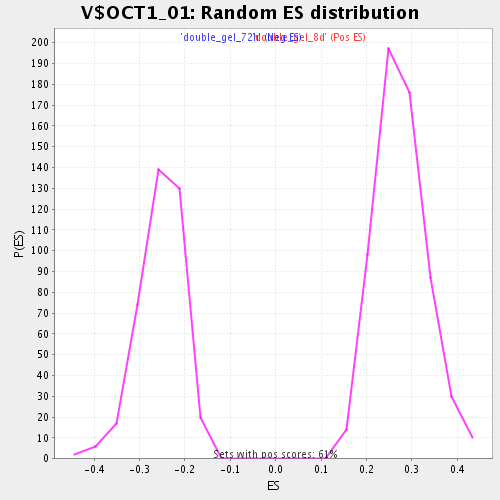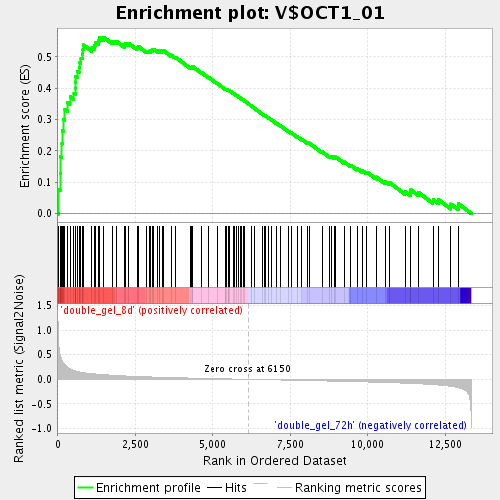
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$OCT1_01 |
| Enrichment Score (ES) | 0.5628102 |
| Normalized Enrichment Score (NES) | 2.0470338 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.00338059 |
| FWER p-Value | 0.094 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPL | LPL Entrez, Source | lipoprotein lipase | 24 | 0.680 | 0.0764 | Yes |
| 2 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 73 | 0.484 | 0.1285 | Yes |
| 3 | SLC24A3 | SLC24A3 Entrez, Source | solute carrier family 24 (sodium/potassium/calcium exchanger), member 3 | 81 | 0.467 | 0.1817 | Yes |
| 4 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 130 | 0.390 | 0.2230 | Yes |
| 5 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 145 | 0.367 | 0.2642 | Yes |
| 6 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 174 | 0.343 | 0.3016 | Yes |
| 7 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 222 | 0.305 | 0.3331 | Yes |
| 8 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 311 | 0.244 | 0.3545 | Yes |
| 9 | ACTG2 | ACTG2 Entrez, Source | actin, gamma 2, smooth muscle, enteric | 391 | 0.212 | 0.3730 | Yes |
| 10 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.3842 | Yes |
| 11 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 559 | 0.171 | 0.4008 | Yes |
| 12 | CPD | CPD Entrez, Source | carboxypeptidase D | 565 | 0.171 | 0.4200 | Yes |
| 13 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 573 | 0.169 | 0.4389 | Yes |
| 14 | GCM1 | GCM1 Entrez, Source | glial cells missing homolog 1 (Drosophila) | 623 | 0.162 | 0.4538 | Yes |
| 15 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 685 | 0.152 | 0.4668 | Yes |
| 16 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 708 | 0.150 | 0.4823 | Yes |
| 17 | NFIA | NFIA Entrez, Source | nuclear factor I/A | 743 | 0.145 | 0.4965 | Yes |
| 18 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 785 | 0.141 | 0.5096 | Yes |
| 19 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 806 | 0.139 | 0.5241 | Yes |
| 20 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 827 | 0.136 | 0.5383 | Yes |
| 21 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 1098 | 0.115 | 0.5311 | Yes |
| 22 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 1183 | 0.110 | 0.5375 | Yes |
| 23 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 1215 | 0.109 | 0.5476 | Yes |
| 24 | DLGAP4 | DLGAP4 Entrez, Source | discs, large (Drosophila) homolog-associated protein 4 | 1317 | 0.103 | 0.5519 | Yes |
| 25 | SYT1 | SYT1 Entrez, Source | synaptotagmin I | 1335 | 0.102 | 0.5623 | Yes |
| 26 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 1475 | 0.095 | 0.5628 | Yes |
| 27 | CDX2 | CDX2 Entrez, Source | caudal type homeobox transcription factor 2 | 1760 | 0.082 | 0.5508 | No |
| 28 | NEUROG1 | NEUROG1 Entrez, Source | neurogenin 1 | 1887 | 0.077 | 0.5502 | No |
| 29 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 2143 | 0.069 | 0.5388 | No |
| 30 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 2181 | 0.068 | 0.5438 | No |
| 31 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 2289 | 0.065 | 0.5432 | No |
| 32 | GPC4 | GPC4 Entrez, Source | glypican 4 | 2553 | 0.058 | 0.5301 | No |
| 33 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 2585 | 0.058 | 0.5344 | No |
| 34 | PDGFRB | PDGFRB Entrez, Source | platelet-derived growth factor receptor, beta polypeptide | 2871 | 0.051 | 0.5188 | No |
| 35 | NFIX | NFIX Entrez, Source | nuclear factor I/X (CCAAT-binding transcription factor) | 2946 | 0.050 | 0.5189 | No |
| 36 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 3000 | 0.048 | 0.5205 | No |
| 37 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 3041 | 0.048 | 0.5230 | No |
| 38 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 3095 | 0.047 | 0.5243 | No |
| 39 | CSN1S1 | CSN1S1 Entrez, Source | casein alpha s1 | 3218 | 0.044 | 0.5202 | No |
| 40 | NUDT3 | NUDT3 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 3 | 3290 | 0.043 | 0.5198 | No |
| 41 | GNB3 | GNB3 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 3 | 3360 | 0.041 | 0.5193 | No |
| 42 | FGF20 | FGF20 Entrez, Source | fibroblast growth factor 20 | 3408 | 0.040 | 0.5204 | No |
| 43 | KCNJ13 | KCNJ13 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 13 | 3652 | 0.036 | 0.5062 | No |
| 44 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 3802 | 0.033 | 0.4988 | No |
| 45 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 4288 | 0.026 | 0.4651 | No |
| 46 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 4310 | 0.025 | 0.4664 | No |
| 47 | ASCL3 | ASCL3 Entrez, Source | achaete-scute complex (Drosophila) homolog-like 3 | 4333 | 0.025 | 0.4677 | No |
| 48 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 4344 | 0.025 | 0.4698 | No |
| 49 | RAB26 | RAB26 Entrez, Source | RAB26, member RAS oncogene family | 4648 | 0.020 | 0.4492 | No |
| 50 | HPCAL4 | HPCAL4 Entrez, Source | hippocalcin like 4 | 4868 | 0.017 | 0.4347 | No |
| 51 | DLL3 | DLL3 Entrez, Source | delta-like 3 (Drosophila) | 5137 | 0.013 | 0.4160 | No |
| 52 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 5410 | 0.010 | 0.3965 | No |
| 53 | SLC24A1 | SLC24A1 Entrez, Source | solute carrier family 24 (sodium/potassium/calcium exchanger), member 1 | 5423 | 0.009 | 0.3967 | No |
| 54 | RHOBTB2 | RHOBTB2 Entrez, Source | Rho-related BTB domain containing 2 | 5450 | 0.009 | 0.3958 | No |
| 55 | FZD4 | FZD4 Entrez, Source | frizzled homolog 4 (Drosophila) | 5495 | 0.008 | 0.3934 | No |
| 56 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 5505 | 0.008 | 0.3937 | No |
| 57 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 5521 | 0.008 | 0.3935 | No |
| 58 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 5661 | 0.006 | 0.3837 | No |
| 59 | NRXN1 | NRXN1 Entrez, Source | neurexin 1 | 5683 | 0.006 | 0.3828 | No |
| 60 | TSGA14 | TSGA14 Entrez, Source | testis specific, 14 | 5776 | 0.005 | 0.3764 | No |
| 61 | IRX2 | IRX2 Entrez, Source | iroquois homeobox protein 2 | 5837 | 0.004 | 0.3723 | No |
| 62 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 5899 | 0.003 | 0.3681 | No |
| 63 | DGKG | DGKG Entrez, Source | diacylglycerol kinase, gamma 90kDa | 5934 | 0.003 | 0.3658 | No |
| 64 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 5987 | 0.002 | 0.3622 | No |
| 65 | KCNN3 | KCNN3 Entrez, Source | potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 | 6008 | 0.002 | 0.3609 | No |
| 66 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 6247 | -0.001 | 0.3431 | No |
| 67 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 6333 | -0.002 | 0.3369 | No |
| 68 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 6617 | -0.006 | 0.3163 | No |
| 69 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 6663 | -0.007 | 0.3136 | No |
| 70 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 6696 | -0.007 | 0.3120 | No |
| 71 | CXXC4 | CXXC4 Entrez, Source | CXXC finger 4 | 6791 | -0.008 | 0.3058 | No |
| 72 | GPR85 | GPR85 Entrez, Source | G protein-coupled receptor 85 | 6883 | -0.009 | 0.3000 | No |
| 73 | CRYZL1 | CRYZL1 Entrez, Source | crystallin, zeta (quinone reductase)-like 1 | 7045 | -0.011 | 0.2892 | No |
| 74 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 7176 | -0.013 | 0.2808 | No |
| 75 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 7445 | -0.016 | 0.2624 | No |
| 76 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 7525 | -0.017 | 0.2584 | No |
| 77 | ATP1B2 | ATP1B2 Entrez, Source | ATPase, Na+/K+ transporting, beta 2 polypeptide | 7724 | -0.020 | 0.2457 | No |
| 78 | ALK | ALK Entrez, Source | anaplastic lymphoma kinase (Ki-1) | 7874 | -0.022 | 0.2370 | No |
| 79 | DNAH7 | DNAH7 Entrez, Source | dynein, axonemal, heavy chain 7 | 8067 | -0.024 | 0.2253 | No |
| 80 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 8105 | -0.025 | 0.2254 | No |
| 81 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 8524 | -0.031 | 0.1974 | No |
| 82 | TCF2 | TCF2 Entrez, Source | transcription factor 2, hepatic; LF-B3; variant hepatic nuclear factor | 8766 | -0.034 | 0.1831 | No |
| 83 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 8836 | -0.035 | 0.1820 | No |
| 84 | TSNAX | TSNAX Entrez, Source | translin-associated factor X | 8938 | -0.037 | 0.1786 | No |
| 85 | GPR4 | GPR4 Entrez, Source | G protein-coupled receptor 4 | 8960 | -0.037 | 0.1813 | No |
| 86 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 9243 | -0.041 | 0.1647 | No |
| 87 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 9444 | -0.044 | 0.1546 | No |
| 88 | PHC2 | PHC2 Entrez, Source | polyhomeotic homolog 2 (Drosophila) | 9654 | -0.048 | 0.1444 | No |
| 89 | NXPH1 | NXPH1 Entrez, Source | neurexophilin 1 | 9817 | -0.050 | 0.1379 | No |
| 90 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 9971 | -0.053 | 0.1325 | No |
| 91 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10269 | -0.058 | 0.1167 | No |
| 92 | NDNL2 | NDNL2 Entrez, Source | necdin-like 2 | 10563 | -0.063 | 0.1019 | No |
| 93 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 10692 | -0.066 | 0.0999 | No |
| 94 | ACBD4 | ACBD4 Entrez, Source | acyl-Coenzyme A binding domain containing 4 | 11202 | -0.078 | 0.0704 | No |
| 95 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 11378 | -0.083 | 0.0668 | No |
| 96 | SYNPR | SYNPR Entrez, Source | synaptoporin | 11391 | -0.084 | 0.0756 | No |
| 97 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 11645 | -0.090 | 0.0668 | No |
| 98 | BAT2 | BAT2 Entrez, Source | HLA-B associated transcript 2 | 12105 | -0.107 | 0.0445 | No |
| 99 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 12283 | -0.115 | 0.0444 | No |
| 100 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 12682 | -0.140 | 0.0305 | No |
| 101 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12922 | -0.168 | 0.0317 | No |