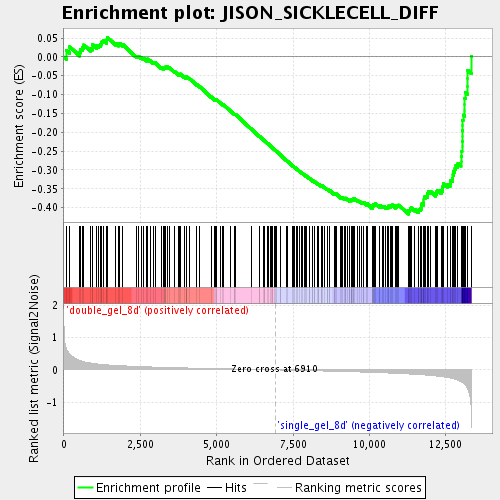
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | JISON_SICKLECELL_DIFF |
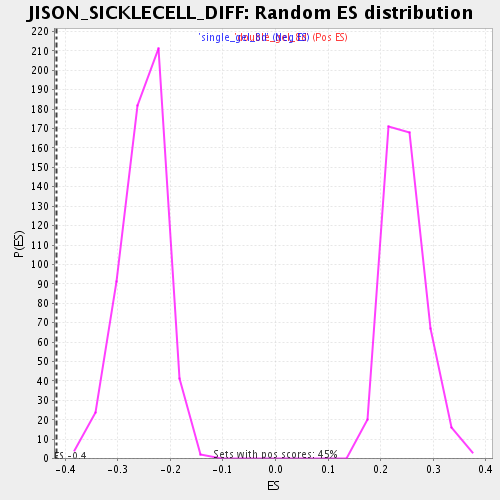
| Enrichment Score (ES) | -0.4171115 |
| Normalized Enrichment Score (NES) | -1.6541272 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.09607506 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LEPR | LEPR Entrez, Source | leptin receptor | 85 | 0.639 | 0.0169 | No |
| 2 | SQLE | SQLE Entrez, Source | squalene epoxidase | 188 | 0.485 | 0.0269 | No |
| 3 | CD9 | CD9 Entrez, Source | CD9 molecule | 512 | 0.295 | 0.0131 | No |
| 4 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 548 | 0.283 | 0.0208 | No |
| 5 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 609 | 0.264 | 0.0259 | No |
| 6 | ATOX1 | ATOX1 Entrez, Source | ATX1 antioxidant protein 1 homolog (yeast) | 653 | 0.251 | 0.0318 | No |
| 7 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 869 | 0.212 | 0.0231 | No |
| 8 | CLIC2 | CLIC2 Entrez, Source | chloride intracellular channel 2 | 935 | 0.202 | 0.0256 | No |
| 9 | P2RX4 | P2RX4 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 4 | 938 | 0.201 | 0.0328 | No |
| 10 | NFE2 | NFE2 Entrez, Source | nuclear factor (erythroid-derived 2), 45kDa | 1071 | 0.185 | 0.0295 | No |
| 11 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 1138 | 0.178 | 0.0310 | No |
| 12 | SLC22A4 | SLC22A4 Entrez, Source | solute carrier family 22 (organic cation transporter), member 4 | 1195 | 0.171 | 0.0330 | No |
| 13 | MCF2L | MCF2L Entrez, Source | MCF.2 cell line derived transforming sequence-like | 1215 | 0.169 | 0.0377 | No |
| 14 | NAB1 | NAB1 Entrez, Source | NGFI-A binding protein 1 (EGR1 binding protein 1) | 1245 | 0.167 | 0.0416 | No |
| 15 | POLG | POLG Entrez, Source | polymerase (DNA directed), gamma | 1288 | 0.165 | 0.0445 | No |
| 16 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 1407 | 0.156 | 0.0412 | No |
| 17 | SELP | SELP Entrez, Source | selectin P (granule membrane protein 140kDa, antigen CD62) | 1408 | 0.156 | 0.0469 | No |
| 18 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1414 | 0.156 | 0.0522 | No |
| 19 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 1691 | 0.137 | 0.0362 | No |
| 20 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 1789 | 0.132 | 0.0337 | No |
| 21 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 1811 | 0.130 | 0.0368 | No |
| 22 | COX17 | COX17 Entrez, Source | COX17 cytochrome c oxidase assembly homolog (S. cerevisiae) | 1921 | 0.125 | 0.0331 | No |
| 23 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 2379 | 0.107 | 0.0023 | No |
| 24 | TALDO1 | TALDO1 Entrez, Source | transaldolase 1 | 2443 | 0.105 | 0.0013 | No |
| 25 | IGF2BP3 | IGF2BP3 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 3 | 2530 | 0.102 | -0.0015 | No |
| 26 | ST3GAL5 | ST3GAL5 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 5 | 2616 | 0.099 | -0.0044 | No |
| 27 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 2707 | 0.095 | -0.0078 | No |
| 28 | TRAF3IP3 | TRAF3IP3 Entrez, Source | TRAF3 interacting protein 3 | 2721 | 0.095 | -0.0053 | No |
| 29 | HBB | HBB Entrez, Source | hemoglobin, beta | 2826 | 0.091 | -0.0099 | No |
| 30 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 2926 | 0.087 | -0.0142 | No |
| 31 | MMP3 | MMP3 Entrez, Source | matrix metallopeptidase 3 (stromelysin 1, progelatinase) | 2981 | 0.085 | -0.0152 | No |
| 32 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 3194 | 0.079 | -0.0285 | No |
| 33 | CPLX2 | CPLX2 Entrez, Source | complexin 2 | 3251 | 0.077 | -0.0299 | No |
| 34 | FGF9 | FGF9 Entrez, Source | fibroblast growth factor 9 (glia-activating factor) | 3279 | 0.077 | -0.0292 | No |
| 35 | F13A1 | F13A1 Entrez, Source | coagulation factor XIII, A1 polypeptide | 3304 | 0.076 | -0.0282 | No |
| 36 | TMEM66 | TMEM66 Entrez, Source | transmembrane protein 66 | 3327 | 0.075 | -0.0272 | No |
| 37 | NOTCH3 | NOTCH3 Entrez, Source | Notch homolog 3 (Drosophila) | 3339 | 0.075 | -0.0253 | No |
| 38 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 3378 | 0.074 | -0.0254 | No |
| 39 | PTPRO | PTPRO Entrez, Source | protein tyrosine phosphatase, receptor type, O | 3457 | 0.072 | -0.0287 | No |
| 40 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 3619 | 0.068 | -0.0385 | No |
| 41 | MSMB | MSMB Entrez, Source | microseminoprotein, beta- | 3764 | 0.065 | -0.0471 | No |
| 42 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 3773 | 0.065 | -0.0453 | No |
| 43 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 3802 | 0.064 | -0.0451 | No |
| 44 | RNF126 | RNF126 Entrez, Source | ring finger protein 126 | 3938 | 0.061 | -0.0532 | No |
| 45 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 3998 | 0.059 | -0.0555 | No |
| 46 | BNIP3L | BNIP3L Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3-like | 4011 | 0.059 | -0.0543 | No |
| 47 | NGFR | NGFR Entrez, Source | nerve growth factor receptor (TNFR superfamily, member 16) | 4017 | 0.059 | -0.0525 | No |
| 48 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 4100 | 0.057 | -0.0566 | No |
| 49 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 4354 | 0.052 | -0.0740 | No |
| 50 | COX6C | COX6C Entrez, Source | cytochrome c oxidase subunit VIc | 4441 | 0.050 | -0.0787 | No |
| 51 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 4819 | 0.041 | -0.1059 | No |
| 52 | IFIT1 | IFIT1 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 1 | 4940 | 0.039 | -0.1136 | No |
| 53 | BLVRA | BLVRA Entrez, Source | biliverdin reductase A | 4973 | 0.038 | -0.1147 | No |
| 54 | CEBPE | CEBPE Entrez, Source | CCAAT/enhancer binding protein (C/EBP), epsilon | 4977 | 0.038 | -0.1135 | No |
| 55 | KLRK1 | KLRK1 Entrez, Source | killer cell lectin-like receptor subfamily K, member 1 | 4998 | 0.038 | -0.1136 | No |
| 56 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 5128 | 0.035 | -0.1222 | No |
| 57 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 5204 | 0.034 | -0.1266 | No |
| 58 | PNMT | PNMT Entrez, Source | phenylethanolamine N-methyltransferase | 5236 | 0.033 | -0.1278 | No |
| 59 | TCTA | TCTA Entrez, Source | T-cell leukemia translocation altered gene | 5450 | 0.029 | -0.1429 | No |
| 60 | SLC36A1 | SLC36A1 Entrez, Source | solute carrier family 36 (proton/amino acid symporter), member 1 | 5594 | 0.026 | -0.1529 | No |
| 61 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 5597 | 0.026 | -0.1521 | No |
| 62 | LYL1 | LYL1 Entrez, Source | lymphoblastic leukemia derived sequence 1 | 5624 | 0.025 | -0.1532 | No |
| 63 | RBL2 | RBL2 Entrez, Source | retinoblastoma-like 2 (p130) | 6125 | 0.016 | -0.1907 | No |
| 64 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 6385 | 0.010 | -0.2100 | No |
| 65 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 6404 | 0.009 | -0.2111 | No |
| 66 | PSMB2 | PSMB2 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 2 | 6524 | 0.007 | -0.2199 | No |
| 67 | HNRPDL | HNRPDL Entrez, Source | heterogeneous nuclear ribonucleoprotein D-like | 6556 | 0.007 | -0.2220 | No |
| 68 | CALR | CALR Entrez, Source | calreticulin | 6558 | 0.007 | -0.2218 | No |
| 69 | SLC4A7 | SLC4A7 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 7 | 6666 | 0.005 | -0.2298 | No |
| 70 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 6683 | 0.004 | -0.2308 | No |
| 71 | PFN2 | PFN2 Entrez, Source | profilin 2 | 6745 | 0.003 | -0.2354 | No |
| 72 | IL5RA | IL5RA Entrez, Source | interleukin 5 receptor, alpha | 6779 | 0.003 | -0.2378 | No |
| 73 | PLEK | PLEK Entrez, Source | pleckstrin | 6834 | 0.002 | -0.2418 | No |
| 74 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 6837 | 0.002 | -0.2419 | No |
| 75 | ANP32B | ANP32B Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member B | 6879 | 0.001 | -0.2450 | No |
| 76 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 6915 | -0.000 | -0.2477 | No |
| 77 | NEK4 | NEK4 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 4 | 6956 | -0.001 | -0.2507 | No |
| 78 | SFRS5 | SFRS5 Entrez, Source | splicing factor, arginine/serine-rich 5 | 7071 | -0.004 | -0.2592 | No |
| 79 | RPL41 | RPL41 Entrez, Source | ribosomal protein L41 | 7088 | -0.004 | -0.2603 | No |
| 80 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 7268 | -0.008 | -0.2737 | No |
| 81 | PABPC4 | PABPC4 Entrez, Source | poly(A) binding protein, cytoplasmic 4 (inducible form) | 7324 | -0.009 | -0.2775 | No |
| 82 | RPL37 | RPL37 Entrez, Source | ribosomal protein L37 | 7466 | -0.012 | -0.2878 | No |
| 83 | VPS52 | VPS52 Entrez, Source | vacuolar protein sorting 52 (S. cerevisiae) | 7507 | -0.013 | -0.2904 | No |
| 84 | GNG5 | GNG5 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 5 | 7559 | -0.014 | -0.2938 | No |
| 85 | VAMP8 | VAMP8 Entrez, Source | vesicle-associated membrane protein 8 (endobrevin) | 7601 | -0.014 | -0.2964 | No |
| 86 | TMED10 | TMED10 Entrez, Source | transmembrane emp24-like trafficking protein 10 (yeast) | 7641 | -0.015 | -0.2988 | No |
| 87 | RBBP6 | RBBP6 Entrez, Source | retinoblastoma binding protein 6 | 7722 | -0.017 | -0.3043 | No |
| 88 | EEF2 | EEF2 Entrez, Source | eukaryotic translation elongation factor 2 | 7765 | -0.018 | -0.3068 | No |
| 89 | RPS6 | RPS6 Entrez, Source | ribosomal protein S6 | 7800 | -0.019 | -0.3087 | No |
| 90 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 7865 | -0.020 | -0.3129 | No |
| 91 | EEF1A1 | EEF1A1 Entrez, Source | eukaryotic translation elongation factor 1 alpha 1 | 7913 | -0.021 | -0.3157 | No |
| 92 | NUCB1 | NUCB1 Entrez, Source | nucleobindin 1 | 7926 | -0.022 | -0.3158 | No |
| 93 | RPLP1 | RPLP1 Entrez, Source | ribosomal protein, large, P1 | 8028 | -0.024 | -0.3226 | No |
| 94 | RHEB | RHEB Entrez, Source | Ras homolog enriched in brain | 8044 | -0.024 | -0.3229 | No |
| 95 | RPL4 | RPL4 Entrez, Source | ribosomal protein L4 | 8146 | -0.026 | -0.3296 | No |
| 96 | TFEC | TFEC Entrez, Source | transcription factor EC | 8147 | -0.026 | -0.3286 | No |
| 97 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 8204 | -0.028 | -0.3319 | No |
| 98 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 8212 | -0.028 | -0.3314 | No |
| 99 | RPS24 | RPS24 Entrez, Source | ribosomal protein S24 | 8294 | -0.030 | -0.3365 | No |
| 100 | RPL9 | RPL9 Entrez, Source | ribosomal protein L9 | 8332 | -0.031 | -0.3382 | No |
| 101 | RPL24 | RPL24 Entrez, Source | ribosomal protein L24 | 8338 | -0.031 | -0.3374 | No |
| 102 | OTUD4 | OTUD4 Entrez, Source | OTU domain containing 4 | 8435 | -0.033 | -0.3435 | No |
| 103 | CTSL | CTSL Entrez, Source | cathepsin L | 8440 | -0.033 | -0.3426 | No |
| 104 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 8446 | -0.033 | -0.3418 | No |
| 105 | RPL5 | RPL5 Entrez, Source | ribosomal protein L5 | 8527 | -0.035 | -0.3466 | No |
| 106 | RPS13 | RPS13 Entrez, Source | ribosomal protein S13 | 8611 | -0.037 | -0.3516 | No |
| 107 | RPL17 | RPL17 Entrez, Source | ribosomal protein L17 | 8625 | -0.037 | -0.3512 | No |
| 108 | TDRD7 | TDRD7 Entrez, Source | tudor domain containing 7 | 8695 | -0.039 | -0.3550 | No |
| 109 | CELSR3 | CELSR3 Entrez, Source | cadherin, EGF LAG seven-pass G-type receptor 3 (flamingo homolog, Drosophila) | 8696 | -0.039 | -0.3536 | No |
| 110 | RPL7 | RPL7 Entrez, Source | ribosomal protein L7 | 8850 | -0.042 | -0.3637 | No |
| 111 | RPL30 | RPL30 Entrez, Source | ribosomal protein L30 | 8866 | -0.043 | -0.3633 | No |
| 112 | RPS27A | RPS27A Entrez, Source | ribosomal protein S27a | 8887 | -0.043 | -0.3632 | No |
| 113 | CASP7 | CASP7 Entrez, Source | caspase 7, apoptosis-related cysteine peptidase | 8912 | -0.044 | -0.3634 | No |
| 114 | RPL19 | RPL19 Entrez, Source | ribosomal protein L19 | 9063 | -0.048 | -0.3731 | No |
| 115 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 9083 | -0.048 | -0.3728 | No |
| 116 | ITGAD | ITGAD Entrez, Source | integrin, alpha D | 9118 | -0.049 | -0.3736 | No |
| 117 | CDS2 | CDS2 Entrez, Source | CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 2 | 9174 | -0.051 | -0.3759 | No |
| 118 | RPL13 | RPL13 Entrez, Source | ribosomal protein L13 | 9201 | -0.051 | -0.3760 | No |
| 119 | PSME2 | PSME2 Entrez, Source | proteasome (prosome, macropain) activator subunit 2 (PA28 beta) | 9205 | -0.051 | -0.3744 | No |
| 120 | RPL6 | RPL6 Entrez, Source | ribosomal protein L6 | 9278 | -0.054 | -0.3779 | No |
| 121 | RPL10A | RPL10A Entrez, Source | ribosomal protein L10a | 9348 | -0.056 | -0.3811 | No |
| 122 | EIF3S6 | EIF3S6 Entrez, Source | eukaryotic translation initiation factor 3, subunit 6 48kDa | 9359 | -0.056 | -0.3798 | No |
| 123 | EIF3S3 | EIF3S3 Entrez, Source | eukaryotic translation initiation factor 3, subunit 3 gamma, 40kDa | 9396 | -0.057 | -0.3805 | No |
| 124 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 9397 | -0.057 | -0.3784 | No |
| 125 | STX7 | STX7 Entrez, Source | syntaxin 7 | 9420 | -0.058 | -0.3780 | No |
| 126 | RPL14 | RPL14 Entrez, Source | ribosomal protein L14 | 9445 | -0.058 | -0.3777 | No |
| 127 | BTF3 | BTF3 Entrez, Source | basic transcription factor 3 | 9484 | -0.059 | -0.3784 | No |
| 128 | EIF3S7 | EIF3S7 Entrez, Source | eukaryotic translation initiation factor 3, subunit 7 zeta, 66/67kDa | 9487 | -0.059 | -0.3764 | No |
| 129 | RPL22 | RPL22 Entrez, Source | ribosomal protein L22 | 9500 | -0.060 | -0.3751 | No |
| 130 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 9623 | -0.063 | -0.3821 | No |
| 131 | GNG11 | GNG11 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 11 | 9666 | -0.064 | -0.3829 | No |
| 132 | GTF2B | GTF2B Entrez, Source | general transcription factor IIB | 9753 | -0.067 | -0.3870 | No |
| 133 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 9794 | -0.068 | -0.3876 | No |
| 134 | AES | AES Entrez, Source | amino-terminal enhancer of split | 9813 | -0.069 | -0.3865 | No |
| 135 | FN1 | FN1 Entrez, Source | fibronectin 1 | 9917 | -0.072 | -0.3917 | No |
| 136 | DDX5 | DDX5 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 5 | 9920 | -0.072 | -0.3892 | No |
| 137 | GSTO1 | GSTO1 Entrez, Source | glutathione S-transferase omega 1 | 10091 | -0.078 | -0.3993 | No |
| 138 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 10098 | -0.078 | -0.3969 | No |
| 139 | RPL11 | RPL11 Entrez, Source | ribosomal protein L11 | 10109 | -0.079 | -0.3947 | No |
| 140 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 10127 | -0.079 | -0.3931 | No |
| 141 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 10129 | -0.079 | -0.3903 | No |
| 142 | FBL | FBL Entrez, Source | fibrillarin | 10172 | -0.080 | -0.3906 | No |
| 143 | EIF4B | EIF4B Entrez, Source | eukaryotic translation initiation factor 4B | 10193 | -0.081 | -0.3891 | No |
| 144 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 10327 | -0.086 | -0.3961 | No |
| 145 | EEF1G | EEF1G Entrez, Source | eukaryotic translation elongation factor 1 gamma | 10329 | -0.086 | -0.3931 | No |
| 146 | EIF4A1 | EIF4A1 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 1 | 10424 | -0.089 | -0.3970 | No |
| 147 | PPP2CB | PPP2CB Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, beta isoform | 10468 | -0.091 | -0.3969 | No |
| 148 | CSNK1E | CSNK1E Entrez, Source | casein kinase 1, epsilon | 10537 | -0.093 | -0.3987 | No |
| 149 | ATP6V1D | ATP6V1D Entrez, Source | ATPase, H+ transporting, lysosomal 34kDa, V1 subunit D | 10581 | -0.095 | -0.3985 | No |
| 150 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 10615 | -0.096 | -0.3976 | No |
| 151 | CDX2 | CDX2 Entrez, Source | caudal type homeobox transcription factor 2 | 10635 | -0.096 | -0.3955 | No |
| 152 | PRDX1 | PRDX1 Entrez, Source | peroxiredoxin 1 | 10681 | -0.098 | -0.3953 | No |
| 153 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 10707 | -0.099 | -0.3936 | No |
| 154 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 10743 | -0.100 | -0.3926 | No |
| 155 | TMED1 | TMED1 Entrez, Source | transmembrane emp24 protein transport domain containing 1 | 10860 | -0.105 | -0.3976 | No |
| 156 | RNF44 | RNF44 Entrez, Source | ring finger protein 44 | 10881 | -0.106 | -0.3953 | No |
| 157 | EMP3 | EMP3 Entrez, Source | epithelial membrane protein 3 | 10906 | -0.107 | -0.3932 | No |
| 158 | HNRPH1 | HNRPH1 Entrez, Source | heterogeneous nuclear ribonucleoprotein H1 (H) | 10957 | -0.109 | -0.3930 | No |
| 159 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 11275 | -0.125 | -0.4125 | Yes |
| 160 | SFPQ | SFPQ Entrez, Source | splicing factor proline/glutamine-rich (polypyrimidine tract binding protein associated) | 11277 | -0.125 | -0.4080 | Yes |
| 161 | TRIM32 | TRIM32 Entrez, Source | tripartite motif-containing 32 | 11322 | -0.127 | -0.4067 | Yes |
| 162 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 11329 | -0.128 | -0.4025 | Yes |
| 163 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 11363 | -0.129 | -0.4003 | Yes |
| 164 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 11479 | -0.136 | -0.4041 | Yes |
| 165 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 11599 | -0.144 | -0.4079 | Yes |
| 166 | GPX1 | GPX1 Entrez, Source | glutathione peroxidase 1 | 11620 | -0.145 | -0.4041 | Yes |
| 167 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 11679 | -0.148 | -0.4031 | Yes |
| 168 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 11694 | -0.149 | -0.3987 | Yes |
| 169 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 11707 | -0.150 | -0.3941 | Yes |
| 170 | BCLAF1 | BCLAF1 Entrez, Source | BCL2-associated transcription factor 1 | 11717 | -0.150 | -0.3893 | Yes |
| 171 | PMM1 | PMM1 Entrez, Source | phosphomannomutase 1 | 11772 | -0.155 | -0.3878 | Yes |
| 172 | ASCC3L1 | ASCC3L1 Entrez, Source | activating signal cointegrator 1 complex subunit 3-like 1 | 11781 | -0.155 | -0.3827 | Yes |
| 173 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 11783 | -0.155 | -0.3771 | Yes |
| 174 | PSMB8 | PSMB8 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7) | 11784 | -0.155 | -0.3714 | Yes |
| 175 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 11832 | -0.159 | -0.3692 | Yes |
| 176 | JAK1 | JAK1 Entrez, Source | Janus kinase 1 (a protein tyrosine kinase) | 11891 | -0.164 | -0.3676 | Yes |
| 177 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 11903 | -0.166 | -0.3624 | Yes |
| 178 | TCN2 | TCN2 Entrez, Source | transcobalamin II; macrocytic anemia | 11922 | -0.167 | -0.3576 | Yes |
| 179 | RAB13 | RAB13 Entrez, Source | RAB13, member RAS oncogene family | 11994 | -0.172 | -0.3567 | Yes |
| 180 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 12163 | -0.188 | -0.3627 | Yes |
| 181 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 12194 | -0.191 | -0.3580 | Yes |
| 182 | IL15 | IL15 Entrez, Source | interleukin 15 | 12233 | -0.197 | -0.3536 | Yes |
| 183 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 12345 | -0.209 | -0.3545 | Yes |
| 184 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 12392 | -0.214 | -0.3501 | Yes |
| 185 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12401 | -0.215 | -0.3429 | Yes |
| 186 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 12413 | -0.216 | -0.3358 | Yes |
| 187 | PPGB | PPGB Entrez, Source | protective protein for beta-galactosidase (galactosialidosis) | 12547 | -0.235 | -0.3373 | Yes |
| 188 | RENBP | RENBP Entrez, Source | renin binding protein | 12639 | -0.251 | -0.3351 | Yes |
| 189 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 12659 | -0.255 | -0.3272 | Yes |
| 190 | CD151 | CD151 Entrez, Source | CD151 molecule (Raph blood group) | 12718 | -0.268 | -0.3218 | Yes |
| 191 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 12732 | -0.272 | -0.3128 | Yes |
| 192 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 12750 | -0.275 | -0.3041 | Yes |
| 193 | ADM | ADM Entrez, Source | adrenomedullin | 12796 | -0.285 | -0.2971 | Yes |
| 194 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 12827 | -0.295 | -0.2886 | Yes |
| 195 | IFI35 | IFI35 Entrez, Source | interferon-induced protein 35 | 12894 | -0.318 | -0.2820 | Yes |
| 196 | SNN | SNN Entrez, Source | stannin | 12996 | -0.362 | -0.2764 | Yes |
| 197 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 12998 | -0.364 | -0.2632 | Yes |
| 198 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 13022 | -0.377 | -0.2511 | Yes |
| 199 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 13029 | -0.381 | -0.2377 | Yes |
| 200 | CLIC1 | CLIC1 Entrez, Source | chloride intracellular channel 1 | 13030 | -0.381 | -0.2237 | Yes |
| 201 | FNBP4 | FNBP4 Entrez, Source | formin binding protein 4 | 13036 | -0.385 | -0.2100 | Yes |
| 202 | NNAT | NNAT Entrez, Source | neuronatin | 13037 | -0.385 | -0.1959 | Yes |
| 203 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 13047 | -0.395 | -0.1822 | Yes |
| 204 | CIRBP | CIRBP Entrez, Source | cold inducible RNA binding protein | 13049 | -0.397 | -0.1677 | Yes |
| 205 | JAK2 | JAK2 Entrez, Source | Janus kinase 2 (a protein tyrosine kinase) | 13065 | -0.406 | -0.1540 | Yes |
| 206 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 13117 | -0.441 | -0.1418 | Yes |
| 207 | GLS | GLS Entrez, Source | glutaminase | 13118 | -0.442 | -0.1256 | Yes |
| 208 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 13120 | -0.443 | -0.1095 | Yes |
| 209 | GPX3 | GPX3 Entrez, Source | glutathione peroxidase 3 (plasma) | 13136 | -0.457 | -0.0939 | Yes |
| 210 | TIMP1 | TIMP1 Entrez, Source | TIMP metallopeptidase inhibitor 1 | 13213 | -0.582 | -0.0784 | Yes |
| 211 | DOK1 | DOK1 Entrez, Source | docking protein 1, 62kDa (downstream of tyrosine kinase 1) | 13216 | -0.591 | -0.0569 | Yes |
| 212 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 13219 | -0.601 | -0.0351 | Yes |
| 213 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 13329 | -1.214 | 0.0010 | Yes |