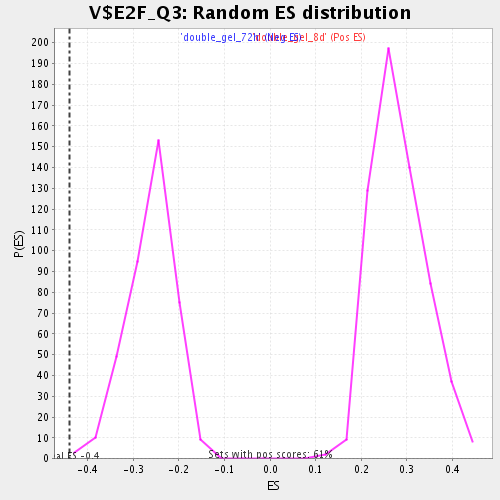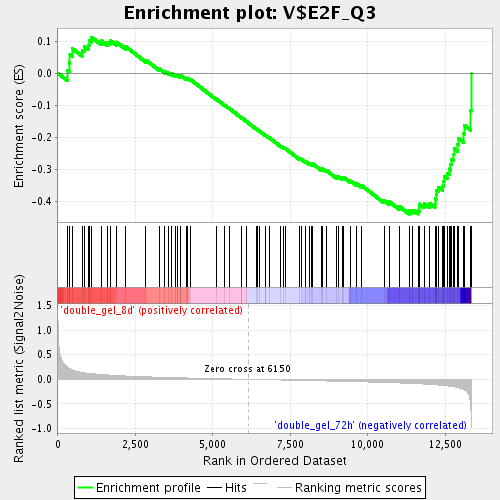
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | V$E2F_Q3 |
| Enrichment Score (ES) | -0.4394078 |
| Normalized Enrichment Score (NES) | -1.6828376 |
| Nominal p-value | 0.002538071 |
| FDR q-value | 0.16707405 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ONECUT1 | ONECUT1 Entrez, Source | one cut domain, family member 1 | 309 | 0.245 | 0.0083 | No |
| 2 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 366 | 0.223 | 0.0328 | No |
| 3 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 389 | 0.213 | 0.0586 | No |
| 4 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 473 | 0.188 | 0.0766 | No |
| 5 | KLF5 | KLF5 Entrez, Source | Kruppel-like factor 5 (intestinal) | 780 | 0.141 | 0.0718 | No |
| 6 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 849 | 0.134 | 0.0840 | No |
| 7 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 1001 | 0.121 | 0.0882 | No |
| 8 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 1021 | 0.119 | 0.1022 | No |
| 9 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 1086 | 0.116 | 0.1123 | No |
| 10 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 1398 | 0.099 | 0.1016 | No |
| 11 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 1606 | 0.089 | 0.0974 | No |
| 12 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1690 | 0.085 | 0.1022 | No |
| 13 | PCSK4 | PCSK4 Entrez, Source | proprotein convertase subtilisin/kexin type 4 | 1884 | 0.077 | 0.0976 | No |
| 14 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 2181 | 0.068 | 0.0840 | No |
| 15 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 2842 | 0.052 | 0.0409 | No |
| 16 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 3272 | 0.043 | 0.0141 | No |
| 17 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 3452 | 0.040 | 0.0057 | No |
| 18 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 3568 | 0.037 | 0.0018 | No |
| 19 | KCNA1 | KCNA1 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 1 (episodic ataxia with myokymia) | 3681 | 0.036 | -0.0020 | No |
| 20 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 3793 | 0.034 | -0.0061 | No |
| 21 | SYT11 | SYT11 Entrez, Source | synaptotagmin XI | 3867 | 0.032 | -0.0074 | No |
| 22 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 3943 | 0.031 | -0.0091 | No |
| 23 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 3967 | 0.031 | -0.0068 | No |
| 24 | BARHL1 | BARHL1 Entrez, Source | BarH-like 1 (Drosophila) | 4137 | 0.028 | -0.0159 | No |
| 25 | SEZ6 | SEZ6 Entrez, Source | seizure related 6 homolog (mouse) | 4170 | 0.028 | -0.0148 | No |
| 26 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 4293 | 0.026 | -0.0207 | No |
| 27 | LHX5 | LHX5 Entrez, Source | LIM homeobox 5 | 5104 | 0.014 | -0.0800 | No |
| 28 | SLC6A4 | SLC6A4 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 5390 | 0.010 | -0.1003 | No |
| 29 | PELP1 | PELP1 Entrez, Source | proline, glutamic acid and leucine rich protein 1 | 5522 | 0.008 | -0.1091 | No |
| 30 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 5938 | 0.003 | -0.1400 | No |
| 31 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 5939 | 0.003 | -0.1397 | No |
| 32 | PVRL1 | PVRL1 Entrez, Source | poliovirus receptor-related 1 (herpesvirus entry mediator C; nectin) | 6070 | 0.001 | -0.1493 | No |
| 33 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 6398 | -0.003 | -0.1736 | No |
| 34 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 6442 | -0.004 | -0.1764 | No |
| 35 | LASP1 | LASP1 Entrez, Source | LIM and SH3 protein 1 | 6500 | -0.004 | -0.1801 | No |
| 36 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 6696 | -0.007 | -0.1939 | No |
| 37 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 6700 | -0.007 | -0.1932 | No |
| 38 | BAT3 | BAT3 Entrez, Source | HLA-B associated transcript 3 | 6821 | -0.008 | -0.2012 | No |
| 39 | KBTBD7 | KBTBD7 Entrez, Source | kelch repeat and BTB (POZ) domain containing 7 | 7191 | -0.013 | -0.2273 | No |
| 40 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 7279 | -0.014 | -0.2321 | No |
| 41 | CTDSP1 | CTDSP1 Entrez, Source | CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 1 | 7334 | -0.015 | -0.2343 | No |
| 42 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 7792 | -0.021 | -0.2661 | No |
| 43 | SLC25A3 | SLC25A3 Entrez, Source | solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 | 7848 | -0.021 | -0.2675 | No |
| 44 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7992 | -0.023 | -0.2753 | No |
| 45 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 8111 | -0.025 | -0.2809 | No |
| 46 | ADCY8 | ADCY8 Entrez, Source | adenylate cyclase 8 (brain) | 8194 | -0.026 | -0.2837 | No |
| 47 | GRIK2 | GRIK2 Entrez, Source | glutamate receptor, ionotropic, kainate 2 | 8215 | -0.027 | -0.2818 | No |
| 48 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 8504 | -0.031 | -0.2995 | No |
| 49 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 8535 | -0.031 | -0.2978 | No |
| 50 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 8663 | -0.033 | -0.3031 | No |
| 51 | SFRS12 | SFRS12 Entrez, Source | splicing factor, arginine/serine-rich 12 | 8997 | -0.038 | -0.3233 | No |
| 52 | UXT | UXT Entrez, Source | ubiquitously-expressed transcript | 9053 | -0.038 | -0.3226 | No |
| 53 | CLTA | CLTA Entrez, Source | clathrin, light chain (Lca) | 9172 | -0.040 | -0.3263 | No |
| 54 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 9219 | -0.040 | -0.3246 | No |
| 55 | NCL | NCL Entrez, Source | nucleolin | 9448 | -0.044 | -0.3361 | No |
| 56 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 9637 | -0.048 | -0.3441 | No |
| 57 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 9789 | -0.050 | -0.3490 | No |
| 58 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 10524 | -0.063 | -0.3963 | No |
| 59 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 10688 | -0.066 | -0.4001 | No |
| 60 | RHD | RHD Entrez, Source | Rh blood group, D antigen | 11014 | -0.074 | -0.4151 | No |
| 61 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 11337 | -0.082 | -0.4288 | Yes |
| 62 | NDUFA11 | NDUFA11 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 11, 14.7kDa | 11457 | -0.085 | -0.4267 | Yes |
| 63 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 11624 | -0.090 | -0.4277 | Yes |
| 64 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 11671 | -0.091 | -0.4195 | Yes |
| 65 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 11674 | -0.091 | -0.4079 | Yes |
| 66 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 11823 | -0.096 | -0.4066 | Yes |
| 67 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 11995 | -0.103 | -0.4063 | Yes |
| 68 | HTF9C | HTF9C Entrez, Source | - | 12171 | -0.110 | -0.4052 | Yes |
| 69 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12174 | -0.110 | -0.3912 | Yes |
| 70 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 12208 | -0.112 | -0.3792 | Yes |
| 71 | CACNA1G | CACNA1G Entrez, Source | calcium channel, voltage-dependent, alpha 1G subunit | 12214 | -0.112 | -0.3652 | Yes |
| 72 | FMO4 | FMO4 Entrez, Source | flavin containing monooxygenase 4 | 12279 | -0.115 | -0.3552 | Yes |
| 73 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 12427 | -0.123 | -0.3504 | Yes |
| 74 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 12451 | -0.124 | -0.3362 | Yes |
| 75 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 12463 | -0.125 | -0.3210 | Yes |
| 76 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12570 | -0.133 | -0.3118 | Yes |
| 77 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 12628 | -0.136 | -0.2986 | Yes |
| 78 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 12677 | -0.140 | -0.2842 | Yes |
| 79 | SSBP3 | SSBP3 Entrez, Source | single stranded DNA binding protein 3 | 12700 | -0.142 | -0.2676 | Yes |
| 80 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 12780 | -0.150 | -0.2542 | Yes |
| 81 | DCK | DCK Entrez, Source | deoxycytidine kinase | 12784 | -0.151 | -0.2350 | Yes |
| 82 | XTP3TPA | XTP3TPA Entrez, Source | - | 12898 | -0.165 | -0.2223 | Yes |
| 83 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12922 | -0.168 | -0.2024 | Yes |
| 84 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 13096 | -0.208 | -0.1886 | Yes |
| 85 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13114 | -0.214 | -0.1623 | Yes |
| 86 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13317 | -0.475 | -0.1162 | Yes |
| 87 | ASCL1 | ASCL1 Entrez, Source | achaete-scute complex-like 1 (Drosophila) | 13340 | -0.915 | 0.0002 | Yes |