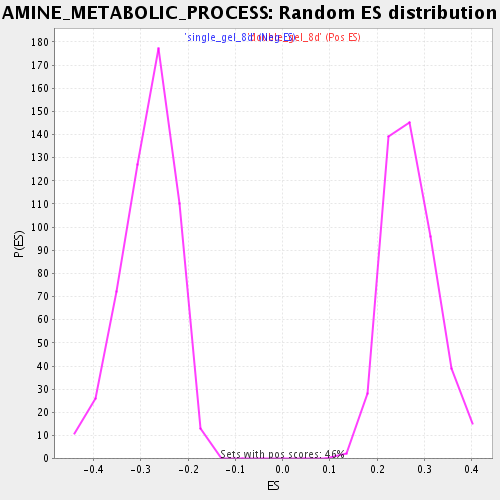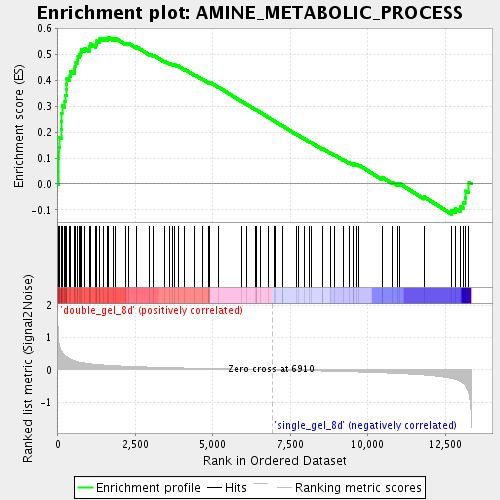
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | AMINE_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.56591094 |
| Normalized Enrichment Score (NES) | 2.0870862 |
| Nominal p-value | 0.0 |
| FDR q-value | 5.700258E-4 |
| FWER p-Value | 0.037 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 16 | 0.933 | 0.0493 | Yes |
| 2 | HGD | HGD Entrez, Source | homogentisate 1,2-dioxygenase (homogentisate oxidase) | 17 | 0.930 | 0.0996 | Yes |
| 3 | MAT1A | MAT1A Entrez, Source | methionine adenosyltransferase I, alpha | 35 | 0.815 | 0.1424 | Yes |
| 4 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 56 | 0.731 | 0.1804 | Yes |
| 5 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 104 | 0.612 | 0.2100 | Yes |
| 6 | SLC25A15 | SLC25A15 Entrez, Source | solute carrier family 25 (mitochondrial carrier; ornithine transporter) member 15 | 113 | 0.597 | 0.2416 | Yes |
| 7 | ASL | ASL Entrez, Source | argininosuccinate lyase | 122 | 0.586 | 0.2727 | Yes |
| 8 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 135 | 0.556 | 0.3019 | Yes |
| 9 | SLC7A2 | SLC7A2 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 | 223 | 0.456 | 0.3200 | Yes |
| 10 | ARG1 | ARG1 Entrez, Source | arginase, liver | 246 | 0.438 | 0.3420 | Yes |
| 11 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 271 | 0.419 | 0.3629 | Yes |
| 12 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 277 | 0.416 | 0.3851 | Yes |
| 13 | GLS2 | GLS2 Entrez, Source | glutaminase 2 (liver, mitochondrial) | 291 | 0.403 | 0.4059 | Yes |
| 14 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 389 | 0.344 | 0.4172 | Yes |
| 15 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 411 | 0.336 | 0.4338 | Yes |
| 16 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 524 | 0.292 | 0.4412 | Yes |
| 17 | SULT1C2 | SULT1C2 Entrez, Source | sulfotransferase family, cytosolic, 1C, member 2 | 550 | 0.282 | 0.4545 | Yes |
| 18 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 580 | 0.273 | 0.4671 | Yes |
| 19 | ADI1 | ADI1 Entrez, Source | acireductone dioxygenase 1 | 623 | 0.260 | 0.4780 | Yes |
| 20 | COLQ | COLQ Entrez, Source | collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase | 646 | 0.252 | 0.4900 | Yes |
| 21 | SLC35A3 | SLC35A3 Entrez, Source | solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3 | 697 | 0.239 | 0.4992 | Yes |
| 22 | SULT1B1 | SULT1B1 Entrez, Source | sulfotransferase family, cytosolic, 1B, member 1 | 743 | 0.229 | 0.5082 | Yes |
| 23 | GAMT | GAMT Entrez, Source | guanidinoacetate N-methyltransferase | 753 | 0.227 | 0.5198 | Yes |
| 24 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 870 | 0.211 | 0.5225 | Yes |
| 25 | SULT1A1 | SULT1A1 Entrez, Source | sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 | 1008 | 0.192 | 0.5225 | Yes |
| 26 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 1027 | 0.189 | 0.5314 | Yes |
| 27 | GCSH | GCSH Entrez, Source | glycine cleavage system protein H (aminomethyl carrier) | 1046 | 0.187 | 0.5402 | Yes |
| 28 | SDS | SDS Entrez, Source | serine dehydratase | 1222 | 0.169 | 0.5361 | Yes |
| 29 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 1252 | 0.167 | 0.5430 | Yes |
| 30 | HDC | HDC Entrez, Source | histidine decarboxylase | 1256 | 0.167 | 0.5518 | Yes |
| 31 | SCLY | SCLY Entrez, Source | selenocysteine lyase | 1345 | 0.161 | 0.5538 | Yes |
| 32 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 1355 | 0.160 | 0.5618 | Yes |
| 33 | CHST3 | CHST3 Entrez, Source | carbohydrate (chondroitin 6) sulfotransferase 3 | 1469 | 0.151 | 0.5615 | Yes |
| 34 | GGTLA1 | GGTLA1 Entrez, Source | gamma-glutamyltransferase-like activity 1 | 1598 | 0.143 | 0.5596 | Yes |
| 35 | NANP | NANP Entrez, Source | N-acetylneuraminic acid phosphatase | 1617 | 0.142 | 0.5659 | Yes |
| 36 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 1785 | 0.132 | 0.5605 | No |
| 37 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 1864 | 0.128 | 0.5615 | No |
| 38 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 2183 | 0.115 | 0.5437 | No |
| 39 | GALNT5 | GALNT5 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (GalNAc-T5) | 2282 | 0.111 | 0.5423 | No |
| 40 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 2527 | 0.102 | 0.5294 | No |
| 41 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 2968 | 0.086 | 0.5009 | No |
| 42 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 3082 | 0.082 | 0.4968 | No |
| 43 | SLC7A8 | SLC7A8 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 8 | 3436 | 0.073 | 0.4741 | No |
| 44 | GSS | GSS Entrez, Source | glutathione synthetase | 3596 | 0.069 | 0.4658 | No |
| 45 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 3691 | 0.067 | 0.4623 | No |
| 46 | SLC7A7 | SLC7A7 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | 3774 | 0.065 | 0.4596 | No |
| 47 | AMT | AMT Entrez, Source | aminomethyltransferase | 3875 | 0.063 | 0.4554 | No |
| 48 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 4100 | 0.057 | 0.4416 | No |
| 49 | XYLT1 | XYLT1 Entrez, Source | xylosyltransferase I | 4421 | 0.050 | 0.4202 | No |
| 50 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 4665 | 0.045 | 0.4043 | No |
| 51 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 4870 | 0.040 | 0.3911 | No |
| 52 | SLC6A6 | SLC6A6 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, taurine), member 6 | 4874 | 0.040 | 0.3930 | No |
| 53 | B4GALT7 | B4GALT7 Entrez, Source | xylosylprotein beta 1,4-galactosyltransferase, polypeptide 7 (galactosyltransferase I) | 4905 | 0.040 | 0.3929 | No |
| 54 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 5183 | 0.034 | 0.3739 | No |
| 55 | SLC5A7 | SLC5A7 Entrez, Source | solute carrier family 5 (choline transporter), member 7 | 5926 | 0.020 | 0.3189 | No |
| 56 | CHST12 | CHST12 Entrez, Source | carbohydrate (chondroitin 4) sulfotransferase 12 | 6090 | 0.016 | 0.3075 | No |
| 57 | MCCC2 | MCCC2 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 2 (beta) | 6381 | 0.010 | 0.2862 | No |
| 58 | GNE | GNE Entrez, Source | glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase | 6397 | 0.010 | 0.2856 | No |
| 59 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 6420 | 0.009 | 0.2844 | No |
| 60 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 6525 | 0.007 | 0.2769 | No |
| 61 | CHIA | CHIA Entrez, Source | chitinase, acidic | 6796 | 0.002 | 0.2567 | No |
| 62 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 6980 | -0.002 | 0.2430 | No |
| 63 | SLC7A9 | SLC7A9 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 | 7015 | -0.002 | 0.2405 | No |
| 64 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 7234 | -0.007 | 0.2245 | No |
| 65 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 7238 | -0.007 | 0.2246 | No |
| 66 | DHPS | DHPS Entrez, Source | deoxyhypusine synthase | 7692 | -0.016 | 0.1913 | No |
| 67 | DARS | DARS Entrez, Source | aspartyl-tRNA synthetase | 7757 | -0.018 | 0.1875 | No |
| 68 | NAGK | NAGK Entrez, Source | N-acetylglucosamine kinase | 7948 | -0.022 | 0.1743 | No |
| 69 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 8124 | -0.026 | 0.1625 | No |
| 70 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 8177 | -0.027 | 0.1601 | No |
| 71 | SLC3A1 | SLC3A1 Entrez, Source | 1 | 8542 | -0.036 | 0.1345 | No |
| 72 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 8548 | -0.036 | 0.1361 | No |
| 73 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 8796 | -0.041 | 0.1197 | No |
| 74 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 8939 | -0.045 | 0.1113 | No |
| 75 | KARS | KARS Entrez, Source | lysyl-tRNA synthetase | 9206 | -0.051 | 0.0941 | No |
| 76 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 9397 | -0.057 | 0.0828 | No |
| 77 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 9551 | -0.061 | 0.0745 | No |
| 78 | GUSB | GUSB Entrez, Source | glucuronidase, beta | 9553 | -0.061 | 0.0778 | No |
| 79 | GGT1 | GGT1 Entrez, Source | gamma-glutamyltransferase 1 | 9642 | -0.064 | 0.0746 | No |
| 80 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 9713 | -0.066 | 0.0729 | No |
| 81 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10462 | -0.090 | 0.0213 | No |
| 82 | CHST1 | CHST1 Entrez, Source | carbohydrate (keratan sulfate Gal-6) sulfotransferase 1 | 10469 | -0.091 | 0.0257 | No |
| 83 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 10791 | -0.102 | 0.0071 | No |
| 84 | SCYE1 | SCYE1 Entrez, Source | small inducible cytokine subfamily E, member 1 (endothelial monocyte-activating) | 10959 | -0.110 | 0.0004 | No |
| 85 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 11033 | -0.113 | 0.0010 | No |
| 86 | EXTL2 | EXTL2 Entrez, Source | exostoses (multiple)-like 2 | 11815 | -0.158 | -0.0494 | No |
| 87 | SPHK1 | SPHK1 Entrez, Source | sphingosine kinase 1 | 12700 | -0.264 | -0.1018 | No |
| 88 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 12832 | -0.296 | -0.0957 | No |
| 89 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 12983 | -0.354 | -0.0879 | No |
| 90 | SMS | SMS Entrez, Source | spermine synthase | 13083 | -0.414 | -0.0729 | No |
| 91 | ST3GAL2 | ST3GAL2 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 2 | 13139 | -0.459 | -0.0522 | No |
| 92 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 13167 | -0.505 | -0.0270 | No |
| 93 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 13266 | -0.741 | 0.0057 | No |