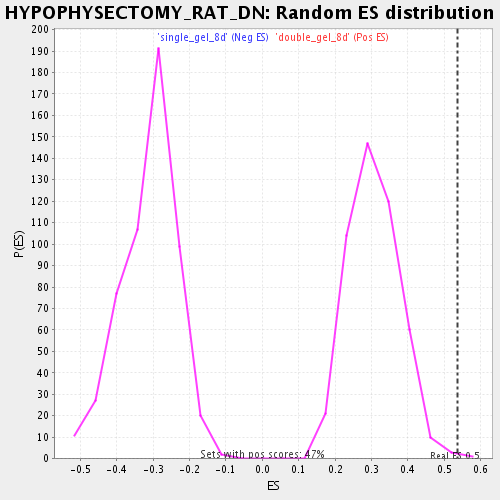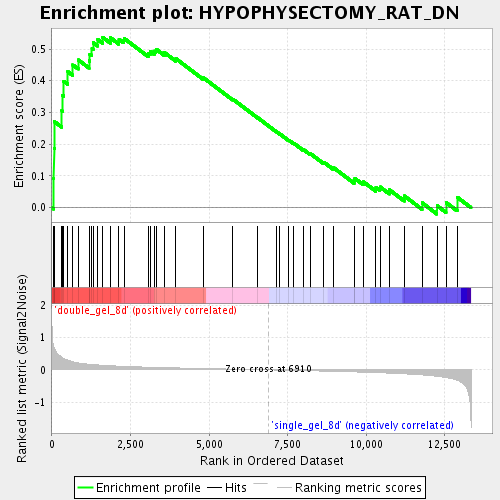
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HYPOPHYSECTOMY_RAT_DN |
| Enrichment Score (ES) | 0.5370138 |
| Normalized Enrichment Score (NES) | 1.7507924 |
| Nominal p-value | 0.0021459227 |
| FDR q-value | 0.021102576 |
| FWER p-Value | 0.979 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PDK4 | PDK4 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 4 | 58 | 0.726 | 0.0928 | Yes |
| 2 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 66 | 0.702 | 0.1863 | Yes |
| 3 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 84 | 0.641 | 0.2709 | Yes |
| 4 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 315 | 0.386 | 0.3052 | Yes |
| 5 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 335 | 0.375 | 0.3540 | Yes |
| 6 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 374 | 0.353 | 0.3984 | Yes |
| 7 | CYB5B | CYB5B Entrez, Source | cytochrome b5 type B (outer mitochondrial membrane) | 505 | 0.298 | 0.4285 | Yes |
| 8 | ALDOB | ALDOB Entrez, Source | aldolase B, fructose-bisphosphate | 666 | 0.249 | 0.4498 | Yes |
| 9 | ALAS1 | ALAS1 Entrez, Source | aminolevulinate, delta-, synthase 1 | 839 | 0.215 | 0.4657 | Yes |
| 10 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 1183 | 0.172 | 0.4629 | Yes |
| 11 | ODC1 | ODC1 Entrez, Source | ornithine decarboxylase 1 | 1205 | 0.170 | 0.4841 | Yes |
| 12 | AQP8 | AQP8 Entrez, Source | aquaporin 8 | 1275 | 0.166 | 0.5011 | Yes |
| 13 | DECR1 | DECR1 Entrez, Source | 2,4-dienoyl CoA reductase 1, mitochondrial | 1315 | 0.163 | 0.5200 | Yes |
| 14 | OAT | OAT Entrez, Source | ornithine aminotransferase (gyrate atrophy) | 1460 | 0.152 | 0.5296 | Yes |
| 15 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1615 | 0.142 | 0.5370 | Yes |
| 16 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 1864 | 0.128 | 0.5355 | No |
| 17 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 2134 | 0.117 | 0.5309 | No |
| 18 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 2294 | 0.111 | 0.5337 | No |
| 19 | PEX19 | PEX19 Entrez, Source | peroxisomal biogenesis factor 19 | 3069 | 0.083 | 0.4866 | No |
| 20 | DDC | DDC Entrez, Source | dopa decarboxylase (aromatic L-amino acid decarboxylase) | 3130 | 0.081 | 0.4929 | No |
| 21 | PADI4 | PADI4 Entrez, Source | peptidyl arginine deiminase, type IV | 3257 | 0.077 | 0.4937 | No |
| 22 | MEP1B | MEP1B Entrez, Source | meprin A, beta | 3313 | 0.076 | 0.4997 | No |
| 23 | NPPA | NPPA Entrez, Source | natriuretic peptide precursor A | 3581 | 0.069 | 0.4889 | No |
| 24 | PIM3 | PIM3 Entrez, Source | pim-3 oncogene | 3946 | 0.061 | 0.4697 | No |
| 25 | DAD1 | DAD1 Entrez, Source | defender against cell death 1 | 4820 | 0.041 | 0.4096 | No |
| 26 | RAB3B | RAB3B Entrez, Source | RAB3B, member RAS oncogene family | 5758 | 0.023 | 0.3422 | No |
| 27 | CALR | CALR Entrez, Source | calreticulin | 6558 | 0.007 | 0.2830 | No |
| 28 | FTH1 | FTH1 Entrez, Source | ferritin, heavy polypeptide 1 | 7147 | -0.005 | 0.2394 | No |
| 29 | AHSG | AHSG Entrez, Source | alpha-2-HS-glycoprotein | 7259 | -0.008 | 0.2321 | No |
| 30 | TKT | TKT Entrez, Source | transketolase (Wernicke-Korsakoff syndrome) | 7534 | -0.013 | 0.2133 | No |
| 31 | FBLN5 | FBLN5 Entrez, Source | fibulin 5 | 7698 | -0.017 | 0.2032 | No |
| 32 | ARF4 | ARF4 Entrez, Source | ADP-ribosylation factor 4 | 7998 | -0.023 | 0.1839 | No |
| 33 | DCN | DCN Entrez, Source | decorin | 8220 | -0.028 | 0.1710 | No |
| 34 | B2M | B2M Entrez, Source | beta-2-microglobulin | 8650 | -0.038 | 0.1438 | No |
| 35 | TFF3 | TFF3 Entrez, Source | trefoil factor 3 (intestinal) | 8966 | -0.045 | 0.1262 | No |
| 36 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 9620 | -0.063 | 0.0855 | No |
| 37 | MGP | MGP Entrez, Source | matrix Gla protein | 9643 | -0.064 | 0.0924 | No |
| 38 | FN1 | FN1 Entrez, Source | fibronectin 1 | 9917 | -0.072 | 0.0816 | No |
| 39 | SLC9A1 | SLC9A1 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 1 (antiporter, Na+/H+, amiloride sensitive) | 10314 | -0.085 | 0.0632 | No |
| 40 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 10444 | -0.090 | 0.0655 | No |
| 41 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 10743 | -0.100 | 0.0565 | No |
| 42 | PDIA3 | PDIA3 Entrez, Source | protein disulfide isomerase family A, member 3 | 11217 | -0.122 | 0.0373 | No |
| 43 | PSMB8 | PSMB8 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7) | 11784 | -0.155 | 0.0155 | No |
| 44 | LIMS2 | LIMS2 Entrez, Source | LIM and senescent cell antigen-like domains 2 | 12259 | -0.199 | 0.0066 | No |
| 45 | PPGB | PPGB Entrez, Source | protective protein for beta-galactosidase (galactosialidosis) | 12547 | -0.235 | 0.0165 | No |
| 46 | COL3A1 | COL3A1 Entrez, Source | collagen, type III, alpha 1 (Ehlers-Danlos syndrome type IV, autosomal dominant) | 12902 | -0.323 | 0.0331 | No |