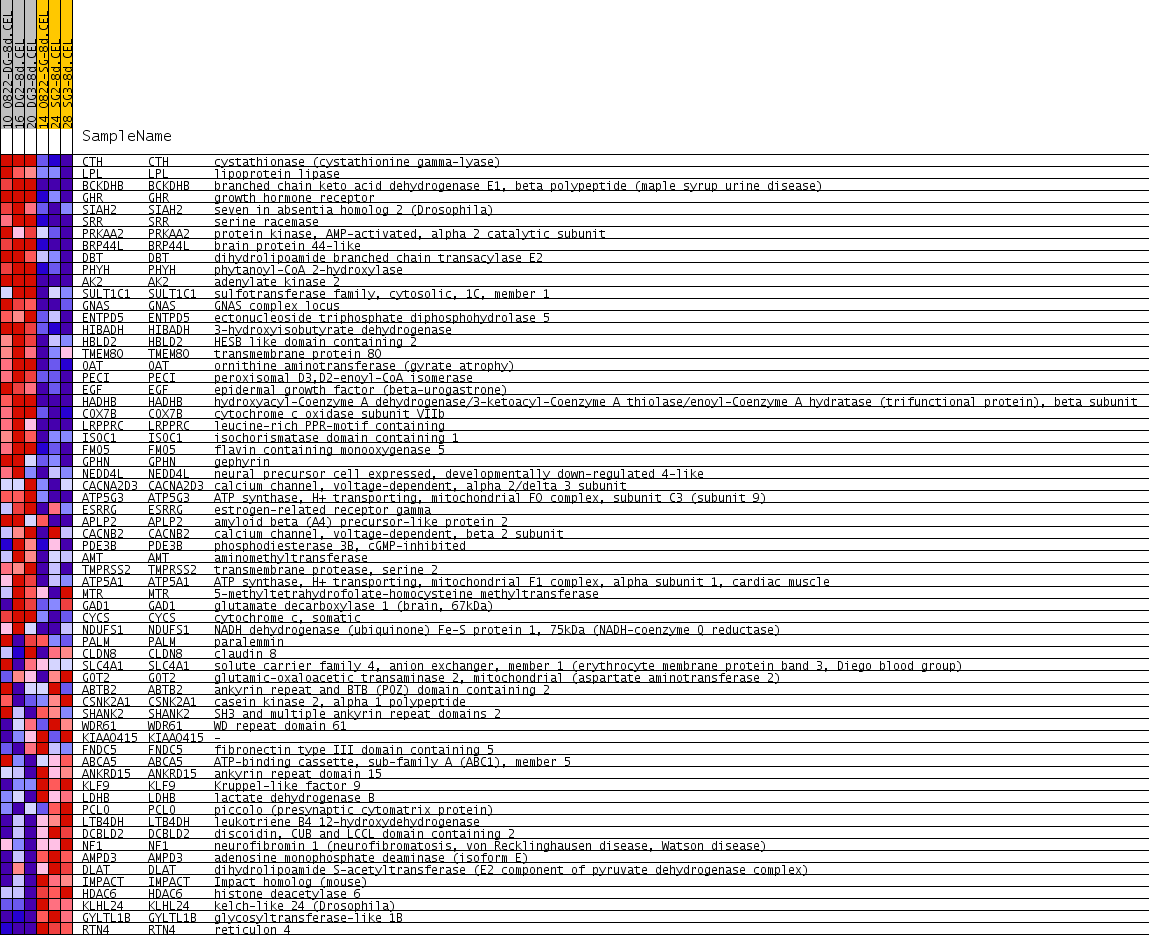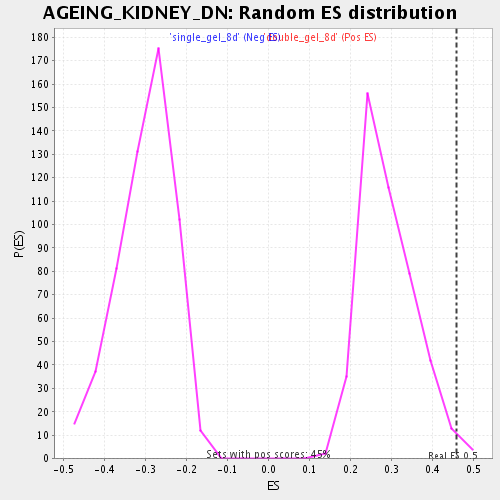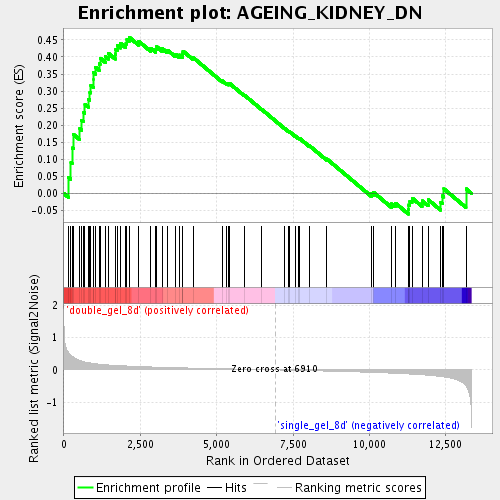
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | AGEING_KIDNEY_DN |
| Enrichment Score (ES) | 0.45768744 |
| Normalized Enrichment Score (NES) | 1.574327 |
| Nominal p-value | 0.013422819 |
| FDR q-value | 0.080461174 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CTH | CTH Entrez, Source | cystathionase (cystathionine gamma-lyase) | 148 | 0.535 | 0.0474 | Yes |
| 2 | LPL | LPL Entrez, Source | lipoprotein lipase | 230 | 0.451 | 0.0907 | Yes |
| 3 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 271 | 0.419 | 0.1335 | Yes |
| 4 | GHR | GHR Entrez, Source | growth hormone receptor | 312 | 0.389 | 0.1730 | Yes |
| 5 | SIAH2 | SIAH2 Entrez, Source | seven in absentia homolog 2 (Drosophila) | 520 | 0.293 | 0.1895 | Yes |
| 6 | SRR | SRR Entrez, Source | serine racemase | 579 | 0.274 | 0.2152 | Yes |
| 7 | PRKAA2 | PRKAA2 Entrez, Source | protein kinase, AMP-activated, alpha 2 catalytic subunit | 649 | 0.252 | 0.2375 | Yes |
| 8 | BRP44L | BRP44L Entrez, Source | brain protein 44-like | 688 | 0.241 | 0.2610 | Yes |
| 9 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 799 | 0.220 | 0.2768 | Yes |
| 10 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 849 | 0.214 | 0.2966 | Yes |
| 11 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 880 | 0.210 | 0.3173 | Yes |
| 12 | SULT1C1 | SULT1C1 Entrez, Source | sulfotransferase family, cytosolic, 1C, member 1 | 964 | 0.199 | 0.3327 | Yes |
| 13 | GNAS | GNAS Entrez, Source | GNAS complex locus | 968 | 0.198 | 0.3542 | Yes |
| 14 | ENTPD5 | ENTPD5 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 5 | 1031 | 0.189 | 0.3702 | Yes |
| 15 | HIBADH | HIBADH Entrez, Source | 3-hydroxyisobutyrate dehydrogenase | 1157 | 0.175 | 0.3799 | Yes |
| 16 | HBLD2 | HBLD2 Entrez, Source | HESB like domain containing 2 | 1186 | 0.172 | 0.3966 | Yes |
| 17 | TMEM80 | TMEM80 Entrez, Source | transmembrane protein 80 | 1365 | 0.160 | 0.4007 | Yes |
| 18 | OAT | OAT Entrez, Source | ornithine aminotransferase (gyrate atrophy) | 1460 | 0.152 | 0.4102 | Yes |
| 19 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1680 | 0.138 | 0.4088 | Yes |
| 20 | EGF | EGF Entrez, Source | epidermal growth factor (beta-urogastrone) | 1695 | 0.137 | 0.4228 | Yes |
| 21 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 1747 | 0.134 | 0.4336 | Yes |
| 22 | COX7B | COX7B Entrez, Source | cytochrome c oxidase subunit VIIb | 1849 | 0.129 | 0.4401 | Yes |
| 23 | LRPPRC | LRPPRC Entrez, Source | leucine-rich PPR-motif containing | 2030 | 0.121 | 0.4398 | Yes |
| 24 | ISOC1 | ISOC1 Entrez, Source | isochorismatase domain containing 1 | 2061 | 0.119 | 0.4505 | Yes |
| 25 | FMO5 | FMO5 Entrez, Source | flavin containing monooxygenase 5 | 2136 | 0.117 | 0.4577 | Yes |
| 26 | GPHN | GPHN Entrez, Source | gephyrin | 2447 | 0.105 | 0.4458 | No |
| 27 | NEDD4L | NEDD4L Entrez, Source | neural precursor cell expressed, developmentally down-regulated 4-like | 2839 | 0.090 | 0.4262 | No |
| 28 | CACNA2D3 | CACNA2D3 Entrez, Source | calcium channel, voltage-dependent, alpha 2/delta 3 subunit | 3006 | 0.085 | 0.4230 | No |
| 29 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 3038 | 0.084 | 0.4298 | No |
| 30 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 3222 | 0.078 | 0.4245 | No |
| 31 | APLP2 | APLP2 Entrez, Source | amyloid beta (A4) precursor-like protein 2 | 3389 | 0.074 | 0.4201 | No |
| 32 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 3644 | 0.068 | 0.4084 | No |
| 33 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 3773 | 0.065 | 0.4058 | No |
| 34 | AMT | AMT Entrez, Source | aminomethyltransferase | 3875 | 0.063 | 0.4050 | No |
| 35 | TMPRSS2 | TMPRSS2 Entrez, Source | transmembrane protease, serine 2 | 3889 | 0.062 | 0.4109 | No |
| 36 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 3895 | 0.062 | 0.4173 | No |
| 37 | MTR | MTR Entrez, Source | 5-methyltetrahydrofolate-homocysteine methyltransferase | 4234 | 0.054 | 0.3978 | No |
| 38 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 5183 | 0.034 | 0.3301 | No |
| 39 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 5320 | 0.032 | 0.3233 | No |
| 40 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 5395 | 0.030 | 0.3211 | No |
| 41 | PALM | PALM Entrez, Source | paralemmin | 5408 | 0.030 | 0.3234 | No |
| 42 | CLDN8 | CLDN8 Entrez, Source | claudin 8 | 5910 | 0.020 | 0.2879 | No |
| 43 | SLC4A1 | SLC4A1 Entrez, Source | solute carrier family 4, anion exchanger, member 1 (erythrocyte membrane protein band 3, Diego blood group) | 6479 | 0.008 | 0.2460 | No |
| 44 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 7234 | -0.007 | 0.1900 | No |
| 45 | ABTB2 | ABTB2 Entrez, Source | ankyrin repeat and BTB (POZ) domain containing 2 | 7363 | -0.010 | 0.1814 | No |
| 46 | CSNK2A1 | CSNK2A1 Entrez, Source | casein kinase 2, alpha 1 polypeptide | 7384 | -0.010 | 0.1810 | No |
| 47 | SHANK2 | SHANK2 Entrez, Source | SH3 and multiple ankyrin repeat domains 2 | 7570 | -0.014 | 0.1686 | No |
| 48 | WDR61 | WDR61 Entrez, Source | WD repeat domain 61 | 7683 | -0.016 | 0.1619 | No |
| 49 | KIAA0415 | KIAA0415 Entrez, Source | - | 7719 | -0.017 | 0.1611 | No |
| 50 | FNDC5 | FNDC5 Entrez, Source | fibronectin type III domain containing 5 | 8027 | -0.024 | 0.1406 | No |
| 51 | ABCA5 | ABCA5 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 5 | 8600 | -0.037 | 0.1016 | No |
| 52 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 10069 | -0.077 | -0.0006 | No |
| 53 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 10129 | -0.079 | 0.0036 | No |
| 54 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 10723 | -0.099 | -0.0302 | No |
| 55 | PCLO | PCLO Entrez, Source | piccolo (presynaptic cytomatrix protein) | 10844 | -0.105 | -0.0278 | No |
| 56 | LTB4DH | LTB4DH Entrez, Source | leukotriene B4 12-hydroxydehydrogenase | 11276 | -0.125 | -0.0465 | No |
| 57 | DCBLD2 | DCBLD2 Entrez, Source | discoidin, CUB and LCCL domain containing 2 | 11288 | -0.126 | -0.0336 | No |
| 58 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 11321 | -0.127 | -0.0221 | No |
| 59 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 11393 | -0.131 | -0.0131 | No |
| 60 | DLAT | DLAT Entrez, Source | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 11723 | -0.150 | -0.0214 | No |
| 61 | IMPACT | IMPACT Entrez, Source | Impact homolog (mouse) | 11924 | -0.167 | -0.0182 | No |
| 62 | HDAC6 | HDAC6 Entrez, Source | histone deacetylase 6 | 12329 | -0.208 | -0.0259 | No |
| 63 | KLHL24 | KLHL24 Entrez, Source | kelch-like 24 (Drosophila) | 12386 | -0.213 | -0.0068 | No |
| 64 | GYLTL1B | GYLTL1B Entrez, Source | glycosyltransferase-like 1B | 12436 | -0.220 | 0.0135 | No |
| 65 | RTN4 | RTN4 Entrez, Source | reticulon 4 | 13160 | -0.500 | 0.0137 | No |