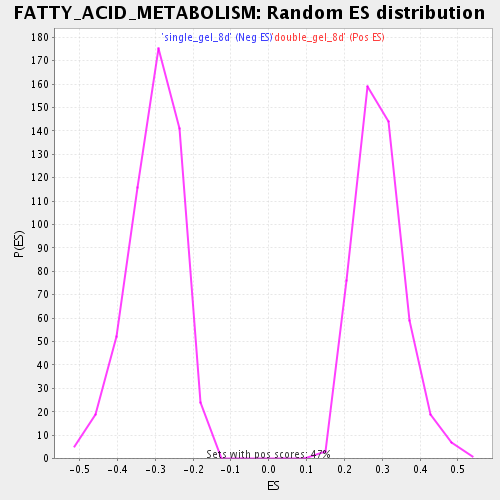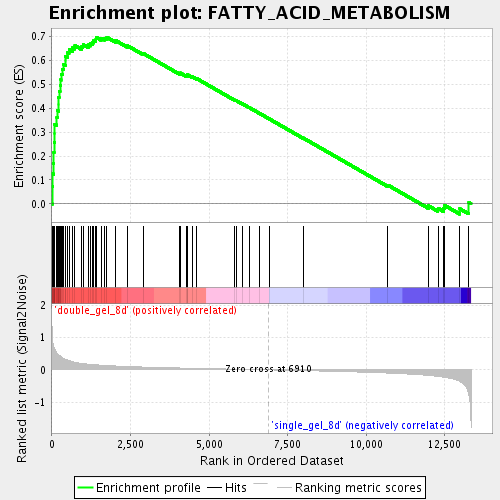
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | FATTY_ACID_METABOLISM |
| Enrichment Score (ES) | 0.6961539 |
| Normalized Enrichment Score (NES) | 2.3798714 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 8 | 1.205 | 0.0737 | Yes |
| 2 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 24 | 0.864 | 0.1259 | Yes |
| 3 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 50 | 0.751 | 0.1703 | Yes |
| 4 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 60 | 0.721 | 0.2141 | Yes |
| 5 | FABP2 | FABP2 Entrez, Source | fatty acid binding protein 2, intestinal | 71 | 0.690 | 0.2559 | Yes |
| 6 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 87 | 0.637 | 0.2940 | Yes |
| 7 | ADHFE1 | ADHFE1 Entrez, Source | alcohol dehydrogenase, iron containing, 1 | 89 | 0.633 | 0.3330 | Yes |
| 8 | CYP51A1 | CYP51A1 Entrez, Source | cytochrome P450, family 51, subfamily A, polypeptide 1 | 157 | 0.524 | 0.3602 | Yes |
| 9 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 172 | 0.499 | 0.3899 | Yes |
| 10 | FASN | FASN Entrez, Source | fatty acid synthase | 213 | 0.460 | 0.4153 | Yes |
| 11 | ADH1A | ADH1A Entrez, Source | alcohol dehydrogenase 1A (class I), alpha polypeptide | 214 | 0.460 | 0.4437 | Yes |
| 12 | LPL | LPL Entrez, Source | lipoprotein lipase | 230 | 0.451 | 0.4704 | Yes |
| 13 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 260 | 0.427 | 0.4946 | Yes |
| 14 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 284 | 0.409 | 0.5180 | Yes |
| 15 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 292 | 0.402 | 0.5423 | Yes |
| 16 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 335 | 0.375 | 0.5623 | Yes |
| 17 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 374 | 0.353 | 0.5812 | Yes |
| 18 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 429 | 0.329 | 0.5974 | Yes |
| 19 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 430 | 0.329 | 0.6177 | Yes |
| 20 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 501 | 0.299 | 0.6309 | Yes |
| 21 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 546 | 0.284 | 0.6451 | Yes |
| 22 | DCI | DCI Entrez, Source | dodecenoyl-Coenzyme A delta isomerase (3,2 trans-enoyl-Coenzyme A isomerase) | 660 | 0.250 | 0.6521 | Yes |
| 23 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 732 | 0.231 | 0.6610 | Yes |
| 24 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 950 | 0.200 | 0.6570 | Yes |
| 25 | PECR | PECR Entrez, Source | peroxisomal trans-2-enoyl-CoA reductase | 989 | 0.195 | 0.6661 | Yes |
| 26 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1158 | 0.175 | 0.6643 | Yes |
| 27 | SDS | SDS Entrez, Source | serine dehydratase | 1222 | 0.169 | 0.6700 | Yes |
| 28 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1296 | 0.164 | 0.6746 | Yes |
| 29 | CYP1A1 | CYP1A1 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 1 | 1331 | 0.162 | 0.6820 | Yes |
| 30 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1399 | 0.157 | 0.6867 | Yes |
| 31 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 1403 | 0.157 | 0.6961 | Yes |
| 32 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 1566 | 0.144 | 0.6928 | Yes |
| 33 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1680 | 0.138 | 0.6928 | Yes |
| 34 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 1747 | 0.134 | 0.6962 | Yes |
| 35 | ACADL | ACADL Entrez, Source | acyl-Coenzyme A dehydrogenase, long chain | 2024 | 0.121 | 0.6828 | No |
| 36 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 2402 | 0.107 | 0.6610 | No |
| 37 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 2909 | 0.088 | 0.6284 | No |
| 38 | LIPE | LIPE Entrez, Source | lipase, hormone-sensitive | 4076 | 0.058 | 0.5441 | No |
| 39 | CYP1A2 | CYP1A2 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 2 | 4085 | 0.058 | 0.5471 | No |
| 40 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 4291 | 0.053 | 0.5349 | No |
| 41 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 4305 | 0.053 | 0.5372 | No |
| 42 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4310 | 0.053 | 0.5401 | No |
| 43 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 4466 | 0.049 | 0.5315 | No |
| 44 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 4613 | 0.046 | 0.5233 | No |
| 45 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 5822 | 0.022 | 0.4338 | No |
| 46 | CYP3A4 | CYP3A4 Entrez, Source | cytochrome P450, family 3, subfamily A, polypeptide 4 | 5860 | 0.021 | 0.4323 | No |
| 47 | CYP4B1 | CYP4B1 Entrez, Source | cytochrome P450, family 4, subfamily B, polypeptide 1 | 6055 | 0.017 | 0.4187 | No |
| 48 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6279 | 0.012 | 0.4027 | No |
| 49 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 6602 | 0.006 | 0.3788 | No |
| 50 | CYP2C9 | CYP2C9 Entrez, Source | cytochrome P450, family 2, subfamily C, polypeptide 9 | 6932 | -0.001 | 0.3541 | No |
| 51 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 7996 | -0.023 | 0.2755 | No |
| 52 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 10696 | -0.098 | 0.0784 | No |
| 53 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 11983 | -0.171 | -0.0078 | No |
| 54 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 12291 | -0.203 | -0.0184 | No |
| 55 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 12469 | -0.223 | -0.0179 | No |
| 56 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 12503 | -0.229 | -0.0063 | No |
| 57 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 12985 | -0.357 | -0.0205 | No |
| 58 | FABP7 | FABP7 Entrez, Source | fatty acid binding protein 7, brain | 13272 | -0.767 | 0.0053 | No |