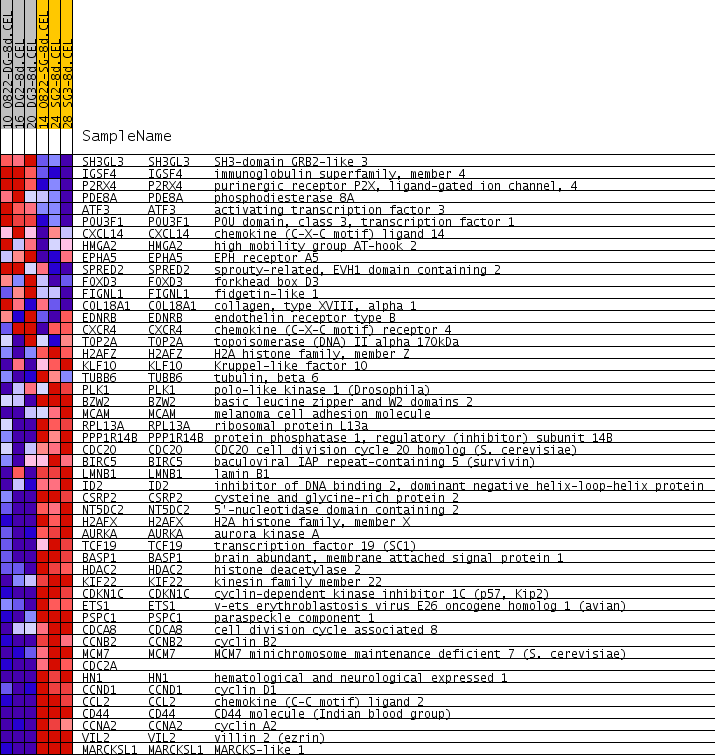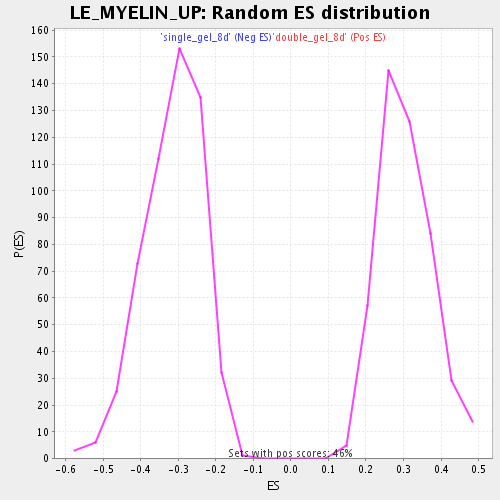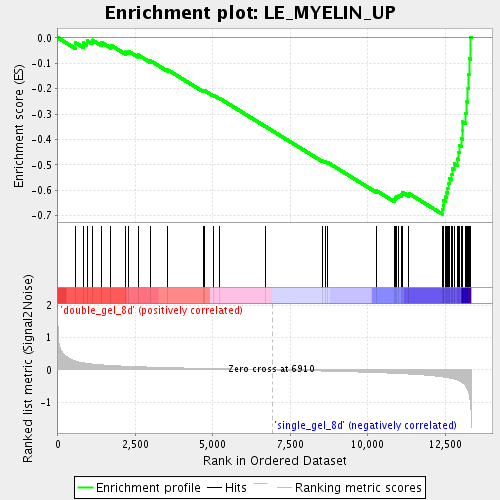
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | LE_MYELIN_UP |
| Enrichment Score (ES) | -0.6937857 |
| Normalized Enrichment Score (NES) | -2.2049801 |
| Nominal p-value | 0.0 |
| FDR q-value | 7.6444185E-4 |
| FWER p-Value | 0.0030 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 574 | 0.276 | -0.0198 | No |
| 2 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 821 | 0.218 | -0.0199 | No |
| 3 | P2RX4 | P2RX4 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 4 | 938 | 0.201 | -0.0115 | No |
| 4 | PDE8A | PDE8A Entrez, Source | phosphodiesterase 8A | 1103 | 0.181 | -0.0085 | No |
| 5 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1414 | 0.156 | -0.0186 | No |
| 6 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 1711 | 0.137 | -0.0294 | No |
| 7 | CXCL14 | CXCL14 Entrez, Source | chemokine (C-X-C motif) ligand 14 | 2190 | 0.115 | -0.0556 | No |
| 8 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 2267 | 0.112 | -0.0519 | No |
| 9 | EPHA5 | EPHA5 Entrez, Source | EPH receptor A5 | 2585 | 0.100 | -0.0673 | No |
| 10 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 2978 | 0.086 | -0.0895 | No |
| 11 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 3547 | 0.070 | -0.1263 | No |
| 12 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 4683 | 0.044 | -0.2080 | No |
| 13 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 4715 | 0.044 | -0.2066 | No |
| 14 | EDNRB | EDNRB Entrez, Source | endothelin receptor type B | 5011 | 0.038 | -0.2256 | No |
| 15 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 5204 | 0.034 | -0.2372 | No |
| 16 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 6709 | 0.004 | -0.3500 | No |
| 17 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 8540 | -0.035 | -0.4846 | No |
| 18 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 8626 | -0.037 | -0.4879 | No |
| 19 | TUBB6 | TUBB6 Entrez, Source | tubulin, beta 6 | 8711 | -0.039 | -0.4909 | No |
| 20 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10276 | -0.084 | -0.6014 | No |
| 21 | BZW2 | BZW2 Entrez, Source | basic leucine zipper and W2 domains 2 | 10870 | -0.106 | -0.6371 | No |
| 22 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 10886 | -0.106 | -0.6293 | No |
| 23 | RPL13A | RPL13A Entrez, Source | ribosomal protein L13a | 10925 | -0.108 | -0.6230 | No |
| 24 | PPP1R14B | PPP1R14B Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 14B | 11006 | -0.112 | -0.6195 | No |
| 25 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 11099 | -0.117 | -0.6165 | No |
| 26 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 11117 | -0.118 | -0.6078 | No |
| 27 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 11329 | -0.128 | -0.6129 | No |
| 28 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12405 | -0.215 | -0.6756 | Yes |
| 29 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 12431 | -0.219 | -0.6589 | Yes |
| 30 | NT5DC2 | NT5DC2 Entrez, Source | 5'-nucleotidase domain containing 2 | 12442 | -0.220 | -0.6410 | Yes |
| 31 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 12504 | -0.229 | -0.6262 | Yes |
| 32 | AURKA | AURKA Entrez, Source | aurora kinase A | 12548 | -0.235 | -0.6096 | Yes |
| 33 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 12573 | -0.239 | -0.5911 | Yes |
| 34 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 12594 | -0.244 | -0.5720 | Yes |
| 35 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 12630 | -0.250 | -0.5535 | Yes |
| 36 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 12713 | -0.267 | -0.5370 | Yes |
| 37 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 12725 | -0.270 | -0.5150 | Yes |
| 38 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12786 | -0.282 | -0.4956 | Yes |
| 39 | PSPC1 | PSPC1 Entrez, Source | paraspeckle component 1 | 12898 | -0.320 | -0.4768 | Yes |
| 40 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 12936 | -0.334 | -0.4513 | Yes |
| 41 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12954 | -0.340 | -0.4238 | Yes |
| 42 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13028 | -0.381 | -0.3971 | Yes |
| 43 | CDC2A | 13061 | -0.405 | -0.3652 | Yes | ||
| 44 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 13064 | -0.406 | -0.3310 | Yes |
| 45 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13162 | -0.500 | -0.2960 | Yes |
| 46 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 13197 | -0.560 | -0.2511 | Yes |
| 47 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 13233 | -0.633 | -0.2002 | Yes |
| 48 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13247 | -0.677 | -0.1438 | Yes |
| 49 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13271 | -0.763 | -0.0810 | Yes |
| 50 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 13312 | -1.018 | 0.0023 | Yes |