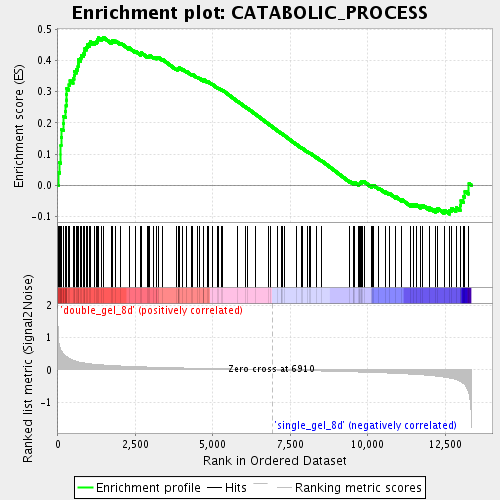
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | CATABOLIC_PROCESS |
| Enrichment Score (ES) | 0.4745522 |
| Normalized Enrichment Score (NES) | 1.8213016 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.011103528 |
| FWER p-Value | 0.824 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HGD | HGD Entrez, Source | homogentisate 1,2-dioxygenase (homogentisate oxidase) | 17 | 0.930 | 0.0415 | Yes |
| 2 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 56 | 0.731 | 0.0723 | Yes |
| 3 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 95 | 0.621 | 0.0980 | Yes |
| 4 | APOA4 | APOA4 Entrez, Source | apolipoprotein A-IV | 96 | 0.620 | 0.1265 | Yes |
| 5 | MIOX | MIOX Entrez, Source | myo-inositol oxygenase | 103 | 0.612 | 0.1542 | Yes |
| 6 | ASL | ASL Entrez, Source | argininosuccinate lyase | 122 | 0.586 | 0.1799 | Yes |
| 7 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 177 | 0.496 | 0.1986 | Yes |
| 8 | ACLY | ACLY Entrez, Source | ATP citrate lyase | 186 | 0.487 | 0.2204 | Yes |
| 9 | ARG1 | ARG1 Entrez, Source | arginase, liver | 246 | 0.438 | 0.2361 | Yes |
| 10 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 260 | 0.427 | 0.2548 | Yes |
| 11 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 271 | 0.419 | 0.2733 | Yes |
| 12 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 277 | 0.416 | 0.2921 | Yes |
| 13 | ZFP36 | ZFP36 Entrez, Source | zinc finger protein 36, C3H type, homolog (mouse) | 286 | 0.407 | 0.3102 | Yes |
| 14 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 356 | 0.360 | 0.3215 | Yes |
| 15 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 389 | 0.344 | 0.3350 | Yes |
| 16 | CES2 | CES2 Entrez, Source | carboxylesterase 2 (intestine, liver) | 489 | 0.305 | 0.3415 | Yes |
| 17 | SIAH2 | SIAH2 Entrez, Source | seven in absentia homolog 2 (Drosophila) | 520 | 0.293 | 0.3527 | Yes |
| 18 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 537 | 0.288 | 0.3648 | Yes |
| 19 | MDM4 | MDM4 Entrez, Source | Mdm4, transformed 3T3 cell double minute 4, p53 binding protein (mouse) | 605 | 0.265 | 0.3719 | Yes |
| 20 | COLQ | COLQ Entrez, Source | collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase | 646 | 0.252 | 0.3805 | Yes |
| 21 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 656 | 0.251 | 0.3914 | Yes |
| 22 | ALDOB | ALDOB Entrez, Source | aldolase B, fructose-bisphosphate | 666 | 0.249 | 0.4021 | Yes |
| 23 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 732 | 0.231 | 0.4079 | Yes |
| 24 | ALDH1L1 | ALDH1L1 Entrez, Source | aldehyde dehydrogenase 1 family, member L1 | 750 | 0.228 | 0.4170 | Yes |
| 25 | APOA5 | APOA5 Entrez, Source | apolipoprotein A-V | 827 | 0.217 | 0.4213 | Yes |
| 26 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 858 | 0.213 | 0.4288 | Yes |
| 27 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 870 | 0.211 | 0.4377 | Yes |
| 28 | TST | TST Entrez, Source | thiosulfate sulfurtransferase (rhodanese) | 926 | 0.203 | 0.4429 | Yes |
| 29 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 950 | 0.200 | 0.4503 | Yes |
| 30 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 1027 | 0.189 | 0.4533 | Yes |
| 31 | GCSH | GCSH Entrez, Source | glycine cleavage system protein H (aminomethyl carrier) | 1046 | 0.187 | 0.4606 | Yes |
| 32 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 1180 | 0.172 | 0.4584 | Yes |
| 33 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 1252 | 0.167 | 0.4607 | Yes |
| 34 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 1278 | 0.165 | 0.4665 | Yes |
| 35 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1296 | 0.164 | 0.4727 | Yes |
| 36 | SDHD | SDHD Entrez, Source | succinate dehydrogenase complex, subunit D, integral membrane protein | 1404 | 0.157 | 0.4719 | Yes |
| 37 | PFKL | PFKL Entrez, Source | phosphofructokinase, liver | 1462 | 0.152 | 0.4746 | Yes |
| 38 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 1743 | 0.134 | 0.4596 | No |
| 39 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 1747 | 0.134 | 0.4655 | No |
| 40 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 1864 | 0.128 | 0.4626 | No |
| 41 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 2032 | 0.120 | 0.4555 | No |
| 42 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 2294 | 0.111 | 0.4409 | No |
| 43 | EGLN2 | EGLN2 Entrez, Source | egl nine homolog 2 (C. elegans) | 2499 | 0.103 | 0.4301 | No |
| 44 | CEL | CEL Entrez, Source | carboxyl ester lipase (bile salt-stimulated lipase) | 2678 | 0.096 | 0.4211 | No |
| 45 | PRSS2 | PRSS2 Entrez, Source | protease, serine, 2 (trypsin 2) | 2681 | 0.096 | 0.4254 | No |
| 46 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2886 | 0.088 | 0.4140 | No |
| 47 | NEU3 | NEU3 Entrez, Source | sialidase 3 (membrane sialidase) | 2937 | 0.087 | 0.4142 | No |
| 48 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 2968 | 0.086 | 0.4159 | No |
| 49 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 3082 | 0.082 | 0.4111 | No |
| 50 | GAPDHS | GAPDHS Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase, spermatogenic | 3168 | 0.079 | 0.4084 | No |
| 51 | BTRC | BTRC Entrez, Source | beta-transducin repeat containing | 3196 | 0.079 | 0.4099 | No |
| 52 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 3231 | 0.078 | 0.4109 | No |
| 53 | UBE2V2 | UBE2V2 Entrez, Source | ubiquitin-conjugating enzyme E2 variant 2 | 3388 | 0.074 | 0.4025 | No |
| 54 | FBXO22 | FBXO22 Entrez, Source | F-box protein 22 | 3823 | 0.063 | 0.3726 | No |
| 55 | AMT | AMT Entrez, Source | aminomethyltransferase | 3875 | 0.063 | 0.3716 | No |
| 56 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 3892 | 0.062 | 0.3733 | No |
| 57 | SDHC | SDHC Entrez, Source | succinate dehydrogenase complex, subunit C, integral membrane protein, 15kDa | 3898 | 0.062 | 0.3758 | No |
| 58 | NPLOC4 | NPLOC4 Entrez, Source | nuclear protein localization 4 homolog (S. cerevisiae) | 3916 | 0.062 | 0.3773 | No |
| 59 | FBXO7 | FBXO7 Entrez, Source | F-box protein 7 | 4032 | 0.059 | 0.3713 | No |
| 60 | SYVN1 | SYVN1 Entrez, Source | synovial apoptosis inhibitor 1, synoviolin | 4157 | 0.056 | 0.3645 | No |
| 61 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4310 | 0.053 | 0.3554 | No |
| 62 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 4353 | 0.052 | 0.3546 | No |
| 63 | DFFA | DFFA Entrez, Source | DNA fragmentation factor, 45kDa, alpha polypeptide | 4493 | 0.049 | 0.3463 | No |
| 64 | PARK2 | PARK2 Entrez, Source | Parkinson disease (autosomal recessive, juvenile) 2, parkin | 4568 | 0.047 | 0.3429 | No |
| 65 | ABCE1 | ABCE1 Entrez, Source | ATP-binding cassette, sub-family E (OABP), member 1 | 4694 | 0.044 | 0.3355 | No |
| 66 | FAF1 | FAF1 Entrez, Source | Fas (TNFRSF6) associated factor 1 | 4712 | 0.044 | 0.3362 | No |
| 67 | MPST | MPST Entrez, Source | mercaptopyruvate sulfurtransferase | 4713 | 0.044 | 0.3382 | No |
| 68 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 4829 | 0.041 | 0.3314 | No |
| 69 | RNASE6 | RNASE6 Entrez, Source | ribonuclease, RNase A family, k6 | 4848 | 0.041 | 0.3319 | No |
| 70 | BLVRA | BLVRA Entrez, Source | biliverdin reductase A | 4973 | 0.038 | 0.3243 | No |
| 71 | SQSTM1 | SQSTM1 Entrez, Source | sequestosome 1 | 5161 | 0.035 | 0.3117 | No |
| 72 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 5183 | 0.034 | 0.3117 | No |
| 73 | UBE2B | UBE2B Entrez, Source | ubiquitin-conjugating enzyme E2B (RAD6 homolog) | 5274 | 0.032 | 0.3064 | No |
| 74 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 5320 | 0.032 | 0.3045 | No |
| 75 | NUDT3 | NUDT3 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 3 | 5804 | 0.022 | 0.2689 | No |
| 76 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 6042 | 0.017 | 0.2518 | No |
| 77 | RNASEH1 | RNASEH1 Entrez, Source | ribonuclease H1 | 6123 | 0.016 | 0.2465 | No |
| 78 | MCCC2 | MCCC2 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 2 (beta) | 6381 | 0.010 | 0.2275 | No |
| 79 | UBE2A | UBE2A Entrez, Source | ubiquitin-conjugating enzyme E2A (RAD6 homolog) | 6804 | 0.002 | 0.1956 | No |
| 80 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 6850 | 0.001 | 0.1923 | No |
| 81 | UBE2N | UBE2N Entrez, Source | ubiquitin-conjugating enzyme E2N (UBC13 homolog, yeast) | 7083 | -0.004 | 0.1749 | No |
| 82 | UBE4A | UBE4A Entrez, Source | ubiquitination factor E4A (UFD2 homolog, yeast) | 7203 | -0.006 | 0.1662 | No |
| 83 | TREH | TREH Entrez, Source | trehalase (brush-border membrane glycoprotein) | 7209 | -0.006 | 0.1661 | No |
| 84 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 7234 | -0.007 | 0.1646 | No |
| 85 | PABPC4 | PABPC4 Entrez, Source | poly(A) binding protein, cytoplasmic 4 (inducible form) | 7324 | -0.009 | 0.1583 | No |
| 86 | DHPS | DHPS Entrez, Source | deoxyhypusine synthase | 7692 | -0.016 | 0.1313 | No |
| 87 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7856 | -0.020 | 0.1198 | No |
| 88 | UBE2G1 | UBE2G1 Entrez, Source | ubiquitin-conjugating enzyme E2G 1 (UBC7 homolog, yeast) | 7901 | -0.021 | 0.1175 | No |
| 89 | PSMC5 | PSMC5 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 5 | 8053 | -0.025 | 0.1072 | No |
| 90 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 8124 | -0.026 | 0.1031 | No |
| 91 | TSG101 | TSG101 Entrez, Source | tumor susceptibility gene 101 | 8160 | -0.027 | 0.1017 | No |
| 92 | GAA | GAA Entrez, Source | glucosidase, alpha; acid (Pompe disease, glycogen storage disease type II) | 8354 | -0.031 | 0.0885 | No |
| 93 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 8494 | -0.034 | 0.0796 | No |
| 94 | ARIH1 | ARIH1 Entrez, Source | ariadne homolog, ubiquitin-conjugating enzyme E2 binding protein, 1 (Drosophila) | 9421 | -0.058 | 0.0121 | No |
| 95 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 9551 | -0.061 | 0.0052 | No |
| 96 | GUSB | GUSB Entrez, Source | glucuronidase, beta | 9553 | -0.061 | 0.0079 | No |
| 97 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 9560 | -0.061 | 0.0103 | No |
| 98 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 9713 | -0.066 | 0.0018 | No |
| 99 | DFFB | DFFB Entrez, Source | DNA fragmentation factor, 40kDa, beta polypeptide (caspase-activated DNase) | 9728 | -0.066 | 0.0038 | No |
| 100 | ALDOA | ALDOA Entrez, Source | aldolase A, fructose-bisphosphate | 9745 | -0.066 | 0.0056 | No |
| 101 | STUB1 | STUB1 Entrez, Source | STIP1 homology and U-box containing protein 1 | 9754 | -0.067 | 0.0081 | No |
| 102 | NEDD8 | NEDD8 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 8 | 9792 | -0.068 | 0.0084 | No |
| 103 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 9794 | -0.068 | 0.0114 | No |
| 104 | ANAPC4 | ANAPC4 Entrez, Source | anaphase promoting complex subunit 4 | 9814 | -0.069 | 0.0132 | No |
| 105 | PPT1 | PPT1 Entrez, Source | palmitoyl-protein thioesterase 1 (ceroid-lipofuscinosis, neuronal 1, infantile) | 9878 | -0.071 | 0.0116 | No |
| 106 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 10127 | -0.079 | -0.0035 | No |
| 107 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 10142 | -0.079 | -0.0009 | No |
| 108 | UBB | UBB Entrez, Source | ubiquitin B | 10179 | -0.081 | 0.0001 | No |
| 109 | VCP | VCP Entrez, Source | valosin-containing protein | 10359 | -0.087 | -0.0095 | No |
| 110 | RNASEH2A | RNASEH2A Entrez, Source | ribonuclease H2, subunit A | 10571 | -0.094 | -0.0211 | No |
| 111 | BAX | BAX Entrez, Source | BCL2-associated X protein | 10699 | -0.098 | -0.0262 | No |
| 112 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 10907 | -0.107 | -0.0369 | No |
| 113 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 11099 | -0.117 | -0.0460 | No |
| 114 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 11393 | -0.131 | -0.0621 | No |
| 115 | NUDT5 | NUDT5 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 5 | 11488 | -0.137 | -0.0630 | No |
| 116 | HGS | HGS Entrez, Source | hepatocyte growth factor-regulated tyrosine kinase substrate | 11569 | -0.141 | -0.0625 | No |
| 117 | USP33 | USP33 Entrez, Source | ubiquitin specific peptidase 33 | 11715 | -0.150 | -0.0666 | No |
| 118 | FRAP1 | FRAP1 Entrez, Source | FK506 binding protein 12-rapamycin associated protein 1 | 11775 | -0.155 | -0.0639 | No |
| 119 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 11989 | -0.172 | -0.0721 | No |
| 120 | CDC23 | CDC23 Entrez, Source | CDC23 (cell division cycle 23, yeast, homolog) | 12182 | -0.190 | -0.0779 | No |
| 121 | ANAPC2 | ANAPC2 Entrez, Source | anaphase promoting complex subunit 2 | 12258 | -0.199 | -0.0744 | No |
| 122 | SMPD3 | SMPD3 Entrez, Source | sphingomyelin phosphodiesterase 3, neutral membrane (neutral sphingomyelinase II) | 12464 | -0.223 | -0.0797 | No |
| 123 | HK1 | HK1 Entrez, Source | hexokinase 1 | 12636 | -0.251 | -0.0811 | No |
| 124 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 12695 | -0.263 | -0.0734 | No |
| 125 | PYGB | PYGB Entrez, Source | phosphorylase, glycogen; brain | 12849 | -0.302 | -0.0710 | No |
| 126 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 12983 | -0.354 | -0.0648 | No |
| 127 | PFKM | PFKM Entrez, Source | phosphofructokinase, muscle | 13007 | -0.369 | -0.0496 | No |
| 128 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 13099 | -0.425 | -0.0369 | No |
| 129 | GPX3 | GPX3 Entrez, Source | glutathione peroxidase 3 (plasma) | 13136 | -0.457 | -0.0186 | No |
| 130 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 13266 | -0.741 | 0.0058 | No |

