0.0222 | No |
| 110 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 12979 | -0.178 | -0.0126 | No |
| 111 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 13133 | -0.220 | -0.0046 | No |
| 112 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 13158 | -0.228 | 0.0139 | No |

