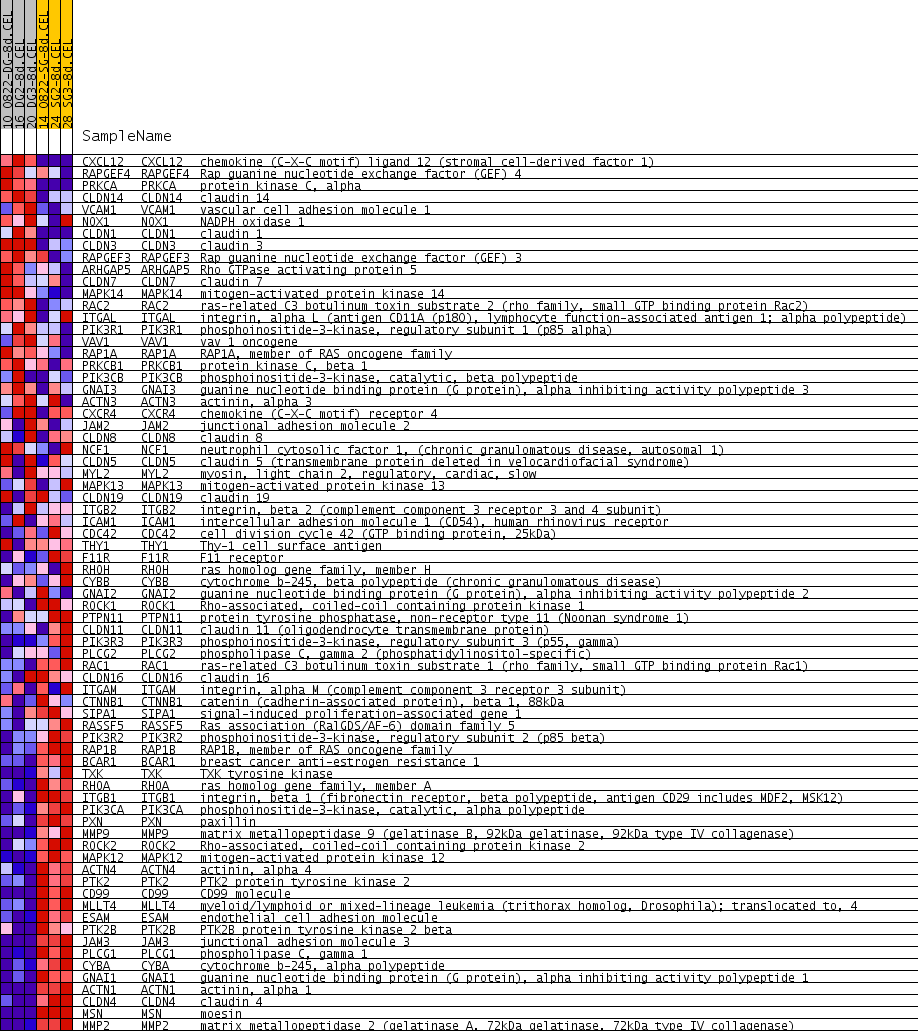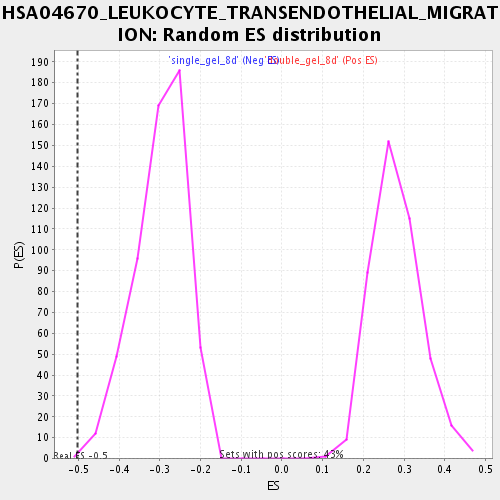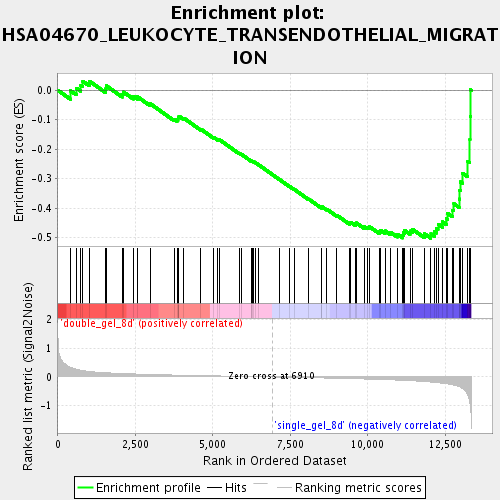
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | HSA04670_LEUKOCYTE_TRANSENDOTHELIAL_MIGRATION |
| Enrichment Score (ES) | -0.50308126 |
| Normalized Enrichment Score (NES) | -1.6895055 |
| Nominal p-value | 0.0017667845 |
| FDR q-value | 0.07812546 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CXCL12 | CXCL12 Entrez, Source | chemokine (C-X-C motif) ligand 12 (stromal cell-derived factor 1) | 407 | 0.337 | -0.0016 | No |
| 2 | RAPGEF4 | RAPGEF4 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 4 | 597 | 0.268 | 0.0073 | No |
| 3 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 736 | 0.230 | 0.0168 | No |
| 4 | CLDN14 | CLDN14 Entrez, Source | claudin 14 | 802 | 0.220 | 0.0308 | No |
| 5 | VCAM1 | VCAM1 Entrez, Source | vascular cell adhesion molecule 1 | 1016 | 0.190 | 0.0312 | No |
| 6 | NOX1 | NOX1 Entrez, Source | NADPH oxidase 1 | 1538 | 0.147 | 0.0046 | No |
| 7 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 1554 | 0.145 | 0.0160 | No |
| 8 | CLDN3 | CLDN3 Entrez, Source | claudin 3 | 2073 | 0.118 | -0.0128 | No |
| 9 | RAPGEF3 | RAPGEF3 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 3 | 2114 | 0.117 | -0.0057 | No |
| 10 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 2435 | 0.105 | -0.0207 | No |
| 11 | CLDN7 | CLDN7 Entrez, Source | claudin 7 | 2563 | 0.101 | -0.0216 | No |
| 12 | MAPK14 | MAPK14 Entrez, Source | mitogen-activated protein kinase 14 | 2979 | 0.086 | -0.0455 | No |
| 13 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 3759 | 0.065 | -0.0986 | No |
| 14 | ITGAL | ITGAL Entrez, Source | integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) | 3846 | 0.063 | -0.0996 | No |
| 15 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 3876 | 0.062 | -0.0964 | No |
| 16 | VAV1 | VAV1 Entrez, Source | vav 1 oncogene | 3880 | 0.062 | -0.0912 | No |
| 17 | RAP1A | RAP1A Entrez, Source | RAP1A, member of RAS oncogene family | 3905 | 0.062 | -0.0877 | No |
| 18 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 4062 | 0.058 | -0.0944 | No |
| 19 | PIK3CB | PIK3CB Entrez, Source | phosphoinositide-3-kinase, catalytic, beta polypeptide | 4611 | 0.046 | -0.1317 | No |
| 20 | GNAI3 | GNAI3 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 3 | 5025 | 0.037 | -0.1596 | No |
| 21 | ACTN3 | ACTN3 Entrez, Source | actinin, alpha 3 | 5159 | 0.035 | -0.1666 | No |
| 22 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 5204 | 0.034 | -0.1670 | No |
| 23 | JAM2 | JAM2 Entrez, Source | junctional adhesion molecule 2 | 5847 | 0.021 | -0.2136 | No |
| 24 | CLDN8 | CLDN8 Entrez, Source | claudin 8 | 5910 | 0.020 | -0.2165 | No |
| 25 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 6242 | 0.013 | -0.2403 | No |
| 26 | CLDN5 | CLDN5 Entrez, Source | claudin 5 (transmembrane protein deleted in velocardiofacial syndrome) | 6250 | 0.013 | -0.2397 | No |
| 27 | MYL2 | MYL2 Entrez, Source | myosin, light chain 2, regulatory, cardiac, slow | 6293 | 0.012 | -0.2419 | No |
| 28 | MAPK13 | MAPK13 Entrez, Source | mitogen-activated protein kinase 13 | 6324 | 0.011 | -0.2431 | No |
| 29 | CLDN19 | CLDN19 Entrez, Source | claudin 19 | 6363 | 0.010 | -0.2451 | No |
| 30 | ITGB2 | ITGB2 Entrez, Source | integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) | 6482 | 0.008 | -0.2533 | No |
| 31 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 7160 | -0.005 | -0.3038 | No |
| 32 | CDC42 | CDC42 Entrez, Source | cell division cycle 42 (GTP binding protein, 25kDa) | 7461 | -0.012 | -0.3255 | No |
| 33 | THY1 | THY1 Entrez, Source | Thy-1 cell surface antigen | 7638 | -0.015 | -0.3374 | No |
| 34 | F11R | F11R Entrez, Source | F11 receptor | 8090 | -0.025 | -0.3692 | No |
| 35 | RHOH | RHOH Entrez, Source | ras homolog gene family, member H | 8506 | -0.035 | -0.3975 | No |
| 36 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 8513 | -0.035 | -0.3949 | No |
| 37 | GNAI2 | GNAI2 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 2 | 8673 | -0.038 | -0.4036 | No |
| 38 | ROCK1 | ROCK1 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 1 | 9002 | -0.046 | -0.4244 | No |
| 39 | PTPN11 | PTPN11 Entrez, Source | protein tyrosine phosphatase, non-receptor type 11 (Noonan syndrome 1) | 9419 | -0.058 | -0.4507 | No |
| 40 | CLDN11 | CLDN11 Entrez, Source | claudin 11 (oligodendrocyte transmembrane protein) | 9454 | -0.058 | -0.4483 | No |
| 41 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 9602 | -0.063 | -0.4539 | No |
| 42 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 9629 | -0.063 | -0.4504 | No |
| 43 | RAC1 | RAC1 Entrez, Source | ras-related C3 botulinum toxin substrate 1 (rho family, small GTP binding protein Rac1) | 9885 | -0.071 | -0.4635 | No |
| 44 | CLDN16 | CLDN16 Entrez, Source | claudin 16 | 9988 | -0.074 | -0.4648 | No |
| 45 | ITGAM | ITGAM Entrez, Source | integrin, alpha M (complement component 3 receptor 3 subunit) | 10046 | -0.076 | -0.4625 | No |
| 46 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 10376 | -0.087 | -0.4797 | No |
| 47 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 10416 | -0.089 | -0.4750 | No |
| 48 | RASSF5 | RASSF5 Entrez, Source | Ras association (RalGDS/AF-6) domain family 5 | 10560 | -0.094 | -0.4777 | No |
| 49 | PIK3R2 | PIK3R2 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 2 (p85 beta) | 10731 | -0.100 | -0.4819 | No |
| 50 | RAP1B | RAP1B Entrez, Source | RAP1B, member of RAS oncogene family | 10952 | -0.109 | -0.4890 | No |
| 51 | BCAR1 | BCAR1 Entrez, Source | breast cancer anti-estrogen resistance 1 | 11123 | -0.118 | -0.4916 | Yes |
| 52 | TXK | TXK Entrez, Source | TXK tyrosine kinase | 11150 | -0.119 | -0.4833 | Yes |
| 53 | RHOA | RHOA Entrez, Source | ras homolog gene family, member A | 11190 | -0.121 | -0.4758 | Yes |
| 54 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 11366 | -0.129 | -0.4778 | Yes |
| 55 | PIK3CA | PIK3CA Entrez, Source | phosphoinositide-3-kinase, catalytic, alpha polypeptide | 11438 | -0.133 | -0.4717 | Yes |
| 56 | PXN | PXN Entrez, Source | paxillin | 11831 | -0.159 | -0.4875 | Yes |
| 57 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12039 | -0.176 | -0.4878 | Yes |
| 58 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 12156 | -0.187 | -0.4804 | Yes |
| 59 | MAPK12 | MAPK12 Entrez, Source | mitogen-activated protein kinase 12 | 12219 | -0.194 | -0.4684 | Yes |
| 60 | ACTN4 | ACTN4 Entrez, Source | actinin, alpha 4 | 12269 | -0.201 | -0.4547 | Yes |
| 61 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 12413 | -0.216 | -0.4468 | Yes |
| 62 | CD99 | CD99 Entrez, Source | CD99 molecule | 12537 | -0.234 | -0.4359 | Yes |
| 63 | MLLT4 | MLLT4 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 4 | 12569 | -0.239 | -0.4176 | Yes |
| 64 | ESAM | ESAM Entrez, Source | endothelial cell adhesion molecule | 12740 | -0.273 | -0.4068 | Yes |
| 65 | PTK2B | PTK2B Entrez, Source | PTK2B protein tyrosine kinase 2 beta | 12778 | -0.280 | -0.3855 | Yes |
| 66 | JAM3 | JAM3 Entrez, Source | junctional adhesion molecule 3 | 12952 | -0.339 | -0.3692 | Yes |
| 67 | PLCG1 | PLCG1 Entrez, Source | phospholipase C, gamma 1 | 12969 | -0.347 | -0.3405 | Yes |
| 68 | CYBA | CYBA Entrez, Source | cytochrome b-245, alpha polypeptide | 12989 | -0.359 | -0.3110 | Yes |
| 69 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13055 | -0.401 | -0.2812 | Yes |
| 70 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 13226 | -0.619 | -0.2406 | Yes |
| 71 | CLDN4 | CLDN4 Entrez, Source | claudin 4 | 13296 | -0.895 | -0.1685 | Yes |
| 72 | MSN | MSN Entrez, Source | moesin | 13302 | -0.941 | -0.0876 | Yes |
| 73 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 13316 | -1.048 | 0.0020 | Yes |