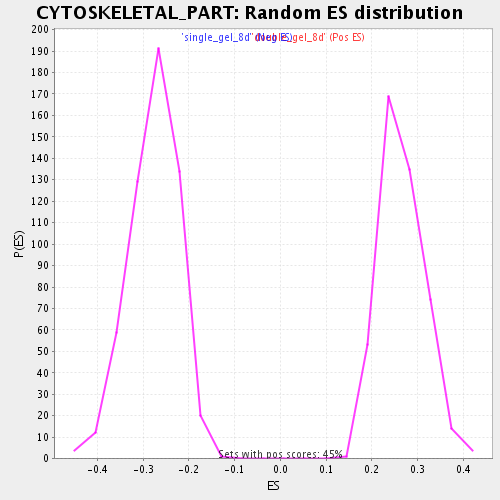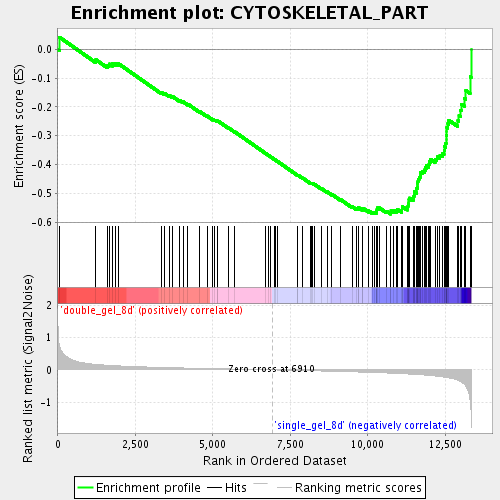
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | CYTOSKELETAL_PART |
| Enrichment Score (ES) | -0.572493 |
| Normalized Enrichment Score (NES) | -2.0679195 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0041405754 |
| FWER p-Value | 0.04 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GYS2 | GYS2 Entrez, Source | glycogen synthase 2 (liver) | 52 | 0.748 | 0.0420 | No |
| 2 | UXT | UXT Entrez, Source | ubiquitously-expressed transcript | 1225 | 0.169 | -0.0361 | No |
| 3 | NARF | NARF Entrez, Source | nuclear prelamin A recognition factor | 1605 | 0.143 | -0.0560 | No |
| 4 | SPTBN4 | SPTBN4 Entrez, Source | spectrin, beta, non-erythrocytic 4 | 1651 | 0.140 | -0.0508 | No |
| 5 | TNNC1 | TNNC1 Entrez, Source | troponin C type 1 (slow) | 1752 | 0.134 | -0.0501 | No |
| 6 | MYL3 | MYL3 Entrez, Source | myosin, light chain 3, alkali; ventricular, skeletal, slow | 1844 | 0.129 | -0.0491 | No |
| 7 | CACNA1C | CACNA1C Entrez, Source | calcium channel, voltage-dependent, L type, alpha 1C subunit | 1956 | 0.124 | -0.0499 | No |
| 8 | DNALI1 | DNALI1 Entrez, Source | dynein, axonemal, light intermediate chain 1 | 3330 | 0.075 | -0.1490 | No |
| 9 | DYNLL2 | DYNLL2 Entrez, Source | dynein, light chain, LC8-type 2 | 3451 | 0.072 | -0.1536 | No |
| 10 | MYH6 | MYH6 Entrez, Source | myosin, heavy chain 6, cardiac muscle, alpha (cardiomyopathy, hypertrophic 1) | 3603 | 0.069 | -0.1608 | No |
| 11 | DYNC1I1 | DYNC1I1 Entrez, Source | dynein, cytoplasmic 1, intermediate chain 1 | 3708 | 0.066 | -0.1646 | No |
| 12 | NUDT21 | NUDT21 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 21 | 3927 | 0.061 | -0.1773 | No |
| 13 | DRD2 | DRD2 Entrez, Source | dopamine receptor D2 | 4037 | 0.059 | -0.1819 | No |
| 14 | TNNI3 | TNNI3 Entrez, Source | troponin I type 3 (cardiac) | 4196 | 0.055 | -0.1905 | No |
| 15 | TNNT2 | TNNT2 Entrez, Source | troponin T type 2 (cardiac) | 4572 | 0.047 | -0.2159 | No |
| 16 | MAPT | MAPT Entrez, Source | microtubule-associated protein tau | 4812 | 0.042 | -0.2314 | No |
| 17 | DCX | DCX Entrez, Source | doublecortex; lissencephaly, X-linked (doublecortin) | 5004 | 0.038 | -0.2435 | No |
| 18 | ACTR1B | ACTR1B Entrez, Source | ARP1 actin-related protein 1 homolog B, centractin beta (yeast) | 5068 | 0.037 | -0.2460 | No |
| 19 | APPBP2 | APPBP2 Entrez, Source | amyloid beta precursor protein (cytoplasmic tail) binding protein 2 | 5141 | 0.035 | -0.2493 | No |
| 20 | ACTN3 | ACTN3 Entrez, Source | actinin, alpha 3 | 5159 | 0.035 | -0.2485 | No |
| 21 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 5506 | 0.028 | -0.2729 | No |
| 22 | EML2 | EML2 Entrez, Source | echinoderm microtubule associated protein like 2 | 5714 | 0.023 | -0.2871 | No |
| 23 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 6709 | 0.004 | -0.3620 | No |
| 24 | ATG4B | ATG4B Entrez, Source | ATG4 autophagy related 4 homolog B (S. cerevisiae) | 6786 | 0.003 | -0.3676 | No |
| 25 | CETN3 | CETN3 Entrez, Source | centrin, EF-hand protein, 3 (CDC31 homolog, yeast) | 6856 | 0.001 | -0.3727 | No |
| 26 | TPT1 | TPT1 Entrez, Source | tumor protein, translationally-controlled 1 | 6974 | -0.002 | -0.3815 | No |
| 27 | MEFV | MEFV Entrez, Source | Mediterranean fever | 7026 | -0.003 | -0.3852 | No |
| 28 | ALS2 | ALS2 Entrez, Source | amyotrophic lateral sclerosis 2 (juvenile) | 7100 | -0.004 | -0.3904 | No |
| 29 | MAP1A | MAP1A Entrez, Source | microtubule-associated protein 1A | 7727 | -0.017 | -0.4367 | No |
| 30 | KATNA1 | KATNA1 Entrez, Source | katanin p60 (ATPase-containing) subunit A 1 | 7746 | -0.018 | -0.4370 | No |
| 31 | MYH7 | MYH7 Entrez, Source | myosin, heavy chain 7, cardiac muscle, beta | 7898 | -0.021 | -0.4471 | No |
| 32 | STAU1 | STAU1 Entrez, Source | staufen, RNA binding protein, homolog 1 (Drosophila) | 8162 | -0.027 | -0.4653 | No |
| 33 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 8176 | -0.027 | -0.4646 | No |
| 34 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 8204 | -0.028 | -0.4649 | No |
| 35 | KIF4A | KIF4A Entrez, Source | kinesin family member 4A | 8274 | -0.029 | -0.4684 | No |
| 36 | ESPN | ESPN Entrez, Source | espin | 8501 | -0.034 | -0.4833 | No |
| 37 | MYOM1 | MYOM1 Entrez, Source | myomesin 1 (skelemin) 185kDa | 8685 | -0.039 | -0.4948 | No |
| 38 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 8842 | -0.042 | -0.5040 | No |
| 39 | GFAP | GFAP Entrez, Source | glial fibrillary acidic protein | 9126 | -0.049 | -0.5223 | No |
| 40 | ARPC5 | ARPC5 Entrez, Source | actin related protein 2/3 complex, subunit 5, 16kDa | 9496 | -0.060 | -0.5466 | No |
| 41 | ZW10 | ZW10 Entrez, Source | ZW10, kinetochore associated, homolog (Drosophila) | 9627 | -0.063 | -0.5525 | No |
| 42 | CABP1 | CABP1 Entrez, Source | calcium binding protein 1 (calbrain) | 9694 | -0.065 | -0.5535 | No |
| 43 | ACTC1 | ACTC1 Entrez, Source | actin, alpha, cardiac muscle 1 | 9699 | -0.065 | -0.5498 | No |
| 44 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 9827 | -0.069 | -0.5551 | No |
| 45 | CAPZA2 | CAPZA2 Entrez, Source | capping protein (actin filament) muscle Z-line, alpha 2 | 9837 | -0.069 | -0.5515 | No |
| 46 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 10026 | -0.075 | -0.5611 | No |
| 47 | MYO9B | MYO9B Entrez, Source | myosin IXB | 10160 | -0.080 | -0.5662 | No |
| 48 | ACTR2 | ACTR2 Entrez, Source | ARP2 actin-related protein 2 homolog (yeast) | 10226 | -0.082 | -0.5661 | No |
| 49 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10276 | -0.084 | -0.5646 | No |
| 50 | SIRT2 | SIRT2 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 2 (S. cerevisiae) | 10295 | -0.085 | -0.5608 | No |
| 51 | APC | APC Entrez, Source | adenomatosis polyposis coli | 10296 | -0.085 | -0.5556 | No |
| 52 | DCTN2 | DCTN2 Entrez, Source | dynactin 2 (p50) | 10303 | -0.085 | -0.5508 | No |
| 53 | ARPC1B | ARPC1B Entrez, Source | actin related protein 2/3 complex, subunit 1B, 41kDa | 10374 | -0.087 | -0.5508 | No |
| 54 | CETN1 | CETN1 Entrez, Source | centrin, EF-hand protein, 1 | 10599 | -0.095 | -0.5618 | No |
| 55 | KIF5A | KIF5A Entrez, Source | kinesin family member 5A | 10741 | -0.100 | -0.5663 | Yes |
| 56 | KATNB1 | KATNB1 Entrez, Source | katanin p80 (WD repeat containing) subunit B 1 | 10749 | -0.101 | -0.5607 | Yes |
| 57 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 10819 | -0.104 | -0.5595 | Yes |
| 58 | CAPZB | CAPZB Entrez, Source | capping protein (actin filament) muscle Z-line, beta | 10922 | -0.108 | -0.5606 | Yes |
| 59 | BMF | BMF Entrez, Source | Bcl2 modifying factor | 10943 | -0.109 | -0.5554 | Yes |
| 60 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 11099 | -0.117 | -0.5599 | Yes |
| 61 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 11105 | -0.117 | -0.5531 | Yes |
| 62 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 11117 | -0.118 | -0.5467 | Yes |
| 63 | AKAP9 | AKAP9 Entrez, Source | A kinase (PRKA) anchor protein (yotiao) 9 | 11273 | -0.125 | -0.5507 | Yes |
| 64 | POLB | POLB Entrez, Source | polymerase (DNA directed), beta | 11294 | -0.126 | -0.5445 | Yes |
| 65 | TBCE | TBCE Entrez, Source | tubulin-specific chaperone e | 11303 | -0.127 | -0.5373 | Yes |
| 66 | DYNC1LI2 | DYNC1LI2 Entrez, Source | dynein, cytoplasmic 1, light intermediate chain 2 | 11327 | -0.128 | -0.5312 | Yes |
| 67 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 11329 | -0.128 | -0.5234 | Yes |
| 68 | DCTN1 | DCTN1 Entrez, Source | dynactin 1 (p150, glued homolog, Drosophila) | 11343 | -0.128 | -0.5165 | Yes |
| 69 | CDC16 | CDC16 Entrez, Source | CDC16 cell division cycle 16 homolog (S. cerevisiae) | 11467 | -0.135 | -0.5175 | Yes |
| 70 | TUBGCP3 | TUBGCP3 Entrez, Source | tubulin, gamma complex associated protein 3 | 11484 | -0.136 | -0.5103 | Yes |
| 71 | TPM3 | TPM3 Entrez, Source | tropomyosin 3 | 11497 | -0.137 | -0.5028 | Yes |
| 72 | GAS8 | GAS8 Entrez, Source | growth arrest-specific 8 | 11506 | -0.138 | -0.4950 | Yes |
| 73 | SEPT7 | SEPT7 Entrez, Source | septin 7 | 11577 | -0.142 | -0.4915 | Yes |
| 74 | ACTR3 | ACTR3 Entrez, Source | ARP3 actin-related protein 3 homolog (yeast) | 11578 | -0.142 | -0.4828 | Yes |
| 75 | NDE1 | NDE1 Entrez, Source | nudE nuclear distribution gene E homolog 1 (A. nidulans) | 11603 | -0.144 | -0.4757 | Yes |
| 76 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 11608 | -0.144 | -0.4672 | Yes |
| 77 | SEPT9 | SEPT9 Entrez, Source | septin 9 | 11618 | -0.145 | -0.4590 | Yes |
| 78 | MYO1C | MYO1C Entrez, Source | myosin IC | 11628 | -0.145 | -0.4507 | Yes |
| 79 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 11658 | -0.148 | -0.4439 | Yes |
| 80 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 11695 | -0.149 | -0.4374 | Yes |
| 81 | DYNLL1 | DYNLL1 Entrez, Source | dynein, light chain, LC8-type 1 | 11702 | -0.150 | -0.4286 | Yes |
| 82 | KIF5B | KIF5B Entrez, Source | kinesin family member 5B | 11762 | -0.153 | -0.4237 | Yes |
| 83 | PXN | PXN Entrez, Source | paxillin | 11831 | -0.159 | -0.4190 | Yes |
| 84 | PCM1 | PCM1 Entrez, Source | pericentriolar material 1 | 11860 | -0.161 | -0.4113 | Yes |
| 85 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 11889 | -0.164 | -0.4033 | Yes |
| 86 | KIFC3 | KIFC3 Entrez, Source | kinesin family member C3 | 11973 | -0.171 | -0.3991 | Yes |
| 87 | CDK5RAP2 | CDK5RAP2 Entrez, Source | CDK5 regulatory subunit associated protein 2 | 11990 | -0.172 | -0.3897 | Yes |
| 88 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 12021 | -0.175 | -0.3813 | Yes |
| 89 | TSGA14 | TSGA14 Entrez, Source | testis specific, 14 | 12175 | -0.189 | -0.3812 | Yes |
| 90 | MAPRE1 | MAPRE1 Entrez, Source | microtubule-associated protein, RP/EB family, member 1 | 12234 | -0.197 | -0.3734 | Yes |
| 91 | HDAC6 | HDAC6 Entrez, Source | histone deacetylase 6 | 12329 | -0.208 | -0.3678 | Yes |
| 92 | LASP1 | LASP1 Entrez, Source | LIM and SH3 protein 1 | 12412 | -0.216 | -0.3607 | Yes |
| 93 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 12463 | -0.223 | -0.3508 | Yes |
| 94 | KIF3C | KIF3C Entrez, Source | kinesin family member 3C | 12481 | -0.225 | -0.3382 | Yes |
| 95 | DCTN4 | DCTN4 Entrez, Source | dynactin 4 (p62) | 12513 | -0.230 | -0.3264 | Yes |
| 96 | NUMA1 | NUMA1 Entrez, Source | nuclear mitotic apparatus protein 1 | 12531 | -0.233 | -0.3134 | Yes |
| 97 | CLASP2 | CLASP2 Entrez, Source | cytoplasmic linker associated protein 2 | 12533 | -0.233 | -0.2991 | Yes |
| 98 | AURKA | AURKA Entrez, Source | aurora kinase A | 12548 | -0.235 | -0.2857 | Yes |
| 99 | TPM4 | TPM4 Entrez, Source | tropomyosin 4 | 12551 | -0.235 | -0.2714 | Yes |
| 100 | MYH9 | MYH9 Entrez, Source | myosin, heavy chain 9, non-muscle | 12585 | -0.241 | -0.2591 | Yes |
| 101 | CKAP5 | CKAP5 Entrez, Source | cytoskeleton associated protein 5 | 12613 | -0.247 | -0.2459 | Yes |
| 102 | CLASP1 | CLASP1 Entrez, Source | cytoplasmic linker associated protein 1 | 12909 | -0.326 | -0.2482 | Yes |
| 103 | MAP1B | MAP1B Entrez, Source | microtubule-associated protein 1B | 12944 | -0.337 | -0.2301 | Yes |
| 104 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 12998 | -0.364 | -0.2117 | Yes |
| 105 | CAPG | CAPG Entrez, Source | capping protein (actin filament), gelsolin-like | 13018 | -0.375 | -0.1900 | Yes |
| 106 | CEP57 | CEP57 Entrez, Source | centrosomal protein 57kDa | 13119 | -0.442 | -0.1704 | Yes |
| 107 | VIM | VIM Entrez, Source | vimentin | 13164 | -0.501 | -0.1429 | Yes |
| 108 | NEFL | NEFL Entrez, Source | neurofilament, light polypeptide 68kDa | 13304 | -0.959 | -0.0944 | Yes |
| 109 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 13340 | -1.581 | 0.0002 | Yes |