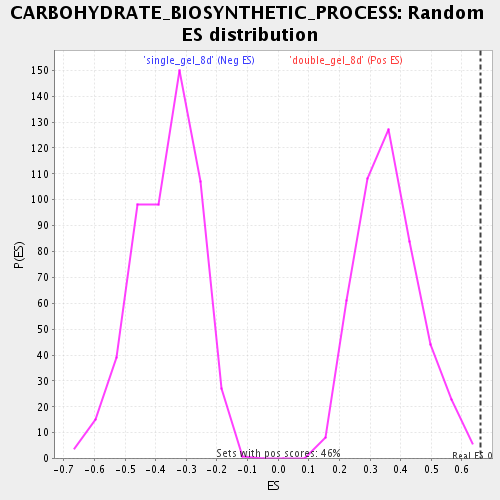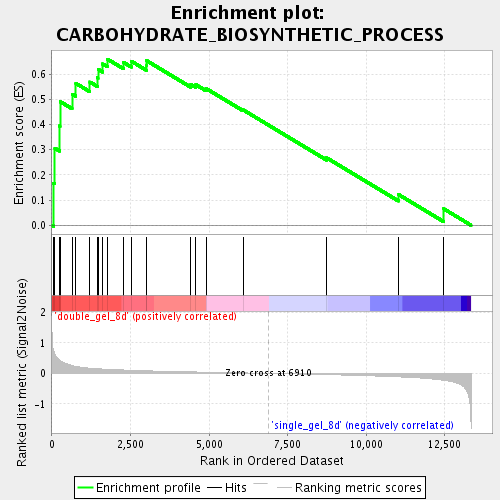
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | CARBOHYDRATE_BIOSYNTHETIC_PROCESS |
| Enrichment Score (ES) | 0.6594724 |
| Normalized Enrichment Score (NES) | 1.8256543 |
| Nominal p-value | 0.0021691974 |
| FDR q-value | 0.011061032 |
| FWER p-Value | 0.805 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GYS2 | GYS2 Entrez, Source | glycogen synthase 2 (liver) | 52 | 0.748 | 0.1670 | Yes |
| 2 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 95 | 0.621 | 0.3058 | Yes |
| 3 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 247 | 0.438 | 0.3945 | Yes |
| 4 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 259 | 0.427 | 0.4913 | Yes |
| 5 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 642 | 0.255 | 0.5210 | Yes |
| 6 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 762 | 0.225 | 0.5636 | Yes |
| 7 | NDST1 | NDST1 Entrez, Source | N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 1 | 1209 | 0.170 | 0.5689 | Yes |
| 8 | ST8SIA2 | ST8SIA2 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 2 | 1450 | 0.153 | 0.5859 | Yes |
| 9 | CHST3 | CHST3 Entrez, Source | carbohydrate (chondroitin 6) sulfotransferase 3 | 1469 | 0.151 | 0.6191 | Yes |
| 10 | NANP | NANP Entrez, Source | N-acetylneuraminic acid phosphatase | 1617 | 0.142 | 0.6406 | Yes |
| 11 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1770 | 0.133 | 0.6595 | Yes |
| 12 | GALNT5 | GALNT5 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (GalNAc-T5) | 2282 | 0.111 | 0.6465 | No |
| 13 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 2527 | 0.102 | 0.6514 | No |
| 14 | MPDU1 | MPDU1 Entrez, Source | mannose-P-dolichol utilization defect 1 | 3005 | 0.085 | 0.6350 | No |
| 15 | ARPP-19 | ARPP-19 Entrez, Source | - | 3013 | 0.084 | 0.6537 | No |
| 16 | XYLT1 | XYLT1 Entrez, Source | xylosyltransferase I | 4421 | 0.050 | 0.5596 | No |
| 17 | B3GAT2 | B3GAT2 Entrez, Source | beta-1,3-glucuronyltransferase 2 (glucuronosyltransferase S) | 4582 | 0.047 | 0.5582 | No |
| 18 | B4GALT7 | B4GALT7 Entrez, Source | xylosylprotein beta 1,4-galactosyltransferase, polypeptide 7 (galactosyltransferase I) | 4905 | 0.040 | 0.5431 | No |
| 19 | CHST12 | CHST12 Entrez, Source | carbohydrate (chondroitin 4) sulfotransferase 12 | 6090 | 0.016 | 0.4580 | No |
| 20 | LALBA | LALBA Entrez, Source | lactalbumin, alpha- | 8747 | -0.040 | 0.2677 | No |
| 21 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 11033 | -0.113 | 0.1221 | No |
| 22 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 12474 | -0.224 | 0.0652 | No |