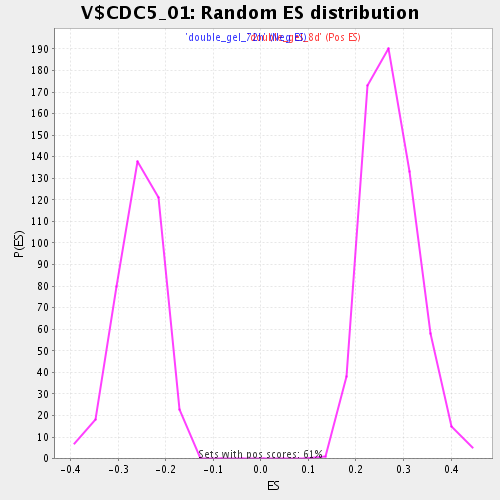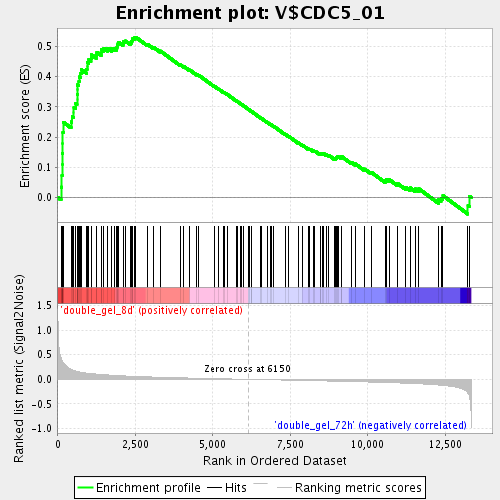
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$CDC5_01 |
| Enrichment Score (ES) | 0.5306131 |
| Normalized Enrichment Score (NES) | 1.9539474 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.008664095 |
| FWER p-Value | 0.362 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 103 | 0.426 | 0.0350 | Yes |
| 2 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 130 | 0.390 | 0.0722 | Yes |
| 3 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 140 | 0.372 | 0.1089 | Yes |
| 4 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 145 | 0.367 | 0.1454 | Yes |
| 5 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 157 | 0.357 | 0.1804 | Yes |
| 6 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 160 | 0.355 | 0.2159 | Yes |
| 7 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 174 | 0.343 | 0.2494 | Yes |
| 8 | TMSB4X | TMSB4X Entrez, Source | thymosin, beta 4, X-linked | 427 | 0.202 | 0.2506 | Yes |
| 9 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 454 | 0.195 | 0.2681 | Yes |
| 10 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 516 | 0.180 | 0.2816 | Yes |
| 11 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.2996 | Yes |
| 12 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 573 | 0.169 | 0.3124 | Yes |
| 13 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 618 | 0.162 | 0.3254 | Yes |
| 14 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 619 | 0.162 | 0.3417 | Yes |
| 15 | BIRC4 | BIRC4 Entrez, Source | baculoviral IAP repeat-containing 4 | 630 | 0.161 | 0.3570 | Yes |
| 16 | FGF10 | FGF10 Entrez, Source | fibroblast growth factor 10 | 632 | 0.160 | 0.3730 | Yes |
| 17 | NDRG4 | NDRG4 Entrez, Source | NDRG family member 4 | 677 | 0.153 | 0.3850 | Yes |
| 18 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 689 | 0.151 | 0.3994 | Yes |
| 19 | PROK2 | PROK2 Entrez, Source | prokineticin 2 | 732 | 0.146 | 0.4109 | Yes |
| 20 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 756 | 0.144 | 0.4236 | Yes |
| 21 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 920 | 0.127 | 0.4240 | Yes |
| 22 | ROD1 | ROD1 Entrez, Source | ROD1 regulator of differentiation 1 (S. pombe) | 942 | 0.126 | 0.4351 | Yes |
| 23 | PBX2 | PBX2 Entrez, Source | pre-B-cell leukemia transcription factor 2 | 944 | 0.126 | 0.4476 | Yes |
| 24 | GFRA1 | GFRA1 Entrez, Source | GDNF family receptor alpha 1 | 997 | 0.122 | 0.4559 | Yes |
| 25 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1068 | 0.116 | 0.4622 | Yes |
| 26 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 1086 | 0.116 | 0.4726 | Yes |
| 27 | MOS | MOS Entrez, Source | v-mos Moloney murine sarcoma viral oncogene homolog | 1237 | 0.108 | 0.4721 | Yes |
| 28 | CHRNA5 | CHRNA5 Entrez, Source | cholinergic receptor, nicotinic, alpha 5 | 1256 | 0.107 | 0.4814 | Yes |
| 29 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 1398 | 0.099 | 0.4807 | Yes |
| 30 | CDH6 | CDH6 Entrez, Source | cadherin 6, type 2, K-cadherin (fetal kidney) | 1405 | 0.098 | 0.4901 | Yes |
| 31 | SLCO2A1 | SLCO2A1 Entrez, Source | solute carrier organic anion transporter family, member 2A1 | 1480 | 0.095 | 0.4940 | Yes |
| 32 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 1606 | 0.089 | 0.4935 | Yes |
| 33 | STAT3 | STAT3 Entrez, Source | signal transducer and activator of transcription 3 (acute-phase response factor) | 1729 | 0.083 | 0.4926 | Yes |
| 34 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 1820 | 0.080 | 0.4938 | Yes |
| 35 | OSR2 | OSR2 Entrez, Source | odd-skipped related 2 (Drosophila) | 1901 | 0.077 | 0.4954 | Yes |
| 36 | PRKAG1 | PRKAG1 Entrez, Source | protein kinase, AMP-activated, gamma 1 non-catalytic subunit | 1908 | 0.076 | 0.5027 | Yes |
| 37 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 1934 | 0.075 | 0.5083 | Yes |
| 38 | FGF12 | FGF12 Entrez, Source | fibroblast growth factor 12 | 1947 | 0.075 | 0.5149 | Yes |
| 39 | SAMSN1 | SAMSN1 Entrez, Source | SAM domain, SH3 domain and nuclear localisation signals, 1 | 2103 | 0.070 | 0.5102 | Yes |
| 40 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 2118 | 0.069 | 0.5161 | Yes |
| 41 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 2178 | 0.068 | 0.5184 | Yes |
| 42 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 2354 | 0.063 | 0.5115 | Yes |
| 43 | DGKB | DGKB Entrez, Source | diacylglycerol kinase, beta 90kDa | 2364 | 0.063 | 0.5172 | Yes |
| 44 | YWHAG | YWHAG Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide | 2394 | 0.062 | 0.5212 | Yes |
| 45 | CHRDL1 | CHRDL1 Entrez, Source | chordin-like 1 | 2416 | 0.062 | 0.5258 | Yes |
| 46 | SLC27A4 | SLC27A4 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 4 | 2459 | 0.061 | 0.5287 | Yes |
| 47 | GPC3 | GPC3 Entrez, Source | glypican 3 | 2514 | 0.059 | 0.5306 | Yes |
| 48 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 2881 | 0.051 | 0.5081 | No |
| 49 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 3095 | 0.047 | 0.4967 | No |
| 50 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 3304 | 0.042 | 0.4852 | No |
| 51 | ODF1 | ODF1 Entrez, Source | outer dense fiber of sperm tails 1 | 3951 | 0.031 | 0.4395 | No |
| 52 | GLI3 | GLI3 Entrez, Source | GLI-Kruppel family member GLI3 (Greig cephalopolysyndactyly syndrome) | 4066 | 0.029 | 0.4338 | No |
| 53 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 4251 | 0.026 | 0.4225 | No |
| 54 | CSNK1G3 | CSNK1G3 Entrez, Source | casein kinase 1, gamma 3 | 4483 | 0.023 | 0.4073 | No |
| 55 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 4536 | 0.022 | 0.4056 | No |
| 56 | COL16A1 | COL16A1 Entrez, Source | collagen, type XVI, alpha 1 | 5051 | 0.015 | 0.3682 | No |
| 57 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 5189 | 0.013 | 0.3591 | No |
| 58 | SLC6A13 | SLC6A13 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, GABA), member 13 | 5341 | 0.011 | 0.3488 | No |
| 59 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 5378 | 0.010 | 0.3471 | No |
| 60 | INPP4B | INPP4B Entrez, Source | inositol polyphosphate-4-phosphatase, type II, 105kDa | 5469 | 0.009 | 0.3411 | No |
| 61 | SHH | SHH Entrez, Source | sonic hedgehog homolog (Drosophila) | 5779 | 0.005 | 0.3183 | No |
| 62 | LMNA | LMNA Entrez, Source | lamin A/C | 5805 | 0.004 | 0.3168 | No |
| 63 | PLCB3 | PLCB3 Entrez, Source | phospholipase C, beta 3 (phosphatidylinositol-specific) | 5900 | 0.003 | 0.3100 | No |
| 64 | TNNI3K | TNNI3K Entrez, Source | TNNI3 interacting kinase | 5925 | 0.003 | 0.3085 | No |
| 65 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 5987 | 0.002 | 0.3042 | No |
| 66 | UNC13D | UNC13D Entrez, Source | unc-13 homolog D (C. elegans) | 6157 | -0.000 | 0.2914 | No |
| 67 | CART1 | CART1 Entrez, Source | cartilage paired-class homeoprotein 1 | 6181 | -0.000 | 0.2897 | No |
| 68 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 6239 | -0.001 | 0.2855 | No |
| 69 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 6247 | -0.001 | 0.2851 | No |
| 70 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 6534 | -0.005 | 0.2640 | No |
| 71 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 6574 | -0.005 | 0.2616 | No |
| 72 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 6769 | -0.008 | 0.2477 | No |
| 73 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 6846 | -0.009 | 0.2428 | No |
| 74 | GPR85 | GPR85 Entrez, Source | G protein-coupled receptor 85 | 6883 | -0.009 | 0.2410 | No |
| 75 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 6971 | -0.010 | 0.2355 | No |
| 76 | CLASP1 | CLASP1 Entrez, Source | cytoplasmic linker associated protein 1 | 7339 | -0.015 | 0.2092 | No |
| 77 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 7452 | -0.016 | 0.2024 | No |
| 78 | GJA3 | GJA3 Entrez, Source | gap junction protein, alpha 3, 46kDa (connexin 46) | 7759 | -0.020 | 0.1813 | No |
| 79 | FUT11 | FUT11 Entrez, Source | fucosyltransferase 11 (alpha (1,3) fucosyltransferase) | 7899 | -0.022 | 0.1730 | No |
| 80 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 8090 | -0.025 | 0.1611 | No |
| 81 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 8131 | -0.026 | 0.1606 | No |
| 82 | NEK6 | NEK6 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 6 | 8247 | -0.027 | 0.1547 | No |
| 83 | SYT4 | SYT4 Entrez, Source | synaptotagmin IV | 8295 | -0.028 | 0.1539 | No |
| 84 | VDR | VDR Entrez, Source | vitamin D (1,25- dihydroxyvitamin D3) receptor | 8460 | -0.030 | 0.1446 | No |
| 85 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 8470 | -0.031 | 0.1469 | No |
| 86 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 8540 | -0.031 | 0.1449 | No |
| 87 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 8579 | -0.032 | 0.1452 | No |
| 88 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 8663 | -0.033 | 0.1422 | No |
| 89 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 8732 | -0.034 | 0.1405 | No |
| 90 | KCNK12 | KCNK12 Entrez, Source | potassium channel, subfamily K, member 12 | 8930 | -0.037 | 0.1293 | No |
| 91 | SLC12A8 | SLC12A8 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 8 | 8971 | -0.037 | 0.1300 | No |
| 92 | SFRS12 | SFRS12 Entrez, Source | splicing factor, arginine/serine-rich 12 | 8997 | -0.038 | 0.1319 | No |
| 93 | OTP | OTP Entrez, Source | orthopedia homolog (Drosophila) | 9009 | -0.038 | 0.1348 | No |
| 94 | HSPA2 | HSPA2 Entrez, Source | heat shock 70kDa protein 2 | 9062 | -0.038 | 0.1348 | No |
| 95 | ANGPT1 | ANGPT1 Entrez, Source | angiopoietin 1 | 9136 | -0.039 | 0.1332 | No |
| 96 | TNNI1 | TNNI1 Entrez, Source | troponin I type 1 (skeletal, slow) | 9147 | -0.039 | 0.1364 | No |
| 97 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 9466 | -0.044 | 0.1168 | No |
| 98 | ACTR10 | ACTR10 Entrez, Source | actin-related protein 10 homolog (S. cerevisiae) | 9598 | -0.047 | 0.1116 | No |
| 99 | ARPC5L | ARPC5L Entrez, Source | actin related protein 2/3 complex, subunit 5-like | 9893 | -0.052 | 0.0946 | No |
| 100 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 10110 | -0.055 | 0.0838 | No |
| 101 | NDNL2 | NDNL2 Entrez, Source | necdin-like 2 | 10563 | -0.063 | 0.0560 | No |
| 102 | MYH6 | MYH6 Entrez, Source | myosin, heavy chain 6, cardiac muscle, alpha (cardiomyopathy, hypertrophic 1) | 10594 | -0.064 | 0.0602 | No |
| 103 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 10686 | -0.066 | 0.0599 | No |
| 104 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 10957 | -0.072 | 0.0468 | No |
| 105 | FNBP1 | FNBP1 Entrez, Source | formin binding protein 1 | 11225 | -0.079 | 0.0345 | No |
| 106 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 11368 | -0.083 | 0.0322 | No |
| 107 | MYL2 | MYL2 Entrez, Source | myosin, light chain 2, regulatory, cardiac, slow | 11531 | -0.087 | 0.0287 | No |
| 108 | SCN7A | SCN7A Entrez, Source | sodium channel, voltage-gated, type VII, alpha | 11650 | -0.090 | 0.0288 | No |
| 109 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 12283 | -0.115 | -0.0074 | No |
| 110 | LIN7A | LIN7A Entrez, Source | lin-7 homolog A (C. elegans) | 12366 | -0.119 | -0.0016 | No |
| 111 | MUSK | MUSK Entrez, Source | muscle, skeletal, receptor tyrosine kinase | 12426 | -0.123 | 0.0062 | No |
| 112 | ENPP2 | ENPP2 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 2 (autotaxin) | 13234 | -0.281 | -0.0266 | No |
| 113 | ARNTL | ARNTL Entrez, Source | aryl hydrocarbon receptor nuclear translocator-like | 13280 | -0.345 | 0.0047 | No |