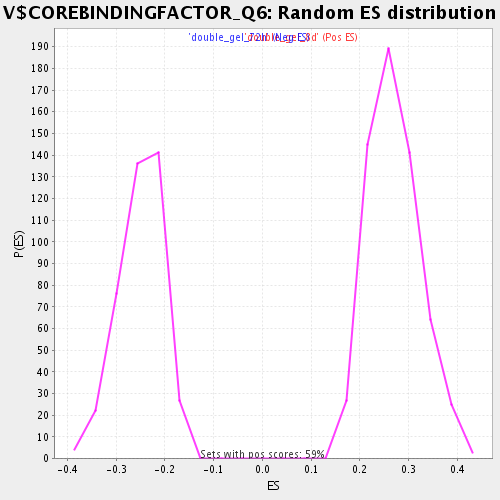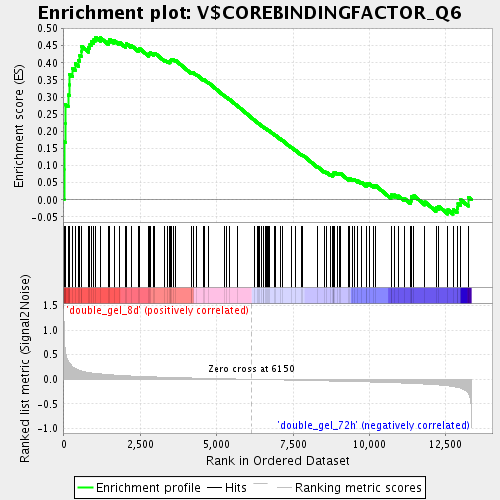
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$COREBINDINGFACTOR_Q6 |
| Enrichment Score (ES) | 0.47480443 |
| Normalized Enrichment Score (NES) | 1.7588161 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.038995 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VIM | VIM Entrez, Source | vimentin | 6 | 0.953 | 0.0891 | Yes |
| 2 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 8 | 0.860 | 0.1699 | Yes |
| 3 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 44 | 0.606 | 0.2242 | Yes |
| 4 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 50 | 0.578 | 0.2782 | Yes |
| 5 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 140 | 0.372 | 0.3064 | Yes |
| 6 | ANXA8 | ANXA8 Entrez, Source | annexin A8 | 177 | 0.340 | 0.3357 | Yes |
| 7 | CCL3 | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 183 | 0.333 | 0.3666 | Yes |
| 8 | PTK2B | PTK2B Entrez, Source | PTK2B protein tyrosine kinase 2 beta | 279 | 0.262 | 0.3841 | Yes |
| 9 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 372 | 0.219 | 0.3977 | Yes |
| 10 | AKAP2 | AKAP2 Entrez, Source | A kinase (PRKA) anchor protein 2 | 481 | 0.186 | 0.4070 | Yes |
| 11 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.4212 | Yes |
| 12 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 568 | 0.170 | 0.4334 | Yes |
| 13 | IL17RE | IL17RE Entrez, Source | interleukin 17 receptor E | 574 | 0.168 | 0.4489 | Yes |
| 14 | PPP1R14C | PPP1R14C Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 14C | 807 | 0.139 | 0.4444 | Yes |
| 15 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 849 | 0.134 | 0.4539 | Yes |
| 16 | CD69 | CD69 Entrez, Source | CD69 molecule | 898 | 0.129 | 0.4625 | Yes |
| 17 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 979 | 0.123 | 0.4680 | Yes |
| 18 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 1037 | 0.118 | 0.4748 | Yes |
| 19 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1192 | 0.110 | 0.4735 | No |
| 20 | SULF1 | SULF1 Entrez, Source | sulfatase 1 | 1470 | 0.095 | 0.4615 | No |
| 21 | LEMD2 | LEMD2 Entrez, Source | LEM domain containing 2 | 1488 | 0.095 | 0.4691 | No |
| 22 | RAMP2 | RAMP2 Entrez, Source | receptor (calcitonin) activity modifying protein 2 | 1641 | 0.087 | 0.4659 | No |
| 23 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 1820 | 0.080 | 0.4599 | No |
| 24 | ADD1 | ADD1 Entrez, Source | adducin 1 (alpha) | 2026 | 0.072 | 0.4512 | No |
| 25 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 2045 | 0.071 | 0.4565 | No |
| 26 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 2200 | 0.067 | 0.4512 | No |
| 27 | DIABLO | DIABLO Entrez, Source | diablo homolog (Drosophila) | 2448 | 0.061 | 0.4382 | No |
| 28 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 2479 | 0.060 | 0.4416 | No |
| 29 | RIN1 | RIN1 Entrez, Source | Ras and Rab interactor 1 | 2778 | 0.053 | 0.4241 | No |
| 30 | ENTPD3 | ENTPD3 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 3 | 2795 | 0.053 | 0.4279 | No |
| 31 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 2819 | 0.052 | 0.4311 | No |
| 32 | NFIX | NFIX Entrez, Source | nuclear factor I/X (CCAAT-binding transcription factor) | 2946 | 0.050 | 0.4262 | No |
| 33 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 2974 | 0.049 | 0.4288 | No |
| 34 | NGB | NGB Entrez, Source | neuroglobin | 3301 | 0.043 | 0.4082 | No |
| 35 | MAMDC1 | MAMDC1 Entrez, Source | MAM domain containing 1 | 3395 | 0.041 | 0.4049 | No |
| 36 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 3468 | 0.039 | 0.4032 | No |
| 37 | ELAVL3 | ELAVL3 Entrez, Source | ELAV (embryonic lethal, abnormal vision, Drosophila)-like 3 (Hu antigen C) | 3483 | 0.039 | 0.4058 | No |
| 38 | IL13 | IL13 Entrez, Source | interleukin 13 | 3501 | 0.039 | 0.4081 | No |
| 39 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 3509 | 0.038 | 0.4112 | No |
| 40 | PDGFC | PDGFC Entrez, Source | platelet derived growth factor C | 3584 | 0.037 | 0.4091 | No |
| 41 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 3662 | 0.036 | 0.4067 | No |
| 42 | CYGB | CYGB Entrez, Source | cytoglobin | 4165 | 0.028 | 0.3713 | No |
| 43 | TGM4 | TGM4 Entrez, Source | transglutaminase 4 (prostate) | 4185 | 0.027 | 0.3724 | No |
| 44 | PSMA1 | PSMA1 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 1 | 4225 | 0.027 | 0.3720 | No |
| 45 | HPCAL1 | HPCAL1 Entrez, Source | hippocalcin-like 1 | 4353 | 0.025 | 0.3647 | No |
| 46 | TRPM8 | TRPM8 Entrez, Source | transient receptor potential cation channel, subfamily M, member 8 | 4554 | 0.022 | 0.3516 | No |
| 47 | LUC7L | LUC7L Entrez, Source | LUC7-like (S. cerevisiae) | 4592 | 0.021 | 0.3508 | No |
| 48 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 4727 | 0.019 | 0.3425 | No |
| 49 | IL23A | IL23A Entrez, Source | interleukin 23, alpha subunit p19 | 5247 | 0.012 | 0.3044 | No |
| 50 | SCNN1A | SCNN1A Entrez, Source | sodium channel, nonvoltage-gated 1 alpha | 5333 | 0.011 | 0.2990 | No |
| 51 | MYL3 | MYL3 Entrez, Source | myosin, light chain 3, alkali; ventricular, skeletal, slow | 5408 | 0.010 | 0.2943 | No |
| 52 | NRXN1 | NRXN1 Entrez, Source | neurexin 1 | 5683 | 0.006 | 0.2741 | No |
| 53 | SLC37A4 | SLC37A4 Entrez, Source | solute carrier family 37 (glycerol-6-phosphate transporter), member 4 | 6249 | -0.001 | 0.2315 | No |
| 54 | CRTAC1 | CRTAC1 Entrez, Source | cartilage acidic protein 1 | 6321 | -0.002 | 0.2264 | No |
| 55 | CD6 | CD6 Entrez, Source | CD6 molecule | 6361 | -0.003 | 0.2237 | No |
| 56 | C1QTNF6 | C1QTNF6 Entrez, Source | C1q and tumor necrosis factor related protein 6 | 6402 | -0.003 | 0.2209 | No |
| 57 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 6475 | -0.004 | 0.2159 | No |
| 58 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 6529 | -0.005 | 0.2123 | No |
| 59 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 6534 | -0.005 | 0.2125 | No |
| 60 | IL3 | IL3 Entrez, Source | interleukin 3 (colony-stimulating factor, multiple) | 6597 | -0.006 | 0.2083 | No |
| 61 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 6625 | -0.006 | 0.2069 | No |
| 62 | FGD1 | FGD1 Entrez, Source | FYVE, RhoGEF and PH domain containing 1 (faciogenital dysplasia) | 6676 | -0.007 | 0.2037 | No |
| 63 | TNFSF4 | TNFSF4 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 4 (tax-transcriptionally activated glycoprotein 1, 34kDa) | 6705 | -0.007 | 0.2023 | No |
| 64 | MAFG | MAFG Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog G (avian) | 6733 | -0.007 | 0.2009 | No |
| 65 | BTG4 | BTG4 Entrez, Source | B-cell translocation gene 4 | 6902 | -0.009 | 0.1891 | No |
| 66 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 6929 | -0.010 | 0.1881 | No |
| 67 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 7071 | -0.012 | 0.1785 | No |
| 68 | SLC12A6 | SLC12A6 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 6 | 7142 | -0.012 | 0.1744 | No |
| 69 | KCNJ1 | KCNJ1 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 1 | 7444 | -0.016 | 0.1531 | No |
| 70 | GRAP2 | GRAP2 Entrez, Source | GRB2-related adaptor protein 2 | 7576 | -0.018 | 0.1449 | No |
| 71 | TBX5 | TBX5 Entrez, Source | T-box 5 | 7774 | -0.020 | 0.1319 | No |
| 72 | FKBP14 | FKBP14 Entrez, Source | FK506 binding protein 14, 22 kDa | 7815 | -0.021 | 0.1309 | No |
| 73 | HEY1 | HEY1 Entrez, Source | hairy/enhancer-of-split related with YRPW motif 1 | 8303 | -0.028 | 0.0967 | No |
| 74 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 8513 | -0.031 | 0.0838 | No |
| 75 | NUTF2 | NUTF2 Entrez, Source | nuclear transport factor 2 | 8594 | -0.032 | 0.0808 | No |
| 76 | CALR | CALR Entrez, Source | calreticulin | 8727 | -0.034 | 0.0740 | No |
| 77 | GABRA6 | GABRA6 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 6 | 8802 | -0.035 | 0.0717 | No |
| 78 | ARF3 | ARF3 Entrez, Source | ADP-ribosylation factor 3 | 8812 | -0.035 | 0.0743 | No |
| 79 | XCL1 | XCL1 Entrez, Source | chemokine (C motif) ligand 1 | 8819 | -0.035 | 0.0771 | No |
| 80 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 8836 | -0.035 | 0.0792 | No |
| 81 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 8862 | -0.036 | 0.0807 | No |
| 82 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 8947 | -0.037 | 0.0778 | No |
| 83 | OTP | OTP Entrez, Source | orthopedia homolog (Drosophila) | 9009 | -0.038 | 0.0767 | No |
| 84 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 9039 | -0.038 | 0.0781 | No |
| 85 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 9316 | -0.042 | 0.0612 | No |
| 86 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 9350 | -0.043 | 0.0627 | No |
| 87 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 9444 | -0.044 | 0.0598 | No |
| 88 | NAPSA | NAPSA Entrez, Source | napsin A aspartic peptidase | 9500 | -0.045 | 0.0599 | No |
| 89 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 9611 | -0.047 | 0.0560 | No |
| 90 | CACNA2D3 | CACNA2D3 Entrez, Source | calcium channel, voltage-dependent, alpha 2/delta 3 subunit | 9729 | -0.049 | 0.0518 | No |
| 91 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 9899 | -0.052 | 0.0439 | No |
| 92 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 9915 | -0.052 | 0.0477 | No |
| 93 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 9985 | -0.053 | 0.0475 | No |
| 94 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 10129 | -0.056 | 0.0419 | No |
| 95 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 10190 | -0.057 | 0.0427 | No |
| 96 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 10706 | -0.066 | 0.0100 | No |
| 97 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 10727 | -0.067 | 0.0147 | No |
| 98 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 10816 | -0.069 | 0.0145 | No |
| 99 | STAT2 | STAT2 Entrez, Source | signal transducer and activator of transcription 2, 113kDa | 10946 | -0.072 | 0.0115 | No |
| 100 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 11134 | -0.077 | 0.0046 | No |
| 101 | SHOX2 | SHOX2 Entrez, Source | short stature homeobox 2 | 11349 | -0.083 | -0.0038 | No |
| 102 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 11362 | -0.083 | 0.0031 | No |
| 103 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 11375 | -0.083 | 0.0100 | No |
| 104 | CHAD | CHAD Entrez, Source | chondroadherin | 11454 | -0.085 | 0.0121 | No |
| 105 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 11816 | -0.096 | -0.0061 | No |
| 106 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 12179 | -0.110 | -0.0231 | No |
| 107 | CD28 | CD28 Entrez, Source | CD28 molecule | 12261 | -0.114 | -0.0185 | No |
| 108 | GPX1 | GPX1 Entrez, Source | glutathione peroxidase 1 | 12567 | -0.133 | -0.0291 | No |
| 109 | DRD3 | DRD3 Entrez, Source | dopamine receptor D3 | 12734 | -0.145 | -0.0280 | No |
| 110 | SOST | SOST Entrez, Source | sclerosteosis | 12868 | -0.161 | -0.0229 | No |
| 111 | PDZK1 | PDZK1 Entrez, Source | PDZ domain containing 1 | 12895 | -0.164 | -0.0095 | No |
| 112 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 12979 | -0.178 | 0.0010 | No |
| 113 | ENPP2 | ENPP2 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 2 (autotaxin) | 13234 | -0.281 | 0.0082 | No |