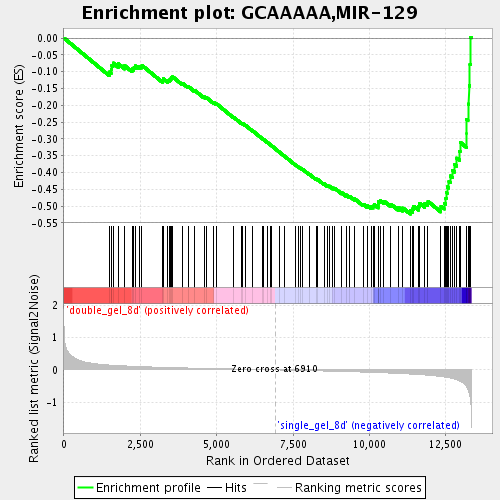
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | GCAAAAA,MIR-129 |
| Enrichment Score (ES) | -0.5238789 |
| Normalized Enrichment Score (NES) | -1.8831345 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02531469 |
| FWER p-Value | 0.548 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PAPOLB | PAPOLB Entrez, Source | poly(A) polymerase beta (testis specific) | 1497 | 0.150 | -0.1009 | No |
| 2 | GRM7 | GRM7 Entrez, Source | glutamate receptor, metabotropic 7 | 1547 | 0.146 | -0.0929 | No |
| 3 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 1566 | 0.144 | -0.0826 | No |
| 4 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 1608 | 0.143 | -0.0742 | No |
| 5 | XKR4 | XKR4 Entrez, Source | XK, Kell blood group complex subunit-related family, member 4 | 1778 | 0.132 | -0.0763 | No |
| 6 | HERC4 | HERC4 Entrez, Source | hect domain and RLD 4 | 1991 | 0.122 | -0.0825 | No |
| 7 | NR2C2 | NR2C2 Entrez, Source | nuclear receptor subfamily 2, group C, member 2 | 2232 | 0.113 | -0.0915 | No |
| 8 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 2292 | 0.111 | -0.0870 | No |
| 9 | TAOK3 | TAOK3 Entrez, Source | TAO kinase 3 | 2351 | 0.108 | -0.0827 | No |
| 10 | TMEM23 | TMEM23 Entrez, Source | transmembrane protein 23 | 2473 | 0.104 | -0.0835 | No |
| 11 | NRXN2 | NRXN2 Entrez, Source | neurexin 2 | 2553 | 0.101 | -0.0813 | No |
| 12 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 3222 | 0.078 | -0.1254 | No |
| 13 | DAB2IP | DAB2IP Entrez, Source | DAB2 interacting protein | 3243 | 0.077 | -0.1207 | No |
| 14 | CACNG2 | CACNG2 Entrez, Source | calcium channel, voltage-dependent, gamma subunit 2 | 3394 | 0.074 | -0.1261 | No |
| 15 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 3452 | 0.072 | -0.1246 | No |
| 16 | SCN8A | SCN8A Entrez, Source | sodium channel, voltage gated, type VIII, alpha | 3492 | 0.071 | -0.1218 | No |
| 17 | STOX2 | STOX2 Entrez, Source | storkhead box 2 | 3509 | 0.071 | -0.1173 | No |
| 18 | OXR1 | OXR1 Entrez, Source | oxidation resistance 1 | 3557 | 0.069 | -0.1153 | No |
| 19 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 3876 | 0.062 | -0.1342 | No |
| 20 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 4062 | 0.058 | -0.1435 | No |
| 21 | PITPNB | PITPNB Entrez, Source | phosphatidylinositol transfer protein, beta | 4286 | 0.053 | -0.1561 | No |
| 22 | SLC17A6 | SLC17A6 Entrez, Source | solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 6 | 4586 | 0.047 | -0.1749 | No |
| 23 | SP3 | SP3 Entrez, Source | Sp3 transcription factor | 4669 | 0.045 | -0.1775 | No |
| 24 | DLGAP2 | DLGAP2 Entrez, Source | discs, large (Drosophila) homolog-associated protein 2 | 4904 | 0.040 | -0.1919 | No |
| 25 | TBC1D15 | TBC1D15 Entrez, Source | TBC1 domain family, member 15 | 4997 | 0.038 | -0.1958 | No |
| 26 | PTP4A1 | PTP4A1 Entrez, Source | protein tyrosine phosphatase type IVA, member 1 | 5554 | 0.026 | -0.2356 | No |
| 27 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 5811 | 0.022 | -0.2532 | No |
| 28 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 5852 | 0.021 | -0.2545 | No |
| 29 | SH3KBP1 | SH3KBP1 Entrez, Source | SH3-domain kinase binding protein 1 | 5941 | 0.019 | -0.2596 | No |
| 30 | SLC25A27 | SLC25A27 Entrez, Source | solute carrier family 25, member 27 | 6177 | 0.015 | -0.2761 | No |
| 31 | GALNT1 | GALNT1 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 1 (GalNAc-T1) | 6485 | 0.008 | -0.2987 | No |
| 32 | SBK1 | SBK1 Entrez, Source | SH3-binding domain kinase 1 | 6546 | 0.007 | -0.3026 | No |
| 33 | EPHA8 | EPHA8 Entrez, Source | EPH receptor A8 | 6653 | 0.005 | -0.3102 | No |
| 34 | PSMA7 | PSMA7 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 7 | 6760 | 0.003 | -0.3180 | No |
| 35 | ABCC5 | ABCC5 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 5 | 6790 | 0.003 | -0.3200 | No |
| 36 | NRG2 | NRG2 Entrez, Source | neuregulin 2 | 7064 | -0.003 | -0.3403 | No |
| 37 | ARL5A | ARL5A Entrez, Source | ADP-ribosylation factor-like 5A | 7225 | -0.007 | -0.3518 | No |
| 38 | LPHN3 | LPHN3 Entrez, Source | latrophilin 3 | 7585 | -0.014 | -0.3778 | No |
| 39 | SFMBT1 | SFMBT1 Entrez, Source | Scm-like with four mbt domains 1 | 7686 | -0.016 | -0.3840 | No |
| 40 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 7740 | -0.017 | -0.3866 | No |
| 41 | ATP2B4 | ATP2B4 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 4 | 7808 | -0.019 | -0.3901 | No |
| 42 | FNDC5 | FNDC5 Entrez, Source | fibronectin type III domain containing 5 | 8027 | -0.024 | -0.4047 | No |
| 43 | NIPBL | NIPBL Entrez, Source | Nipped-B homolog (Drosophila) | 8250 | -0.029 | -0.4191 | No |
| 44 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 8301 | -0.030 | -0.4205 | No |
| 45 | GFRA2 | GFRA2 Entrez, Source | GDNF family receptor alpha 2 | 8512 | -0.035 | -0.4335 | No |
| 46 | SYT1 | SYT1 Entrez, Source | synaptotagmin I | 8610 | -0.037 | -0.4379 | No |
| 47 | MYT1L | MYT1L Entrez, Source | myelin transcription factor 1-like | 8684 | -0.039 | -0.4403 | No |
| 48 | NPEPPS | NPEPPS Entrez, Source | aminopeptidase puromycin sensitive | 8805 | -0.041 | -0.4460 | No |
| 49 | STAT5B | STAT5B Entrez, Source | signal transducer and activator of transcription 5B | 8858 | -0.043 | -0.4465 | No |
| 50 | GNAZ | GNAZ Entrez, Source | guanine nucleotide binding protein (G protein), alpha z polypeptide | 9092 | -0.048 | -0.4602 | No |
| 51 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 9236 | -0.052 | -0.4668 | No |
| 52 | HOXC10 | HOXC10 Entrez, Source | homeobox C10 | 9351 | -0.056 | -0.4709 | No |
| 53 | CBX7 | CBX7 Entrez, Source | chromobox homolog 7 | 9510 | -0.060 | -0.4780 | No |
| 54 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 9794 | -0.068 | -0.4939 | No |
| 55 | ARNT | ARNT Entrez, Source | aryl hydrocarbon receptor nuclear translocator | 9940 | -0.073 | -0.4990 | No |
| 56 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 10052 | -0.076 | -0.5012 | No |
| 57 | PITPNA | PITPNA Entrez, Source | phosphatidylinositol transfer protein, alpha | 10116 | -0.079 | -0.4996 | No |
| 58 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 10149 | -0.080 | -0.4956 | No |
| 59 | NR3C2 | NR3C2 Entrez, Source | nuclear receptor subfamily 3, group C, member 2 | 10284 | -0.084 | -0.4990 | No |
| 60 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 10287 | -0.084 | -0.4924 | No |
| 61 | APC | APC Entrez, Source | adenomatosis polyposis coli | 10296 | -0.085 | -0.4861 | No |
| 62 | VCP | VCP Entrez, Source | valosin-containing protein | 10359 | -0.087 | -0.4838 | No |
| 63 | VPS26A | VPS26A Entrez, Source | vacuolar protein sorting 26 homolog A (yeast) | 10474 | -0.091 | -0.4851 | No |
| 64 | HNRPK | HNRPK Entrez, Source | heterogeneous nuclear ribonucleoprotein K | 10700 | -0.098 | -0.4942 | No |
| 65 | SP4 | SP4 Entrez, Source | Sp4 transcription factor | 10950 | -0.109 | -0.5042 | No |
| 66 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 11094 | -0.116 | -0.5056 | No |
| 67 | CIB2 | CIB2 Entrez, Source | calcium and integrin binding family member 2 | 11337 | -0.128 | -0.5136 | Yes |
| 68 | RAB21 | RAB21 Entrez, Source | RAB21, member RAS oncogene family | 11411 | -0.132 | -0.5085 | Yes |
| 69 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 11444 | -0.134 | -0.5001 | Yes |
| 70 | MATR3 | MATR3 Entrez, Source | matrin 3 | 11605 | -0.144 | -0.5006 | Yes |
| 71 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 11635 | -0.146 | -0.4910 | Yes |
| 72 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 11811 | -0.158 | -0.4916 | Yes |
| 73 | KCTD1 | KCTD1 Entrez, Source | potassium channel tetramerisation domain containing 1 | 11913 | -0.167 | -0.4858 | Yes |
| 74 | LMNA | LMNA Entrez, Source | lamin A/C | 12337 | -0.209 | -0.5009 | Yes |
| 75 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 12443 | -0.220 | -0.4911 | Yes |
| 76 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 12496 | -0.228 | -0.4767 | Yes |
| 77 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 12514 | -0.231 | -0.4594 | Yes |
| 78 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 12545 | -0.235 | -0.4428 | Yes |
| 79 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 12599 | -0.245 | -0.4271 | Yes |
| 80 | BTBD10 | BTBD10 Entrez, Source | BTB (POZ) domain containing 10 | 12646 | -0.253 | -0.4102 | Yes |
| 81 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 12719 | -0.269 | -0.3940 | Yes |
| 82 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12786 | -0.282 | -0.3762 | Yes |
| 83 | ZBTB10 | ZBTB10 Entrez, Source | zinc finger and BTB domain containing 10 | 12857 | -0.306 | -0.3569 | Yes |
| 84 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 12949 | -0.338 | -0.3366 | Yes |
| 85 | PLCG1 | PLCG1 Entrez, Source | phospholipase C, gamma 1 | 12969 | -0.347 | -0.3101 | Yes |
| 86 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 13173 | -0.513 | -0.2841 | Yes |
| 87 | PTPRZ1 | PTPRZ1 Entrez, Source | protein tyrosine phosphatase, receptor-type, Z polypeptide 1 | 13176 | -0.521 | -0.2423 | Yes |
| 88 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 13226 | -0.619 | -0.1962 | Yes |
| 89 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 13257 | -0.714 | -0.1411 | Yes |
| 90 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 13287 | -0.812 | -0.0779 | Yes |
| 91 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 13312 | -1.018 | 0.0023 | Yes |

