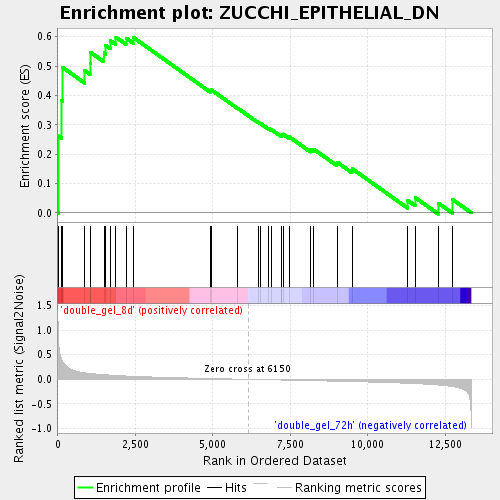
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ZUCCHI_EPITHELIAL_DN |
| Enrichment Score (ES) | 0.59825265 |
| Normalized Enrichment Score (NES) | 1.7360146 |
| Nominal p-value | 0.0053097345 |
| FDR q-value | 0.04525409 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 10 | 0.815 | 0.2622 | Yes |
| 2 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 118 | 0.402 | 0.3837 | Yes |
| 3 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 160 | 0.355 | 0.4949 | Yes |
| 4 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 866 | 0.132 | 0.4847 | Yes |
| 5 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 1037 | 0.118 | 0.5100 | Yes |
| 6 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 1045 | 0.118 | 0.5475 | Yes |
| 7 | PDLIM1 | PDLIM1 Entrez, Source | PDZ and LIM domain 1 (elfin) | 1487 | 0.095 | 0.5449 | Yes |
| 8 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1541 | 0.092 | 0.5705 | Yes |
| 9 | CST3 | CST3 Entrez, Source | cystatin C (amyloid angiopathy and cerebral hemorrhage) | 1692 | 0.085 | 0.5867 | Yes |
| 10 | RASD1 | RASD1 Entrez, Source | RAS, dexamethasone-induced 1 | 1872 | 0.077 | 0.5983 | Yes |
| 11 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 2200 | 0.067 | 0.5954 | No |
| 12 | CLDN4 | CLDN4 Entrez, Source | claudin 4 | 2429 | 0.061 | 0.5980 | No |
| 13 | PNRC1 | PNRC1 Entrez, Source | proline-rich nuclear receptor coactivator 1 | 4913 | 0.016 | 0.4167 | No |
| 14 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 4953 | 0.016 | 0.4189 | No |
| 15 | LMNA | LMNA Entrez, Source | lamin A/C | 5805 | 0.004 | 0.3564 | No |
| 16 | JUND | JUND Entrez, Source | jun D proto-oncogene | 6457 | -0.004 | 0.3087 | No |
| 17 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 6529 | -0.005 | 0.3049 | No |
| 18 | PPM1G | PPM1G Entrez, Source | protein phosphatase 1G (formerly 2C), magnesium-dependent, gamma isoform | 6810 | -0.008 | 0.2866 | No |
| 19 | DDX5 | DDX5 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 5 | 6880 | -0.009 | 0.2844 | No |
| 20 | HMGN1 | HMGN1 Entrez, Source | high-mobility group nucleosome binding domain 1 | 7215 | -0.013 | 0.2636 | No |
| 21 | PFN1 | PFN1 Entrez, Source | profilin 1 | 7229 | -0.013 | 0.2669 | No |
| 22 | AREG | AREG Entrez, Source | amphiregulin (schwannoma-derived growth factor) | 7265 | -0.014 | 0.2687 | No |
| 23 | ZYX | ZYX Entrez, Source | zyxin | 7461 | -0.016 | 0.2593 | No |
| 24 | SCGB1D2 | SCGB1D2 Entrez, Source | secretoglobin, family 1D, member 2 | 8154 | -0.026 | 0.2157 | No |
| 25 | YWHAE | YWHAE Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide | 8261 | -0.027 | 0.2165 | No |
| 26 | EIF4A1 | EIF4A1 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 1 | 9007 | -0.038 | 0.1727 | No |
| 27 | DDB1 | DDB1 Entrez, Source | damage-specific DNA binding protein 1, 127kDa | 9492 | -0.045 | 0.1508 | No |
| 28 | EIF3S4 | EIF3S4 Entrez, Source | eukaryotic translation initiation factor 3, subunit 4 delta, 44kDa | 11289 | -0.081 | 0.0420 | No |
| 29 | ST13 | ST13 Entrez, Source | suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) | 11527 | -0.087 | 0.0524 | No |
| 30 | LGALS3 | LGALS3 Entrez, Source | lectin, galactoside-binding, soluble, 3 (galectin 3) | 12281 | -0.115 | 0.0329 | No |
| 31 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 12736 | -0.145 | 0.0455 | No |

