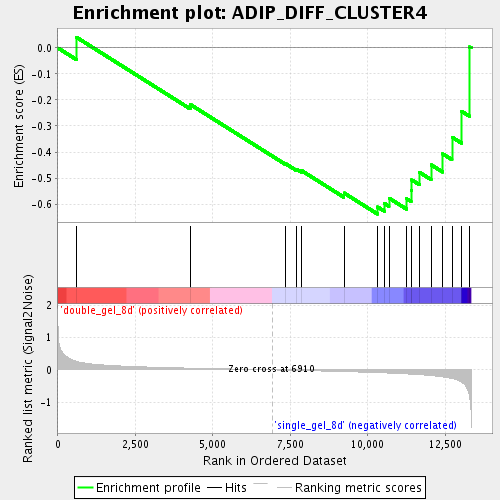
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | ADIP_DIFF_CLUSTER4 |
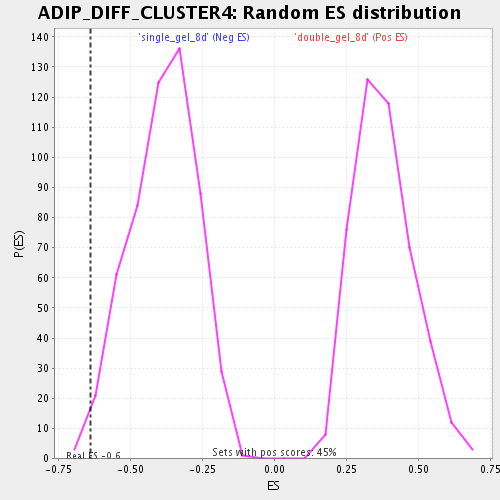
| Enrichment Score (ES) | -0.63870627 |
| Normalized Enrichment Score (NES) | -1.6528276 |
| Nominal p-value | 0.00729927 |
| FDR q-value | 0.09675509 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ASS1 | ASS1 Entrez, Source | argininosuccinate synthetase 1 | 598 | 0.267 | 0.0412 | No |
| 2 | ABCF1 | ABCF1 Entrez, Source | ATP-binding cassette, sub-family F (GCN20), member 1 | 4274 | 0.053 | -0.2174 | No |
| 3 | GRWD1 | GRWD1 Entrez, Source | glutamate-rich WD repeat containing 1 | 7357 | -0.009 | -0.4456 | No |
| 4 | ECM1 | ECM1 Entrez, Source | extracellular matrix protein 1 | 7713 | -0.017 | -0.4668 | No |
| 5 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 7873 | -0.020 | -0.4723 | No |
| 6 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 9233 | -0.052 | -0.5574 | No |
| 7 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 10317 | -0.085 | -0.6112 | Yes |
| 8 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 10538 | -0.093 | -0.5978 | Yes |
| 9 | NAP1L4 | NAP1L4 Entrez, Source | nucleosome assembly protein 1-like 4 | 10686 | -0.098 | -0.5773 | Yes |
| 10 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 11253 | -0.124 | -0.5797 | Yes |
| 11 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 11400 | -0.131 | -0.5485 | Yes |
| 12 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 11420 | -0.132 | -0.5073 | Yes |
| 13 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 11672 | -0.148 | -0.4784 | Yes |
| 14 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 12047 | -0.177 | -0.4494 | Yes |
| 15 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 12419 | -0.217 | -0.4072 | Yes |
| 16 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12720 | -0.269 | -0.3430 | Yes |
| 17 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13028 | -0.381 | -0.2434 | Yes |
| 18 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13289 | -0.828 | 0.0040 | Yes |