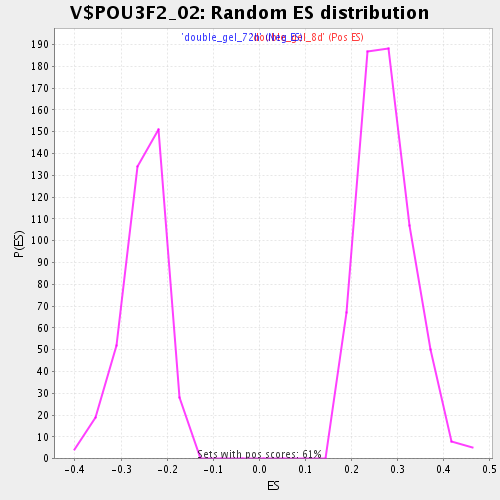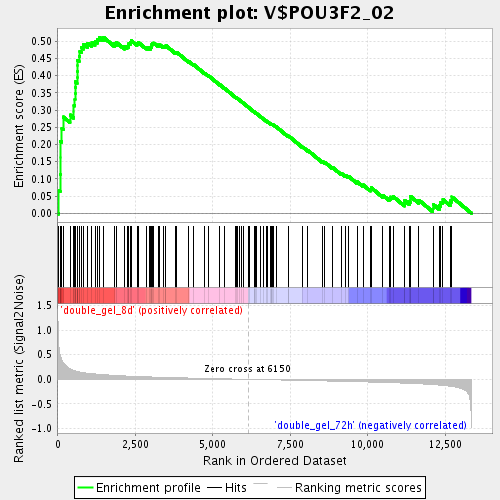
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$POU3F2_02 |
| Enrichment Score (ES) | 0.5115655 |
| Normalized Enrichment Score (NES) | 1.8546596 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02287026 |
| FWER p-Value | 0.842 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ADM | ADM Entrez, Source | adrenomedullin | 34 | 0.649 | 0.0652 | Yes |
| 2 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 73 | 0.484 | 0.1130 | Yes |
| 3 | SLC24A3 | SLC24A3 Entrez, Source | solute carrier family 24 (sodium/potassium/calcium exchanger), member 3 | 81 | 0.467 | 0.1613 | Yes |
| 4 | LOX | LOX Entrez, Source | lysyl oxidase | 82 | 0.465 | 0.2099 | Yes |
| 5 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 128 | 0.391 | 0.2474 | Yes |
| 6 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 174 | 0.343 | 0.2799 | Yes |
| 7 | ACTG2 | ACTG2 Entrez, Source | actin, gamma 2, smooth muscle, enteric | 391 | 0.212 | 0.2857 | Yes |
| 8 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 516 | 0.180 | 0.2952 | Yes |
| 9 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 517 | 0.180 | 0.3140 | Yes |
| 10 | RAB3C | RAB3C Entrez, Source | RAB3C, member RAS oncogene family | 549 | 0.172 | 0.3296 | Yes |
| 11 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 553 | 0.172 | 0.3474 | Yes |
| 12 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 559 | 0.171 | 0.3649 | Yes |
| 13 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 568 | 0.170 | 0.3820 | Yes |
| 14 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 619 | 0.162 | 0.3952 | Yes |
| 15 | GCM1 | GCM1 Entrez, Source | glial cells missing homolog 1 (Drosophila) | 623 | 0.162 | 0.4119 | Yes |
| 16 | BIRC4 | BIRC4 Entrez, Source | baculoviral IAP repeat-containing 4 | 630 | 0.161 | 0.4283 | Yes |
| 17 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 633 | 0.160 | 0.4449 | Yes |
| 18 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 685 | 0.152 | 0.4569 | Yes |
| 19 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 708 | 0.150 | 0.4709 | Yes |
| 20 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 756 | 0.144 | 0.4824 | Yes |
| 21 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 827 | 0.136 | 0.4914 | Yes |
| 22 | GABRG2 | GABRG2 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, gamma 2 | 962 | 0.124 | 0.4943 | Yes |
| 23 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 1098 | 0.115 | 0.4961 | Yes |
| 24 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 1198 | 0.109 | 0.5001 | Yes |
| 25 | LPHN1 | LPHN1 Entrez, Source | latrophilin 1 | 1271 | 0.105 | 0.5057 | Yes |
| 26 | SYT1 | SYT1 Entrez, Source | synaptotagmin I | 1335 | 0.102 | 0.5116 | Yes |
| 27 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 1475 | 0.095 | 0.5110 | No |
| 28 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 1820 | 0.080 | 0.4933 | No |
| 29 | NEUROG1 | NEUROG1 Entrez, Source | neurogenin 1 | 1887 | 0.077 | 0.4964 | No |
| 30 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 2143 | 0.069 | 0.4843 | No |
| 31 | CXADR | CXADR Entrez, Source | coxsackie virus and adenovirus receptor | 2231 | 0.066 | 0.4847 | No |
| 32 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 2279 | 0.065 | 0.4879 | No |
| 33 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 2289 | 0.065 | 0.4940 | No |
| 34 | FKRP | FKRP Entrez, Source | fukutin related protein | 2335 | 0.064 | 0.4973 | No |
| 35 | LGI1 | LGI1 Entrez, Source | leucine-rich, glioma inactivated 1 | 2365 | 0.063 | 0.5017 | No |
| 36 | GPC4 | GPC4 Entrez, Source | glypican 4 | 2553 | 0.058 | 0.4936 | No |
| 37 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 2585 | 0.058 | 0.4973 | No |
| 38 | TFB2M | TFB2M Entrez, Source | transcription factor B2, mitochondrial | 2853 | 0.052 | 0.4826 | No |
| 39 | NFIX | NFIX Entrez, Source | nuclear factor I/X (CCAAT-binding transcription factor) | 2946 | 0.050 | 0.4808 | No |
| 40 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 3000 | 0.048 | 0.4819 | No |
| 41 | HOXA4 | HOXA4 Entrez, Source | homeobox A4 | 3020 | 0.048 | 0.4855 | No |
| 42 | DCN | DCN Entrez, Source | decorin | 3029 | 0.048 | 0.4899 | No |
| 43 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 3041 | 0.048 | 0.4940 | No |
| 44 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 3095 | 0.047 | 0.4949 | No |
| 45 | GBX2 | GBX2 Entrez, Source | gastrulation brain homeobox 2 | 3234 | 0.044 | 0.4891 | No |
| 46 | NUDT3 | NUDT3 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 3 | 3290 | 0.043 | 0.4894 | No |
| 47 | FGF20 | FGF20 Entrez, Source | fibroblast growth factor 20 | 3408 | 0.040 | 0.4848 | No |
| 48 | SLC22A17 | SLC22A17 Entrez, Source | solute carrier family 22 (organic cation transporter), member 17 | 3460 | 0.039 | 0.4850 | No |
| 49 | GDI1 | GDI1 Entrez, Source | GDP dissociation inhibitor 1 | 3486 | 0.039 | 0.4872 | No |
| 50 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 3802 | 0.033 | 0.4669 | No |
| 51 | NEO1 | NEO1 Entrez, Source | neogenin homolog 1 (chicken) | 3835 | 0.033 | 0.4679 | No |
| 52 | PSMA1 | PSMA1 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 1 | 4225 | 0.027 | 0.4413 | No |
| 53 | UBB | UBB Entrez, Source | ubiquitin B | 4379 | 0.024 | 0.4322 | No |
| 54 | POU1F1 | POU1F1 Entrez, Source | POU domain, class 1, transcription factor 1 (Pit1, growth hormone factor 1) | 4741 | 0.019 | 0.4070 | No |
| 55 | HPCAL4 | HPCAL4 Entrez, Source | hippocalcin like 4 | 4868 | 0.017 | 0.3992 | No |
| 56 | TEAD3 | TEAD3 Entrez, Source | TEA domain family member 3 | 5228 | 0.012 | 0.3734 | No |
| 57 | CYP4B1 | CYP4B1 Entrez, Source | cytochrome P450, family 4, subfamily B, polypeptide 1 | 5380 | 0.010 | 0.3630 | No |
| 58 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 5737 | 0.005 | 0.3367 | No |
| 59 | SHH | SHH Entrez, Source | sonic hedgehog homolog (Drosophila) | 5779 | 0.005 | 0.3341 | No |
| 60 | LMNA | LMNA Entrez, Source | lamin A/C | 5805 | 0.004 | 0.3326 | No |
| 61 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 5875 | 0.004 | 0.3278 | No |
| 62 | DGKG | DGKG Entrez, Source | diacylglycerol kinase, gamma 90kDa | 5934 | 0.003 | 0.3237 | No |
| 63 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 5987 | 0.002 | 0.3200 | No |
| 64 | UNC13D | UNC13D Entrez, Source | unc-13 homolog D (C. elegans) | 6157 | -0.000 | 0.3073 | No |
| 65 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 6189 | -0.000 | 0.3050 | No |
| 66 | KCNK2 | KCNK2 Entrez, Source | potassium channel, subfamily K, member 2 | 6328 | -0.002 | 0.2948 | No |
| 67 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 6333 | -0.002 | 0.2947 | No |
| 68 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 6392 | -0.003 | 0.2907 | No |
| 69 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 6415 | -0.003 | 0.2893 | No |
| 70 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 6534 | -0.005 | 0.2809 | No |
| 71 | MAL2 | MAL2 Entrez, Source | mal, T-cell differentiation protein 2 | 6641 | -0.006 | 0.2736 | No |
| 72 | PANK4 | PANK4 Entrez, Source | pantothenate kinase 4 | 6721 | -0.007 | 0.2684 | No |
| 73 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 6769 | -0.008 | 0.2656 | No |
| 74 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 6860 | -0.009 | 0.2598 | No |
| 75 | GPR85 | GPR85 Entrez, Source | G protein-coupled receptor 85 | 6883 | -0.009 | 0.2591 | No |
| 76 | XYLT2 | XYLT2 Entrez, Source | xylosyltransferase II | 6905 | -0.009 | 0.2585 | No |
| 77 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 6929 | -0.010 | 0.2578 | No |
| 78 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 6971 | -0.010 | 0.2557 | No |
| 79 | CRYZL1 | CRYZL1 Entrez, Source | crystallin, zeta (quinone reductase)-like 1 | 7045 | -0.011 | 0.2514 | No |
| 80 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 7439 | -0.016 | 0.2234 | No |
| 81 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 7445 | -0.016 | 0.2247 | No |
| 82 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 7901 | -0.022 | 0.1926 | No |
| 83 | DNAH7 | DNAH7 Entrez, Source | dynein, axonemal, heavy chain 7 | 8067 | -0.024 | 0.1827 | No |
| 84 | MAP2K5 | MAP2K5 Entrez, Source | mitogen-activated protein kinase kinase 5 | 8547 | -0.031 | 0.1497 | No |
| 85 | PLCD4 | PLCD4 Entrez, Source | phospholipase C, delta 4 | 8610 | -0.032 | 0.1484 | No |
| 86 | SOSTDC1 | SOSTDC1 Entrez, Source | sclerostin domain containing 1 | 8869 | -0.036 | 0.1327 | No |
| 87 | TNNI1 | TNNI1 Entrez, Source | troponin I type 1 (skeletal, slow) | 9147 | -0.039 | 0.1159 | No |
| 88 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 9285 | -0.042 | 0.1099 | No |
| 89 | AGTR2 | AGTR2 Entrez, Source | angiotensin II receptor, type 2 | 9378 | -0.043 | 0.1074 | No |
| 90 | PHC2 | PHC2 Entrez, Source | polyhomeotic homolog 2 (Drosophila) | 9654 | -0.048 | 0.0916 | No |
| 91 | PNMA1 | PNMA1 Entrez, Source | paraneoplastic antigen MA1 | 9848 | -0.051 | 0.0824 | No |
| 92 | CXXC5 | CXXC5 Entrez, Source | CXXC finger 5 | 10086 | -0.055 | 0.0702 | No |
| 93 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 10110 | -0.055 | 0.0742 | No |
| 94 | HTR4 | HTR4 Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 4 | 10483 | -0.062 | 0.0526 | No |
| 95 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 10692 | -0.066 | 0.0438 | No |
| 96 | CDC2L1 | CDC2L1 Entrez, Source | cell division cycle 2-like 1 (PITSLRE proteins) | 10741 | -0.067 | 0.0472 | No |
| 97 | SLC18A3 | SLC18A3 Entrez, Source | solute carrier family 18 (vesicular acetylcholine), member 3 | 10815 | -0.069 | 0.0488 | No |
| 98 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 11186 | -0.078 | 0.0290 | No |
| 99 | GIP | GIP Entrez, Source | gastric inhibitory polypeptide | 11194 | -0.078 | 0.0366 | No |
| 100 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 11343 | -0.083 | 0.0341 | No |
| 101 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 11375 | -0.083 | 0.0405 | No |
| 102 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 11378 | -0.083 | 0.0490 | No |
| 103 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 11645 | -0.090 | 0.0384 | No |
| 104 | BAT2 | BAT2 Entrez, Source | HLA-B associated transcript 2 | 12105 | -0.107 | 0.0149 | No |
| 105 | TAPBP | TAPBP Entrez, Source | TAP binding protein (tapasin) | 12109 | -0.108 | 0.0259 | No |
| 106 | MAGED2 | MAGED2 Entrez, Source | melanoma antigen family D, 2 | 12317 | -0.117 | 0.0225 | No |
| 107 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 12353 | -0.119 | 0.0323 | No |
| 108 | MUSK | MUSK Entrez, Source | muscle, skeletal, receptor tyrosine kinase | 12426 | -0.123 | 0.0397 | No |
| 109 | DIXDC1 | DIXDC1 Entrez, Source | DIX domain containing 1 | 12665 | -0.138 | 0.0361 | No |
| 110 | RARB | RARB Entrez, Source | retinoic acid receptor, beta | 12717 | -0.143 | 0.0472 | No |