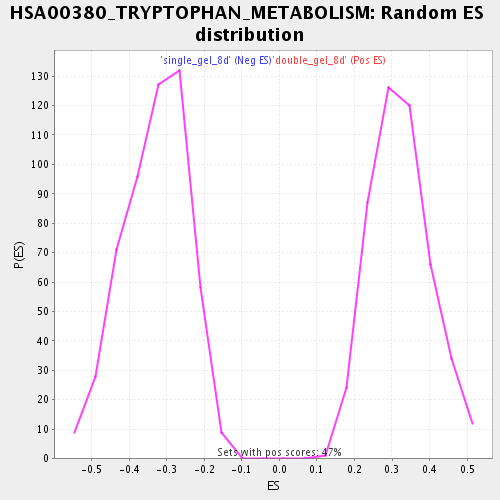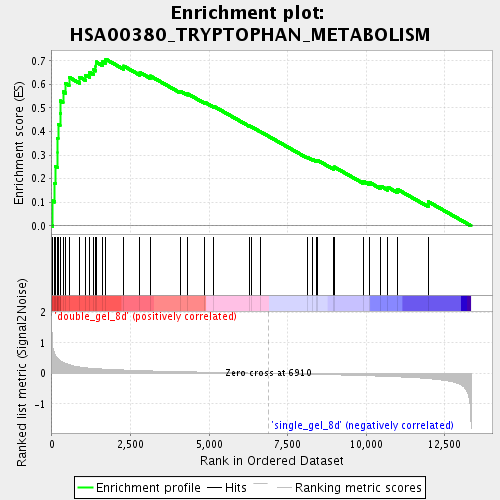
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HSA00380_TRYPTOPHAN_METABOLISM |
| Enrichment Score (ES) | 0.70711184 |
| Normalized Enrichment Score (NES) | 2.1936076 |
| Nominal p-value | 0.0 |
| FDR q-value | 7.3575684E-5 |
| FWER p-Value | 0.0030 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 24 | 0.864 | 0.1056 | Yes |
| 2 | AADAT | AADAT Entrez, Source | aminoadipate aminotransferase | 88 | 0.633 | 0.1795 | Yes |
| 3 | ABP1 | ABP1 Entrez, Source | amiloride binding protein 1 (amine oxidase (copper-containing)) | 109 | 0.606 | 0.2533 | Yes |
| 4 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 172 | 0.499 | 0.3106 | Yes |
| 5 | KMO | KMO Entrez, Source | kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) | 174 | 0.497 | 0.3723 | Yes |
| 6 | HAAO | HAAO Entrez, Source | 3-hydroxyanthranilate 3,4-dioxygenase | 196 | 0.479 | 0.4303 | Yes |
| 7 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 260 | 0.427 | 0.4786 | Yes |
| 8 | ACMSD | ACMSD Entrez, Source | aminocarboxymuconate semialdehyde decarboxylase | 269 | 0.420 | 0.5301 | Yes |
| 9 | TDO2 | TDO2 Entrez, Source | tryptophan 2,3-dioxygenase | 355 | 0.360 | 0.5685 | Yes |
| 10 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 430 | 0.329 | 0.6038 | Yes |
| 11 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 548 | 0.283 | 0.6301 | Yes |
| 12 | CAT | CAT Entrez, Source | catalase | 884 | 0.209 | 0.6310 | Yes |
| 13 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 1074 | 0.185 | 0.6397 | Yes |
| 14 | AANAT | AANAT Entrez, Source | arylalkylamine N-acetyltransferase | 1198 | 0.171 | 0.6517 | Yes |
| 15 | CYP1A1 | CYP1A1 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 1 | 1331 | 0.162 | 0.6619 | Yes |
| 16 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1399 | 0.157 | 0.6763 | Yes |
| 17 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 1403 | 0.157 | 0.6956 | Yes |
| 18 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1615 | 0.142 | 0.6974 | Yes |
| 19 | MAOB | MAOB Entrez, Source | monoamine oxidase B | 1712 | 0.137 | 0.7071 | Yes |
| 20 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 2281 | 0.111 | 0.6782 | No |
| 21 | MAOA | MAOA Entrez, Source | monoamine oxidase A | 2802 | 0.091 | 0.6505 | No |
| 22 | DDC | DDC Entrez, Source | dopa decarboxylase (aromatic L-amino acid decarboxylase) | 3130 | 0.081 | 0.6359 | No |
| 23 | CYP1A2 | CYP1A2 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 2 | 4085 | 0.058 | 0.5713 | No |
| 24 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 4305 | 0.053 | 0.5614 | No |
| 25 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 4870 | 0.040 | 0.5240 | No |
| 26 | NFX1 | NFX1 Entrez, Source | nuclear transcription factor, X-box binding 1 | 5152 | 0.035 | 0.5073 | No |
| 27 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6279 | 0.012 | 0.4241 | No |
| 28 | PRMT3 | PRMT3 Entrez, Source | protein arginine methyltransferase 3 | 6340 | 0.011 | 0.4210 | No |
| 29 | TPH1 | TPH1 Entrez, Source | tryptophan hydroxylase 1 (tryptophan 5-monooxygenase) | 6630 | 0.005 | 0.3999 | No |
| 30 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 8124 | -0.026 | 0.2909 | No |
| 31 | CARM1 | CARM1 Entrez, Source | coactivator-associated arginine methyltransferase 1 | 8288 | -0.030 | 0.2823 | No |
| 32 | METTL6 | METTL6 Entrez, Source | methyltransferase like 6 | 8429 | -0.033 | 0.2759 | No |
| 33 | AOX1 | AOX1 Entrez, Source | aldehyde oxidase 1 | 8449 | -0.033 | 0.2786 | No |
| 34 | LCMT2 | LCMT2 Entrez, Source | leucine carboxyl methyltransferase 2 | 8973 | -0.045 | 0.2449 | No |
| 35 | ASMT | ASMT Entrez, Source | acetylserotonin O-methyltransferase | 8983 | -0.045 | 0.2499 | No |
| 36 | PRMT2 | PRMT2 Entrez, Source | protein arginine methyltransferase 2 | 9909 | -0.072 | 0.1893 | No |
| 37 | LCMT1 | LCMT1 Entrez, Source | leucine carboxyl methyltransferase 1 | 10118 | -0.079 | 0.1834 | No |
| 38 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10462 | -0.090 | 0.1689 | No |
| 39 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 10696 | -0.098 | 0.1636 | No |
| 40 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 11000 | -0.112 | 0.1547 | No |
| 41 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 11983 | -0.171 | 0.1022 | No |