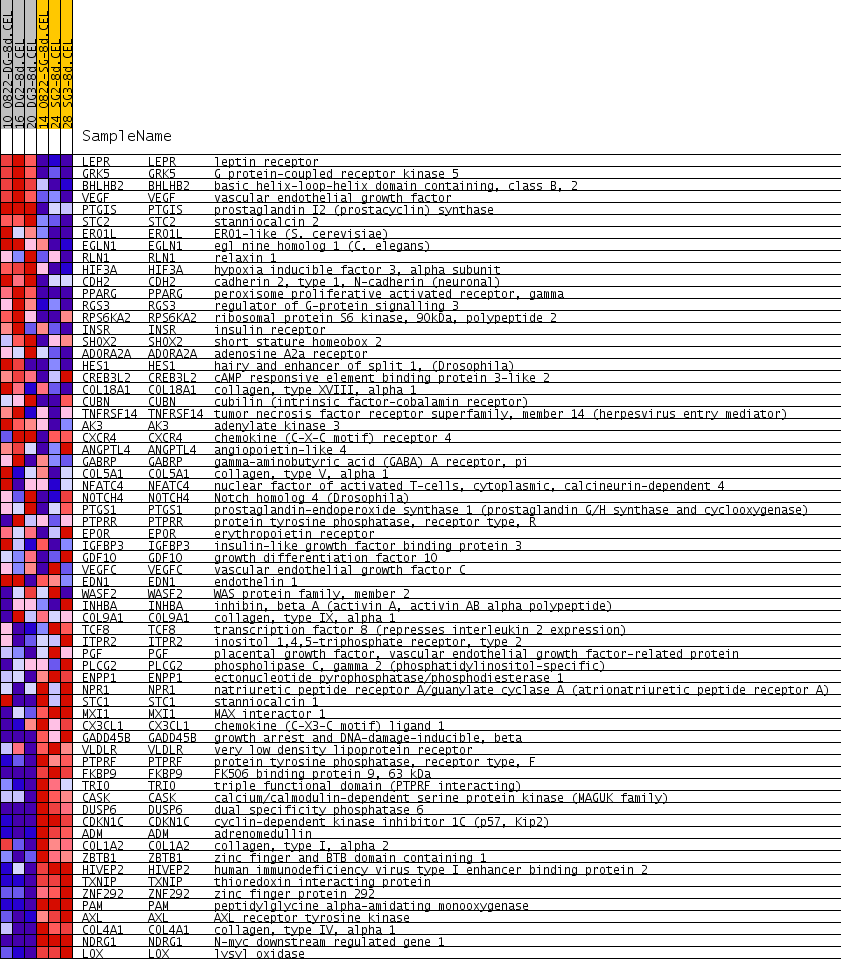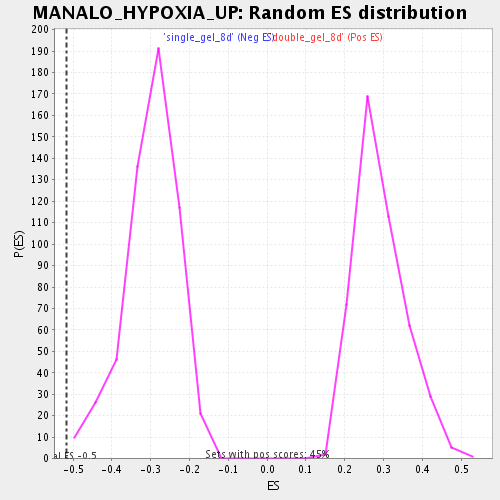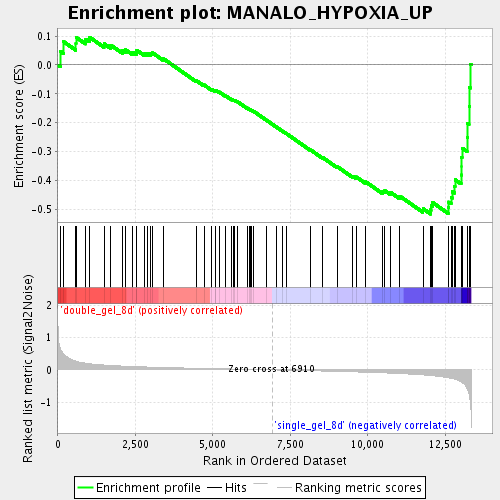
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | MANALO_HYPOXIA_UP |
| Enrichment Score (ES) | -0.5166258 |
| Normalized Enrichment Score (NES) | -1.728507 |
| Nominal p-value | 0.0018281536 |
| FDR q-value | 0.06829199 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LEPR | LEPR Entrez, Source | leptin receptor | 85 | 0.639 | 0.0468 | No |
| 2 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 176 | 0.496 | 0.0813 | No |
| 3 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 581 | 0.272 | 0.0735 | No |
| 4 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 585 | 0.271 | 0.0959 | No |
| 5 | PTGIS | PTGIS Entrez, Source | prostaglandin I2 (prostacyclin) synthase | 894 | 0.208 | 0.0900 | No |
| 6 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 1024 | 0.190 | 0.0961 | No |
| 7 | ERO1L | ERO1L Entrez, Source | ERO1-like (S. cerevisiae) | 1489 | 0.150 | 0.0736 | No |
| 8 | EGLN1 | EGLN1 Entrez, Source | egl nine homolog 1 (C. elegans) | 1704 | 0.137 | 0.0689 | No |
| 9 | RLN1 | RLN1 Entrez, Source | relaxin 1 | 2088 | 0.118 | 0.0499 | No |
| 10 | HIF3A | HIF3A Entrez, Source | hypoxia inducible factor 3, alpha subunit | 2179 | 0.115 | 0.0527 | No |
| 11 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 2409 | 0.106 | 0.0443 | No |
| 12 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 2538 | 0.101 | 0.0431 | No |
| 13 | RGS3 | RGS3 Entrez, Source | regulator of G-protein signalling 3 | 2542 | 0.101 | 0.0513 | No |
| 14 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 2807 | 0.091 | 0.0390 | No |
| 15 | INSR | INSR Entrez, Source | insulin receptor | 2897 | 0.088 | 0.0396 | No |
| 16 | SHOX2 | SHOX2 Entrez, Source | short stature homeobox 2 | 2987 | 0.085 | 0.0400 | No |
| 17 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 3051 | 0.083 | 0.0422 | No |
| 18 | HES1 | HES1 Entrez, Source | hairy and enhancer of split 1, (Drosophila) | 3410 | 0.073 | 0.0213 | No |
| 19 | CREB3L2 | CREB3L2 Entrez, Source | cAMP responsive element binding protein 3-like 2 | 4457 | 0.049 | -0.0533 | No |
| 20 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 4715 | 0.044 | -0.0691 | No |
| 21 | CUBN | CUBN Entrez, Source | cubilin (intrinsic factor-cobalamin receptor) | 4959 | 0.039 | -0.0842 | No |
| 22 | TNFRSF14 | TNFRSF14 Entrez, Source | tumor necrosis factor receptor superfamily, member 14 (herpesvirus entry mediator) | 5071 | 0.037 | -0.0895 | No |
| 23 | AK3 | AK3 Entrez, Source | adenylate kinase 3 | 5082 | 0.036 | -0.0872 | No |
| 24 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 5204 | 0.034 | -0.0935 | No |
| 25 | ANGPTL4 | ANGPTL4 Entrez, Source | angiopoietin-like 4 | 5417 | 0.030 | -0.1070 | No |
| 26 | GABRP | GABRP Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, pi | 5609 | 0.025 | -0.1193 | No |
| 27 | COL5A1 | COL5A1 Entrez, Source | collagen, type V, alpha 1 | 5666 | 0.024 | -0.1215 | No |
| 28 | NFATC4 | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 5709 | 0.024 | -0.1227 | No |
| 29 | NOTCH4 | NOTCH4 Entrez, Source | Notch homolog 4 (Drosophila) | 5781 | 0.022 | -0.1262 | No |
| 30 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 6103 | 0.016 | -0.1490 | No |
| 31 | PTPRR | PTPRR Entrez, Source | protein tyrosine phosphatase, receptor type, R | 6189 | 0.014 | -0.1542 | No |
| 32 | EPOR | EPOR Entrez, Source | erythropoietin receptor | 6228 | 0.014 | -0.1560 | No |
| 33 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 6247 | 0.013 | -0.1562 | No |
| 34 | GDF10 | GDF10 Entrez, Source | growth differentiation factor 10 | 6319 | 0.012 | -0.1606 | No |
| 35 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 6727 | 0.004 | -0.1910 | No |
| 36 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 7069 | -0.004 | -0.2164 | No |
| 37 | WASF2 | WASF2 Entrez, Source | WAS protein family, member 2 | 7261 | -0.008 | -0.2301 | No |
| 38 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 7375 | -0.010 | -0.2378 | No |
| 39 | COL9A1 | COL9A1 Entrez, Source | collagen, type IX, alpha 1 | 8152 | -0.027 | -0.2941 | No |
| 40 | TCF8 | TCF8 Entrez, Source | transcription factor 8 (represses interleukin 2 expression) | 8543 | -0.036 | -0.3205 | No |
| 41 | ITPR2 | ITPR2 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 2 | 9021 | -0.046 | -0.3526 | No |
| 42 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 9515 | -0.060 | -0.3847 | No |
| 43 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 9629 | -0.063 | -0.3879 | No |
| 44 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 9921 | -0.072 | -0.4038 | No |
| 45 | NPR1 | NPR1 Entrez, Source | natriuretic peptide receptor A/guanylate cyclase A (atrionatriuretic peptide receptor A) | 10478 | -0.091 | -0.4382 | No |
| 46 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 10550 | -0.093 | -0.4358 | No |
| 47 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 10722 | -0.099 | -0.4404 | No |
| 48 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 11035 | -0.114 | -0.4544 | No |
| 49 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 11783 | -0.155 | -0.4978 | No |
| 50 | VLDLR | VLDLR Entrez, Source | very low density lipoprotein receptor | 12034 | -0.176 | -0.5020 | Yes |
| 51 | PTPRF | PTPRF Entrez, Source | protein tyrosine phosphatase, receptor type, F | 12043 | -0.177 | -0.4879 | Yes |
| 52 | FKBP9 | FKBP9 Entrez, Source | FK506 binding protein 9, 63 kDa | 12085 | -0.180 | -0.4760 | Yes |
| 53 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 12599 | -0.245 | -0.4942 | Yes |
| 54 | CASK | CASK Entrez, Source | calcium/calmodulin-dependent serine protein kinase (MAGUK family) | 12614 | -0.247 | -0.4747 | Yes |
| 55 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 12692 | -0.262 | -0.4587 | Yes |
| 56 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 12725 | -0.270 | -0.4386 | Yes |
| 57 | ADM | ADM Entrez, Source | adrenomedullin | 12796 | -0.285 | -0.4202 | Yes |
| 58 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 12818 | -0.293 | -0.3974 | Yes |
| 59 | ZBTB1 | ZBTB1 Entrez, Source | zinc finger and BTB domain containing 1 | 13014 | -0.373 | -0.3811 | Yes |
| 60 | HIVEP2 | HIVEP2 Entrez, Source | human immunodeficiency virus type I enhancer binding protein 2 | 13020 | -0.376 | -0.3502 | Yes |
| 61 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 13022 | -0.377 | -0.3189 | Yes |
| 62 | ZNF292 | ZNF292 Entrez, Source | zinc finger protein 292 | 13045 | -0.393 | -0.2879 | Yes |
| 63 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 13208 | -0.576 | -0.2521 | Yes |
| 64 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 13219 | -0.601 | -0.2029 | Yes |
| 65 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 13269 | -0.760 | -0.1433 | Yes |
| 66 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 13282 | -0.795 | -0.0781 | Yes |
| 67 | LOX | LOX Entrez, Source | lysyl oxidase | 13308 | -0.991 | 0.0026 | Yes |