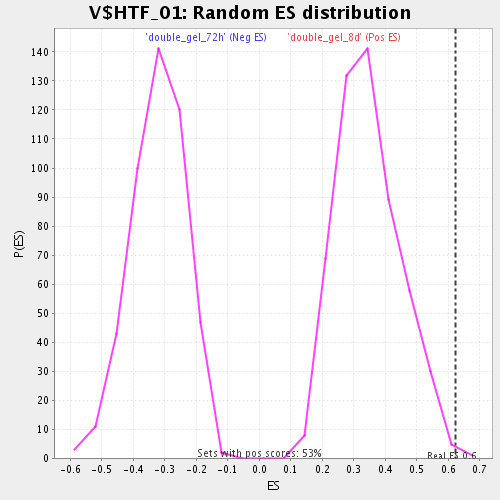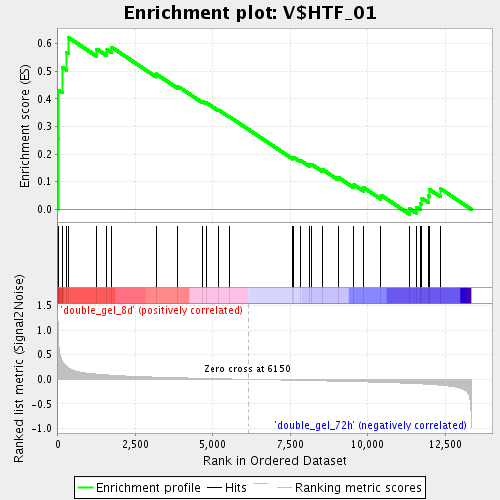
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$HTF_01 |
| Enrichment Score (ES) | 0.6226459 |
| Normalized Enrichment Score (NES) | 1.796662 |
| Nominal p-value | 0.0037523452 |
| FDR q-value | 0.032474656 |
| FWER p-Value | 0.972 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 3 | 1.046 | 0.2576 | Yes |
| 2 | ARMCX2 | ARMCX2 Entrez, Source | armadillo repeat containing, X-linked 2 | 19 | 0.707 | 0.4308 | Yes |
| 3 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 140 | 0.372 | 0.5136 | Yes |
| 4 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 278 | 0.263 | 0.5681 | Yes |
| 5 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 330 | 0.237 | 0.6226 | Yes |
| 6 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 1250 | 0.107 | 0.5801 | No |
| 7 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 1582 | 0.090 | 0.5774 | No |
| 8 | STAT3 | STAT3 Entrez, Source | signal transducer and activator of transcription 3 (acute-phase response factor) | 1729 | 0.083 | 0.5870 | No |
| 9 | POLG | POLG Entrez, Source | polymerase (DNA directed), gamma | 3170 | 0.045 | 0.4899 | No |
| 10 | SYT11 | SYT11 Entrez, Source | synaptotagmin XI | 3867 | 0.032 | 0.4456 | No |
| 11 | UFC1 | UFC1 Entrez, Source | ubiquitin-fold modifier conjugating enzyme 1 | 4668 | 0.020 | 0.3904 | No |
| 12 | DNM3 | DNM3 Entrez, Source | dynamin 3 | 4784 | 0.018 | 0.3864 | No |
| 13 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 5189 | 0.013 | 0.3592 | No |
| 14 | PIP5K1A | PIP5K1A Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, alpha | 5528 | 0.008 | 0.3357 | No |
| 15 | MORF4L2 | MORF4L2 Entrez, Source | mortality factor 4 like 2 | 7580 | -0.018 | 0.1860 | No |
| 16 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 7602 | -0.018 | 0.1889 | No |
| 17 | FKBP14 | FKBP14 Entrez, Source | FK506 binding protein 14, 22 kDa | 7815 | -0.021 | 0.1782 | No |
| 18 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 8111 | -0.025 | 0.1622 | No |
| 19 | UBQLN1 | UBQLN1 Entrez, Source | ubiquilin 1 | 8177 | -0.026 | 0.1638 | No |
| 20 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 8524 | -0.031 | 0.1455 | No |
| 21 | NUMBL | NUMBL Entrez, Source | numb homolog (Drosophila)-like | 9040 | -0.038 | 0.1162 | No |
| 22 | SRP54 | SRP54 Entrez, Source | signal recognition particle 54kDa | 9555 | -0.046 | 0.0889 | No |
| 23 | NLGN2 | NLGN2 Entrez, Source | neuroligin 2 | 9868 | -0.051 | 0.0781 | No |
| 24 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 10426 | -0.061 | 0.0512 | No |
| 25 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 11340 | -0.083 | 0.0030 | No |
| 26 | SND1 | SND1 Entrez, Source | - | 11565 | -0.088 | 0.0079 | No |
| 27 | SEC61A1 | SEC61A1 Entrez, Source | Sec61 alpha 1 subunit (S. cerevisiae) | 11691 | -0.092 | 0.0211 | No |
| 28 | COG3 | COG3 Entrez, Source | component of oligomeric golgi complex 3 | 11746 | -0.094 | 0.0401 | No |
| 29 | VDP | VDP Entrez, Source | - | 11959 | -0.101 | 0.0491 | No |
| 30 | DHDDS | DHDDS Entrez, Source | dehydrodolichyl diphosphate synthase | 11997 | -0.103 | 0.0717 | No |
| 31 | CREB3L1 | CREB3L1 Entrez, Source | cAMP responsive element binding protein 3-like 1 | 12355 | -0.119 | 0.0741 | No |