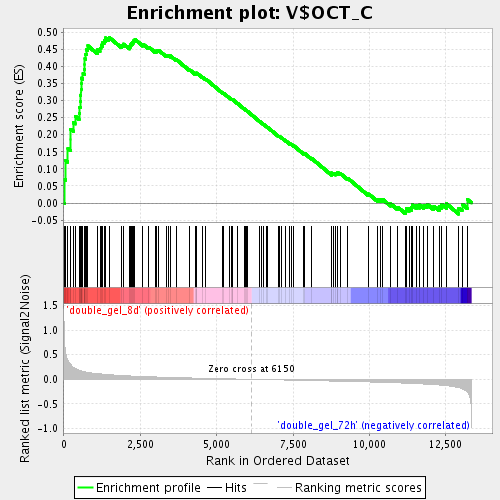
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$OCT_C |
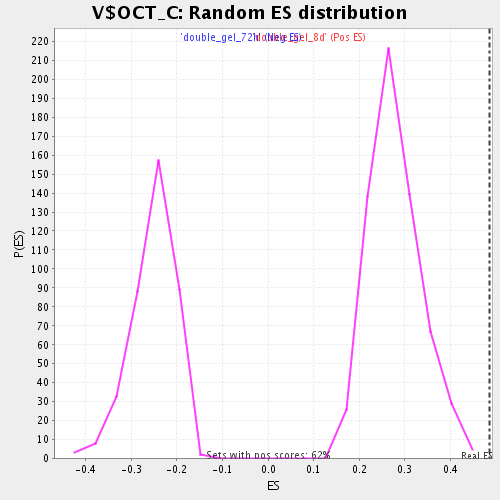
| Enrichment Score (ES) | 0.4856234 |
| Normalized Enrichment Score (NES) | 1.7423365 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04323908 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPL | LPL Entrez, Source | lipoprotein lipase | 24 | 0.680 | 0.0689 | Yes |
| 2 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 55 | 0.564 | 0.1252 | Yes |
| 3 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 130 | 0.390 | 0.1602 | Yes |
| 4 | GPM6A | GPM6A Entrez, Source | glycoprotein M6A | 221 | 0.305 | 0.1851 | Yes |
| 5 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 222 | 0.305 | 0.2168 | Yes |
| 6 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 311 | 0.244 | 0.2355 | Yes |
| 7 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 366 | 0.223 | 0.2546 | Yes |
| 8 | DPYSL3 | DPYSL3 Entrez, Source | dihydropyrimidinase-like 3 | 499 | 0.182 | 0.2636 | Yes |
| 9 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.2810 | Yes |
| 10 | GNA14 | GNA14 Entrez, Source | guanine nucleotide binding protein (G protein), alpha 14 | 536 | 0.175 | 0.2978 | Yes |
| 11 | ADORA1 | ADORA1 Entrez, Source | adenosine A1 receptor | 547 | 0.173 | 0.3150 | Yes |
| 12 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 559 | 0.171 | 0.3319 | Yes |
| 13 | PER2 | PER2 Entrez, Source | period homolog 2 (Drosophila) | 566 | 0.171 | 0.3492 | Yes |
| 14 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 573 | 0.169 | 0.3663 | Yes |
| 15 | GCM1 | GCM1 Entrez, Source | glial cells missing homolog 1 (Drosophila) | 623 | 0.162 | 0.3794 | Yes |
| 16 | SLC10A2 | SLC10A2 Entrez, Source | solute carrier family 10 (sodium/bile acid cotransporter family), member 2 | 675 | 0.153 | 0.3915 | Yes |
| 17 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 685 | 0.152 | 0.4067 | Yes |
| 18 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 688 | 0.151 | 0.4222 | Yes |
| 19 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 708 | 0.150 | 0.4364 | Yes |
| 20 | NFIA | NFIA Entrez, Source | nuclear factor I/A | 743 | 0.145 | 0.4489 | Yes |
| 21 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 785 | 0.141 | 0.4605 | Yes |
| 22 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 1098 | 0.115 | 0.4489 | Yes |
| 23 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 1183 | 0.110 | 0.4540 | Yes |
| 24 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 1215 | 0.109 | 0.4629 | Yes |
| 25 | LPHN1 | LPHN1 Entrez, Source | latrophilin 1 | 1271 | 0.105 | 0.4697 | Yes |
| 26 | DLGAP4 | DLGAP4 Entrez, Source | discs, large (Drosophila) homolog-associated protein 4 | 1317 | 0.103 | 0.4771 | Yes |
| 27 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 1367 | 0.100 | 0.4838 | Yes |
| 28 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 1475 | 0.095 | 0.4856 | Yes |
| 29 | NEUROG1 | NEUROG1 Entrez, Source | neurogenin 1 | 1887 | 0.077 | 0.4626 | No |
| 30 | FGF12 | FGF12 Entrez, Source | fibroblast growth factor 12 | 1947 | 0.075 | 0.4659 | No |
| 31 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 2143 | 0.069 | 0.4583 | No |
| 32 | VSNL1 | VSNL1 Entrez, Source | visinin-like 1 | 2169 | 0.068 | 0.4635 | No |
| 33 | TAS2R13 | TAS2R13 Entrez, Source | taste receptor, type 2, member 13 | 2202 | 0.067 | 0.4681 | No |
| 34 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 2250 | 0.066 | 0.4714 | No |
| 35 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 2289 | 0.065 | 0.4752 | No |
| 36 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 2324 | 0.064 | 0.4793 | No |
| 37 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 2585 | 0.058 | 0.4657 | No |
| 38 | MRAS | MRAS Entrez, Source | muscle RAS oncogene homolog | 2774 | 0.053 | 0.4570 | No |
| 39 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 3000 | 0.048 | 0.4450 | No |
| 40 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 3041 | 0.048 | 0.4470 | No |
| 41 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 3095 | 0.047 | 0.4478 | No |
| 42 | CDH10 | CDH10 Entrez, Source | cadherin 10, type 2 (T2-cadherin) | 3356 | 0.041 | 0.4325 | No |
| 43 | FGF20 | FGF20 Entrez, Source | fibroblast growth factor 20 | 3408 | 0.040 | 0.4328 | No |
| 44 | EGLN2 | EGLN2 Entrez, Source | egl nine homolog 2 (C. elegans) | 3481 | 0.039 | 0.4314 | No |
| 45 | GAP43 | GAP43 Entrez, Source | growth associated protein 43 | 3668 | 0.036 | 0.4211 | No |
| 46 | SLC39A13 | SLC39A13 Entrez, Source | solute carrier family 39 (zinc transporter), member 13 | 4119 | 0.028 | 0.3900 | No |
| 47 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 4310 | 0.025 | 0.3783 | No |
| 48 | ASCL3 | ASCL3 Entrez, Source | achaete-scute complex (Drosophila) homolog-like 3 | 4333 | 0.025 | 0.3792 | No |
| 49 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 4344 | 0.025 | 0.3810 | No |
| 50 | BLNK | BLNK Entrez, Source | B-cell linker | 4539 | 0.022 | 0.3687 | No |
| 51 | RAB26 | RAB26 Entrez, Source | RAB26, member RAS oncogene family | 4648 | 0.020 | 0.3626 | No |
| 52 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 5189 | 0.013 | 0.3231 | No |
| 53 | TEAD3 | TEAD3 Entrez, Source | TEA domain family member 3 | 5228 | 0.012 | 0.3215 | No |
| 54 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 5410 | 0.010 | 0.3089 | No |
| 55 | FZD4 | FZD4 Entrez, Source | frizzled homolog 4 (Drosophila) | 5495 | 0.008 | 0.3034 | No |
| 56 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 5505 | 0.008 | 0.3036 | No |
| 57 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 5521 | 0.008 | 0.3033 | No |
| 58 | NRXN1 | NRXN1 Entrez, Source | neurexin 1 | 5683 | 0.006 | 0.2917 | No |
| 59 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 5899 | 0.003 | 0.2758 | No |
| 60 | DGKG | DGKG Entrez, Source | diacylglycerol kinase, gamma 90kDa | 5934 | 0.003 | 0.2735 | No |
| 61 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 5987 | 0.002 | 0.2699 | No |
| 62 | KCNN3 | KCNN3 Entrez, Source | potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 | 6008 | 0.002 | 0.2686 | No |
| 63 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 6415 | -0.003 | 0.2382 | No |
| 64 | JUND | JUND Entrez, Source | jun D proto-oncogene | 6457 | -0.004 | 0.2355 | No |
| 65 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 6534 | -0.005 | 0.2303 | No |
| 66 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 6617 | -0.006 | 0.2247 | No |
| 67 | SEMA6C | SEMA6C Entrez, Source | sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6C | 6673 | -0.007 | 0.2213 | No |
| 68 | FGF14 | FGF14 Entrez, Source | fibroblast growth factor 14 | 7024 | -0.011 | 0.1960 | No |
| 69 | CRYZL1 | CRYZL1 Entrez, Source | crystallin, zeta (quinone reductase)-like 1 | 7045 | -0.011 | 0.1956 | No |
| 70 | ARHGAP4 | ARHGAP4 Entrez, Source | Rho GTPase activating protein 4 | 7114 | -0.012 | 0.1917 | No |
| 71 | TMOD2 | TMOD2 Entrez, Source | tropomodulin 2 (neuronal) | 7253 | -0.014 | 0.1827 | No |
| 72 | BSND | BSND Entrez, Source | Bartter syndrome, infantile, with sensorineural deafness (Barttin) | 7398 | -0.015 | 0.1734 | No |
| 73 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 7445 | -0.016 | 0.1716 | No |
| 74 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 7525 | -0.017 | 0.1674 | No |
| 75 | GALNT1 | GALNT1 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 1 (GalNAc-T1) | 7850 | -0.021 | 0.1452 | No |
| 76 | ALK | ALK Entrez, Source | anaplastic lymphoma kinase (Ki-1) | 7874 | -0.022 | 0.1457 | No |
| 77 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 8105 | -0.025 | 0.1309 | No |
| 78 | MYBPC1 | MYBPC1 Entrez, Source | myosin binding protein C, slow type | 8761 | -0.034 | 0.0850 | No |
| 79 | TCF2 | TCF2 Entrez, Source | transcription factor 2, hepatic; LF-B3; variant hepatic nuclear factor | 8766 | -0.034 | 0.0883 | No |
| 80 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 8836 | -0.035 | 0.0867 | No |
| 81 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 8900 | -0.036 | 0.0857 | No |
| 82 | TSNAX | TSNAX Entrez, Source | translin-associated factor X | 8938 | -0.037 | 0.0868 | No |
| 83 | GPR4 | GPR4 Entrez, Source | G protein-coupled receptor 4 | 8960 | -0.037 | 0.0890 | No |
| 84 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 9039 | -0.038 | 0.0871 | No |
| 85 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 9285 | -0.042 | 0.0729 | No |
| 86 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 9971 | -0.053 | 0.0267 | No |
| 87 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10269 | -0.058 | 0.0103 | No |
| 88 | POU2F3 | POU2F3 Entrez, Source | POU domain, class 2, transcription factor 3 | 10351 | -0.059 | 0.0103 | No |
| 89 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 10426 | -0.061 | 0.0110 | No |
| 90 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 10692 | -0.066 | -0.0021 | No |
| 91 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 10932 | -0.072 | -0.0127 | No |
| 92 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 11186 | -0.078 | -0.0238 | No |
| 93 | CRISP1 | CRISP1 Entrez, Source | cysteine-rich secretory protein 1 | 11196 | -0.078 | -0.0163 | No |
| 94 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 11295 | -0.081 | -0.0153 | No |
| 95 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 11378 | -0.083 | -0.0128 | No |
| 96 | SYNPR | SYNPR Entrez, Source | synaptoporin | 11391 | -0.084 | -0.0050 | No |
| 97 | IGJ | IGJ Entrez, Source | immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides | 11533 | -0.087 | -0.0065 | No |
| 98 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 11645 | -0.090 | -0.0056 | No |
| 99 | GRID2 | GRID2 Entrez, Source | glutamate receptor, ionotropic, delta 2 | 11782 | -0.095 | -0.0060 | No |
| 100 | SMPX | SMPX Entrez, Source | small muscle protein, X-linked | 11886 | -0.099 | -0.0035 | No |
| 101 | BAT2 | BAT2 Entrez, Source | HLA-B associated transcript 2 | 12105 | -0.107 | -0.0088 | No |
| 102 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 12283 | -0.115 | -0.0102 | No |
| 103 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 12353 | -0.119 | -0.0031 | No |
| 104 | ABTB2 | ABTB2 Entrez, Source | ankyrin repeat and BTB (POZ) domain containing 2 | 12516 | -0.129 | -0.0019 | No |
| 105 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12922 | -0.168 | -0.0150 | No |
| 106 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 13033 | -0.194 | -0.0032 | No |
| 107 | FGFR2 | FGFR2 Entrez, Source | ffer syndrome, Jackson-Weiss syndrome) | 13200 | -0.255 | 0.0107 | No |