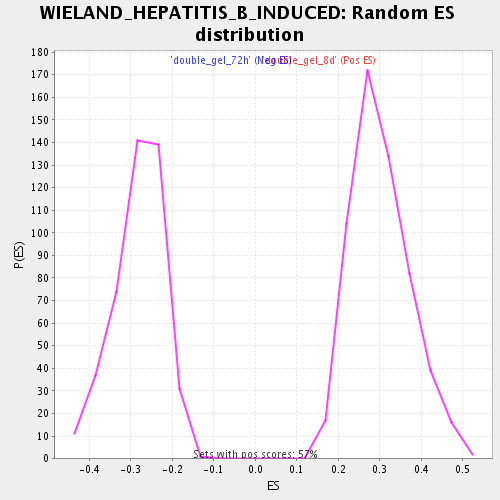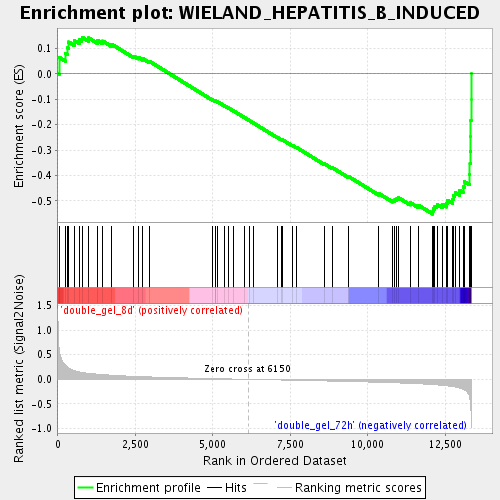
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | WIELAND_HEPATITIS_B_INDUCED |
| Enrichment Score (ES) | -0.5515342 |
| Normalized Enrichment Score (NES) | -1.9674461 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.041946035 |
| FWER p-Value | 0.247 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HLA-DMA | HLA-DMA Entrez, Source | major histocompatibility complex, class II, DM alpha | 40 | 0.620 | 0.0646 | No |
| 2 | CD48 | CD48 Entrez, Source | CD48 molecule | 239 | 0.291 | 0.0815 | No |
| 3 | LPXN | LPXN Entrez, Source | leupaxin | 317 | 0.240 | 0.1019 | No |
| 4 | UCP2 | UCP2 Entrez, Source | uncoupling protein 2 (mitochondrial, proton carrier) | 344 | 0.231 | 0.1252 | No |
| 5 | IGSF6 | IGSF6 Entrez, Source | immunoglobulin superfamily, member 6 | 537 | 0.175 | 0.1298 | No |
| 6 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 697 | 0.150 | 0.1342 | No |
| 7 | HLA-DMB | HLA-DMB Entrez, Source | major histocompatibility complex, class II, DM beta | 777 | 0.142 | 0.1437 | No |
| 8 | GZMK | GZMK Entrez, Source | granzyme K (granzyme 3; tryptase II) | 989 | 0.123 | 0.1412 | No |
| 9 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 1285 | 0.105 | 0.1304 | No |
| 10 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 1444 | 0.096 | 0.1291 | No |
| 11 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 1734 | 0.083 | 0.1164 | No |
| 12 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 2447 | 0.061 | 0.0694 | No |
| 13 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 2597 | 0.058 | 0.0645 | No |
| 14 | GZMA | GZMA Entrez, Source | granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) | 2740 | 0.054 | 0.0597 | No |
| 15 | PPT1 | PPT1 Entrez, Source | palmitoyl-protein thioesterase 1 (ceroid-lipofuscinosis, neuronal 1, infantile) | 2945 | 0.050 | 0.0498 | No |
| 16 | B2M | B2M Entrez, Source | beta-2-microglobulin | 4992 | 0.015 | -0.1026 | No |
| 17 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 5075 | 0.014 | -0.1072 | No |
| 18 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 5082 | 0.014 | -0.1061 | No |
| 19 | SP110 | SP110 Entrez, Source | SP110 nuclear body protein | 5160 | 0.013 | -0.1105 | No |
| 20 | TYROBP | TYROBP Entrez, Source | TYRO protein tyrosine kinase binding protein | 5388 | 0.010 | -0.1265 | No |
| 21 | CD53 | CD53 Entrez, Source | CD53 molecule | 5502 | 0.008 | -0.1341 | No |
| 22 | RAB27A | RAB27A Entrez, Source | RAB27A, member RAS oncogene family | 5665 | 0.006 | -0.1456 | No |
| 23 | IL10RA | IL10RA Entrez, Source | interleukin 10 receptor, alpha | 6007 | 0.002 | -0.1711 | No |
| 24 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 6191 | -0.000 | -0.1848 | No |
| 25 | CD3D | CD3D Entrez, Source | CD3d molecule, delta (CD3-TCR complex) | 6312 | -0.002 | -0.1936 | No |
| 26 | CD38 | CD38 Entrez, Source | CD38 molecule | 7086 | -0.012 | -0.2505 | No |
| 27 | CFL1 | CFL1 Entrez, Source | cofilin 1 (non-muscle) | 7218 | -0.013 | -0.2589 | No |
| 28 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 7239 | -0.014 | -0.2589 | No |
| 29 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 7569 | -0.018 | -0.2818 | No |
| 30 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 7684 | -0.019 | -0.2883 | No |
| 31 | EMR1 | EMR1 Entrez, Source | egf-like module containing, mucin-like, hormone receptor-like 1 | 8587 | -0.032 | -0.3527 | No |
| 32 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 8852 | -0.035 | -0.3687 | No |
| 33 | C3AR1 | C3AR1 Entrez, Source | complement component 3a receptor 1 | 9367 | -0.043 | -0.4027 | No |
| 34 | CTSB | CTSB Entrez, Source | cathepsin B | 10352 | -0.059 | -0.4703 | No |
| 35 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 10803 | -0.068 | -0.4967 | No |
| 36 | GIMAP5 | GIMAP5 Entrez, Source | GTPase, IMAP family member 5 | 10876 | -0.070 | -0.4945 | No |
| 37 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 10939 | -0.072 | -0.4913 | No |
| 38 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 10993 | -0.073 | -0.4873 | No |
| 39 | CTSC | CTSC Entrez, Source | cathepsin C | 11373 | -0.083 | -0.5068 | No |
| 40 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 11635 | -0.090 | -0.5166 | No |
| 41 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 12100 | -0.107 | -0.5398 | Yes |
| 42 | TAPBP | TAPBP Entrez, Source | TAP binding protein (tapasin) | 12109 | -0.108 | -0.5287 | Yes |
| 43 | PSME1 | PSME1 Entrez, Source | proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) | 12165 | -0.110 | -0.5208 | Yes |
| 44 | PSME2 | PSME2 Entrez, Source | proteasome (prosome, macropain) activator subunit 2 (PA28 beta) | 12234 | -0.113 | -0.5137 | Yes |
| 45 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 12406 | -0.122 | -0.5133 | Yes |
| 46 | AURKA | AURKA Entrez, Source | aurora kinase A | 12546 | -0.131 | -0.5094 | Yes |
| 47 | CD68 | CD68 Entrez, Source | CD68 molecule | 12573 | -0.133 | -0.4969 | Yes |
| 48 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 12736 | -0.145 | -0.4932 | Yes |
| 49 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 12760 | -0.148 | -0.4789 | Yes |
| 50 | PSMB10 | PSMB10 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 10 | 12815 | -0.154 | -0.4661 | Yes |
| 51 | LAP3 | LAP3 Entrez, Source | leucine aminopeptidase 3 | 12964 | -0.175 | -0.4582 | Yes |
| 52 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 13079 | -0.204 | -0.4445 | Yes |
| 53 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13114 | -0.214 | -0.4237 | Yes |
| 54 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 13292 | -0.382 | -0.3954 | Yes |
| 55 | CXCL9 | CXCL9 Entrez, Source | chemokine (C-X-C motif) ligand 9 | 13299 | -0.402 | -0.3520 | Yes |
| 56 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 13311 | -0.439 | -0.3049 | Yes |
| 57 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 13324 | -0.530 | -0.2480 | Yes |
| 58 | SLC7A7 | SLC7A7 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | 13328 | -0.584 | -0.1846 | Yes |
| 59 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 13337 | -0.787 | -0.0993 | Yes |
| 60 | CD5L | CD5L Entrez, Source | CD5 molecule-like | 13339 | -0.913 | 0.0002 | Yes |