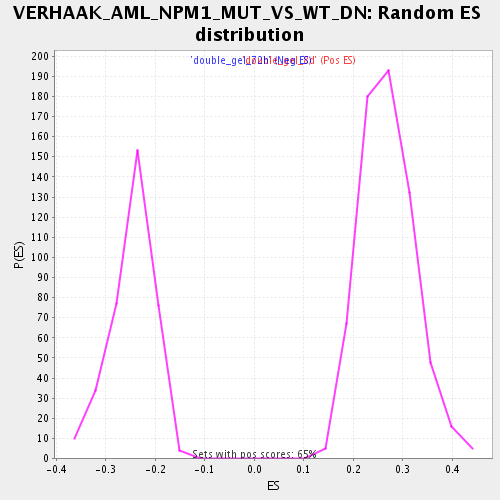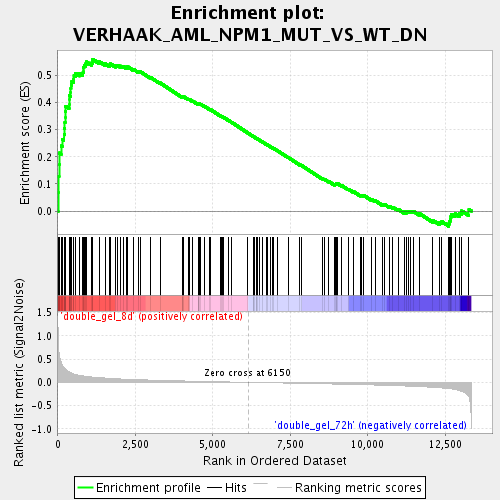
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | VERHAAK_AML_NPM1_MUT_VS_WT_DN |
| Enrichment Score (ES) | 0.5603625 |
| Normalized Enrichment Score (NES) | 2.0792 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0026945102 |
| FWER p-Value | 0.054 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 5 | 0.972 | 0.0695 | Yes |
| 2 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 11 | 0.815 | 0.1277 | Yes |
| 3 | RAMP1 | RAMP1 Entrez, Source | receptor (calcitonin) activity modifying protein 1 | 37 | 0.635 | 0.1715 | Yes |
| 4 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 44 | 0.606 | 0.2146 | Yes |
| 5 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 104 | 0.424 | 0.2406 | Yes |
| 6 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 140 | 0.372 | 0.2647 | Yes |
| 7 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 196 | 0.324 | 0.2838 | Yes |
| 8 | FOXO1A | FOXO1A Entrez, Source | forkhead box O1A (rhabdomyosarcoma) | 213 | 0.311 | 0.3050 | Yes |
| 9 | LAMC1 | LAMC1 Entrez, Source | laminin, gamma 1 (formerly LAMB2) | 224 | 0.303 | 0.3260 | Yes |
| 10 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 234 | 0.297 | 0.3466 | Yes |
| 11 | CD48 | CD48 Entrez, Source | CD48 molecule | 239 | 0.291 | 0.3673 | Yes |
| 12 | TRPM4 | TRPM4 Entrez, Source | transient receptor potential cation channel, subfamily M, member 4 | 250 | 0.282 | 0.3868 | Yes |
| 13 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 361 | 0.225 | 0.3946 | Yes |
| 14 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 366 | 0.223 | 0.4104 | Yes |
| 15 | TMOD1 | TMOD1 Entrez, Source | tropomodulin 1 | 370 | 0.220 | 0.4259 | Yes |
| 16 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 412 | 0.206 | 0.4377 | Yes |
| 17 | GSN | GSN Entrez, Source | gelsolin (amyloidosis, Finnish type) | 417 | 0.205 | 0.4521 | Yes |
| 18 | P2RX5 | P2RX5 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 5 | 423 | 0.203 | 0.4663 | Yes |
| 19 | PROM1 | PROM1 Entrez, Source | prominin 1 | 448 | 0.196 | 0.4786 | Yes |
| 20 | F2RL1 | F2RL1 Entrez, Source | coagulation factor II (thrombin) receptor-like 1 | 508 | 0.181 | 0.4871 | Yes |
| 21 | EHD3 | EHD3 Entrez, Source | EH-domain containing 3 | 518 | 0.179 | 0.4994 | Yes |
| 22 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 568 | 0.170 | 0.5079 | Yes |
| 23 | ITM2C | ITM2C Entrez, Source | integral membrane protein 2C | 696 | 0.150 | 0.5091 | Yes |
| 24 | MLLT3 | MLLT3 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 | 808 | 0.139 | 0.5107 | Yes |
| 25 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 825 | 0.137 | 0.5193 | Yes |
| 26 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 827 | 0.136 | 0.5290 | Yes |
| 27 | FCER1A | FCER1A Entrez, Source | Fc fragment of IgE, high affinity I, receptor for; alpha polypeptide | 844 | 0.135 | 0.5375 | Yes |
| 28 | CD69 | CD69 Entrez, Source | CD69 molecule | 898 | 0.129 | 0.5428 | Yes |
| 29 | TGFB1I1 | TGFB1I1 Entrez, Source | transforming growth factor beta 1 induced transcript 1 | 919 | 0.127 | 0.5504 | Yes |
| 30 | HSPB1 | HSPB1 Entrez, Source | heat shock 27kDa protein 1 | 1084 | 0.116 | 0.5463 | Yes |
| 31 | NBL1 | NBL1 Entrez, Source | neuroblastoma, suppression of tumorigenicity 1 | 1115 | 0.114 | 0.5523 | Yes |
| 32 | B4GALT6 | B4GALT6 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 6 | 1117 | 0.114 | 0.5604 | Yes |
| 33 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 1333 | 0.102 | 0.5514 | No |
| 34 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 1522 | 0.093 | 0.5439 | No |
| 35 | IL5RA | IL5RA Entrez, Source | interleukin 5 receptor, alpha | 1677 | 0.086 | 0.5384 | No |
| 36 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1690 | 0.085 | 0.5437 | No |
| 37 | SERINC5 | SERINC5 Entrez, Source | serine incorporator 5 | 1858 | 0.078 | 0.5366 | No |
| 38 | WASF1 | WASF1 Entrez, Source | WAS protein family, member 1 | 1914 | 0.076 | 0.5379 | No |
| 39 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 2019 | 0.073 | 0.5353 | No |
| 40 | ALAS2 | ALAS2 Entrez, Source | aminolevulinate, delta-, synthase 2 (sideroblastic/hypochromic anemia) | 2100 | 0.070 | 0.5342 | No |
| 41 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 2200 | 0.067 | 0.5316 | No |
| 42 | FZD6 | FZD6 Entrez, Source | frizzled homolog 6 (Drosophila) | 2258 | 0.066 | 0.5320 | No |
| 43 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 2424 | 0.061 | 0.5239 | No |
| 44 | GMPR | GMPR Entrez, Source | guanosine monophosphate reductase | 2611 | 0.057 | 0.5140 | No |
| 45 | ASS1 | ASS1 Entrez, Source | argininosuccinate synthetase 1 | 2650 | 0.056 | 0.5152 | No |
| 46 | BZRAP1 | BZRAP1 Entrez, Source | benzodiazapine receptor (peripheral) associated protein 1 | 2978 | 0.049 | 0.4939 | No |
| 47 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 3298 | 0.043 | 0.4729 | No |
| 48 | TMEM158 | TMEM158 Entrez, Source | transmembrane protein 158 | 4032 | 0.030 | 0.4196 | No |
| 49 | TRH | TRH Entrez, Source | thyrotropin-releasing hormone | 4036 | 0.030 | 0.4215 | No |
| 50 | SNCA | SNCA Entrez, Source | synuclein, alpha (non A4 component of amyloid precursor) | 4052 | 0.029 | 0.4225 | No |
| 51 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 4207 | 0.027 | 0.4127 | No |
| 52 | SH3BP4 | SH3BP4 Entrez, Source | SH3-domain binding protein 4 | 4238 | 0.026 | 0.4124 | No |
| 53 | KIT | KIT Entrez, Source | v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog | 4345 | 0.025 | 0.4061 | No |
| 54 | GNG7 | GNG7 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 7 | 4525 | 0.022 | 0.3942 | No |
| 55 | BLNK | BLNK Entrez, Source | B-cell linker | 4539 | 0.022 | 0.3948 | No |
| 56 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 4547 | 0.022 | 0.3958 | No |
| 57 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 4566 | 0.022 | 0.3960 | No |
| 58 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 4610 | 0.021 | 0.3942 | No |
| 59 | CD2 | CD2 Entrez, Source | CD2 molecule | 4735 | 0.019 | 0.3862 | No |
| 60 | MYH11 | MYH11 Entrez, Source | myosin, heavy chain 11, smooth muscle | 4899 | 0.017 | 0.3751 | No |
| 61 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 4934 | 0.016 | 0.3737 | No |
| 62 | NCAM1 | NCAM1 Entrez, Source | neural cell adhesion molecule 1 | 5249 | 0.012 | 0.3508 | No |
| 63 | FHL1 | FHL1 Entrez, Source | four and a half LIM domains 1 | 5266 | 0.012 | 0.3504 | No |
| 64 | OPTN | OPTN Entrez, Source | optineurin | 5299 | 0.011 | 0.3488 | No |
| 65 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 5356 | 0.010 | 0.3453 | No |
| 66 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 5501 | 0.008 | 0.3350 | No |
| 67 | BPGM | BPGM Entrez, Source | 2,3-bisphosphoglycerate mutase | 5600 | 0.007 | 0.3281 | No |
| 68 | NRXN2 | NRXN2 Entrez, Source | neurexin 2 | 6107 | 0.001 | 0.2899 | No |
| 69 | MSLN | MSLN Entrez, Source | mesothelin | 6309 | -0.002 | 0.2748 | No |
| 70 | CD3D | CD3D Entrez, Source | CD3d molecule, delta (CD3-TCR complex) | 6312 | -0.002 | 0.2748 | No |
| 71 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 6337 | -0.002 | 0.2732 | No |
| 72 | EVL | EVL Entrez, Source | Enah/Vasp-like | 6399 | -0.003 | 0.2688 | No |
| 73 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 6410 | -0.003 | 0.2683 | No |
| 74 | MST1 | MST1 Entrez, Source | macrophage stimulating 1 (hepatocyte growth factor-like) | 6456 | -0.004 | 0.2651 | No |
| 75 | RAP2A | RAP2A Entrez, Source | RAP2A, member of RAS oncogene family | 6516 | -0.005 | 0.2610 | No |
| 76 | BAALC | BAALC Entrez, Source | brain and acute leukemia, cytoplasmic | 6612 | -0.006 | 0.2542 | No |
| 77 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6734 | -0.007 | 0.2456 | No |
| 78 | EGFL7 | EGFL7 Entrez, Source | EGF-like-domain, multiple 7 | 6762 | -0.008 | 0.2441 | No |
| 79 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 6866 | -0.009 | 0.2370 | No |
| 80 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 6929 | -0.010 | 0.2330 | No |
| 81 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 6967 | -0.010 | 0.2309 | No |
| 82 | CD38 | CD38 Entrez, Source | CD38 molecule | 7086 | -0.012 | 0.2229 | No |
| 83 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 7435 | -0.016 | 0.1977 | No |
| 84 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 7809 | -0.021 | 0.1710 | No |
| 85 | CIITA | CIITA Entrez, Source | class II, major histocompatibility complex, transactivator | 7855 | -0.021 | 0.1691 | No |
| 86 | ADCY3 | ADCY3 Entrez, Source | adenylate cyclase 3 | 8531 | -0.031 | 0.1203 | No |
| 87 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 8603 | -0.032 | 0.1172 | No |
| 88 | DLK1 | DLK1 Entrez, Source | delta-like 1 homolog (Drosophila) | 8739 | -0.034 | 0.1095 | No |
| 89 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 8926 | -0.036 | 0.0980 | No |
| 90 | SCCPDH | SCCPDH Entrez, Source | saccharopine dehydrogenase (putative) | 8949 | -0.037 | 0.0990 | No |
| 91 | CCR2 | CCR2 Entrez, Source | chemokine (C-C motif) receptor 2 | 8978 | -0.037 | 0.0996 | No |
| 92 | TPSAB1 | TPSAB1 Entrez, Source | tryptase alpha/beta 1 | 8986 | -0.037 | 0.1017 | No |
| 93 | ROBO1 | ROBO1 Entrez, Source | roundabout, axon guidance receptor, homolog 1 (Drosophila) | 9015 | -0.038 | 0.1023 | No |
| 94 | NARFL | NARFL Entrez, Source | nuclear prelamin A recognition factor-like | 9164 | -0.040 | 0.0940 | No |
| 95 | ST18 | ST18 Entrez, Source | suppression of tumorigenicity 18 (breast carcinoma) (zinc finger protein) | 9371 | -0.043 | 0.0815 | No |
| 96 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 9537 | -0.046 | 0.0723 | No |
| 97 | PRODH | PRODH Entrez, Source | proline dehydrogenase (oxidase) 1 | 9764 | -0.050 | 0.0587 | No |
| 98 | CD200 | CD200 Entrez, Source | CD200 molecule | 9804 | -0.050 | 0.0594 | No |
| 99 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 9869 | -0.051 | 0.0583 | No |
| 100 | PALM | PALM Entrez, Source | paralemmin | 10122 | -0.056 | 0.0432 | No |
| 101 | JUP | JUP Entrez, Source | junction plakoglobin | 10241 | -0.057 | 0.0384 | No |
| 102 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 10486 | -0.062 | 0.0244 | No |
| 103 | LRP3 | LRP3 Entrez, Source | low density lipoprotein receptor-related protein 3 | 10545 | -0.063 | 0.0245 | No |
| 104 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 10686 | -0.066 | 0.0187 | No |
| 105 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 10807 | -0.069 | 0.0145 | No |
| 106 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 10978 | -0.073 | 0.0069 | No |
| 107 | SPON1 | SPON1 Entrez, Source | spondin 1, extracellular matrix protein | 11183 | -0.078 | -0.0030 | No |
| 108 | SYNGR1 | SYNGR1 Entrez, Source | synaptogyrin 1 | 11243 | -0.080 | -0.0017 | No |
| 109 | NT5E | NT5E Entrez, Source | 5'-nucleotidase, ecto (CD73) | 11299 | -0.081 | 0.0000 | No |
| 110 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 11375 | -0.083 | 0.0003 | No |
| 111 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 11468 | -0.085 | -0.0005 | No |
| 112 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 11674 | -0.091 | -0.0095 | No |
| 113 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 12100 | -0.107 | -0.0339 | No |
| 114 | FGF13 | FGF13 Entrez, Source | fibroblast growth factor 13 | 12314 | -0.117 | -0.0416 | No |
| 115 | C1ORF24 | C1ORF24 Entrez, Source | chromosome 1 open reading frame 24 | 12389 | -0.121 | -0.0385 | No |
| 116 | SNRPN /// SNURF | 12618 | -0.136 | -0.0460 | No | ||
| 117 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12642 | -0.137 | -0.0379 | No |
| 118 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 12659 | -0.138 | -0.0292 | No |
| 119 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 12681 | -0.140 | -0.0207 | No |
| 120 | ARHGEF12 | ARHGEF12 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 12 | 12697 | -0.141 | -0.0117 | No |
| 121 | AK3L1 | AK3L1 Entrez, Source | adenylate kinase 3-like 1 | 12819 | -0.155 | -0.0098 | No |
| 122 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 12962 | -0.175 | -0.0079 | No |
| 123 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 13032 | -0.193 | 0.0007 | No |
| 124 | VLDLR | VLDLR Entrez, Source | very low density lipoprotein receptor | 13265 | -0.315 | 0.0058 | No |