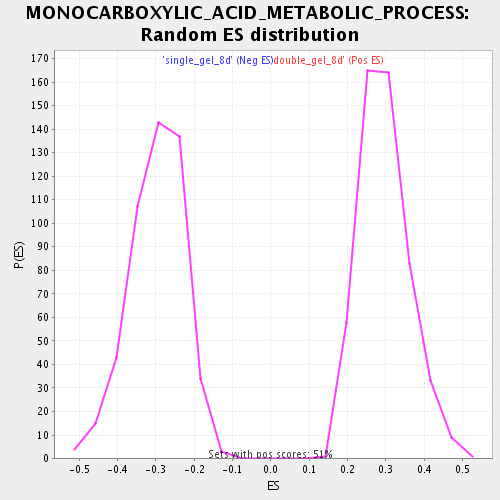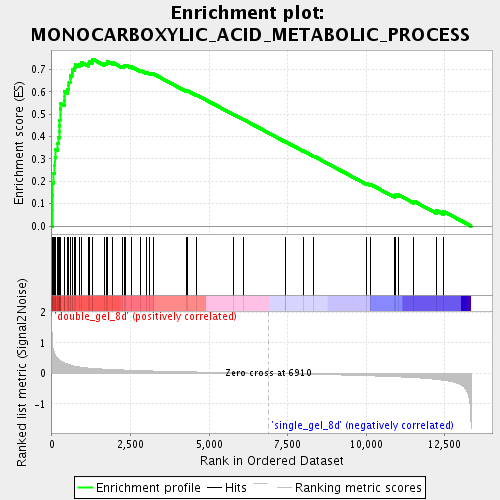
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | MONOCARBOXYLIC_ACID_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.7461566 |
| Normalized Enrichment Score (NES) | 2.5125148 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FTCD | FTCD Entrez, Source | formiminotransferase cyclodeaminase | 9 | 1.171 | 0.0706 | Yes |
| 2 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 10 | 1.121 | 0.1388 | Yes |
| 3 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 16 | 0.933 | 0.1952 | Yes |
| 4 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 60 | 0.721 | 0.2358 | Yes |
| 5 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 95 | 0.621 | 0.2710 | Yes |
| 6 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 98 | 0.617 | 0.3083 | Yes |
| 7 | AGXT | AGXT Entrez, Source | alanine-glyoxylate aminotransferase (oxalosis I; hyperoxaluria I; glycolicaciduria; serine-pyruvate aminotransferase) | 108 | 0.606 | 0.3445 | Yes |
| 8 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 177 | 0.496 | 0.3696 | Yes |
| 9 | ACOT12 | ACOT12 Entrez, Source | acyl-CoA thioesterase 12 | 212 | 0.461 | 0.3951 | Yes |
| 10 | ACOT2 | ACOT2 Entrez, Source | acyl-CoA thioesterase 2 | 231 | 0.450 | 0.4211 | Yes |
| 11 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 236 | 0.445 | 0.4479 | Yes |
| 12 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 247 | 0.438 | 0.4738 | Yes |
| 13 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 259 | 0.427 | 0.4989 | Yes |
| 14 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 260 | 0.427 | 0.5249 | Yes |
| 15 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 284 | 0.409 | 0.5480 | Yes |
| 16 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 392 | 0.341 | 0.5607 | Yes |
| 17 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 401 | 0.338 | 0.5807 | Yes |
| 18 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 403 | 0.337 | 0.6011 | Yes |
| 19 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 501 | 0.299 | 0.6121 | Yes |
| 20 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 537 | 0.288 | 0.6269 | Yes |
| 21 | NR1H4 | NR1H4 Entrez, Source | nuclear receptor subfamily 1, group H, member 4 | 543 | 0.285 | 0.6439 | Yes |
| 22 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 580 | 0.273 | 0.6578 | Yes |
| 23 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 584 | 0.271 | 0.6741 | Yes |
| 24 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 642 | 0.255 | 0.6853 | Yes |
| 25 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 656 | 0.251 | 0.6996 | Yes |
| 26 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 732 | 0.231 | 0.7080 | Yes |
| 27 | ALDH1L1 | ALDH1L1 Entrez, Source | aldehyde dehydrogenase 1 family, member L1 | 750 | 0.228 | 0.7206 | Yes |
| 28 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 870 | 0.211 | 0.7245 | Yes |
| 29 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 950 | 0.200 | 0.7307 | Yes |
| 30 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1158 | 0.175 | 0.7258 | Yes |
| 31 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 1180 | 0.172 | 0.7347 | Yes |
| 32 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 1278 | 0.165 | 0.7374 | Yes |
| 33 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1296 | 0.164 | 0.7462 | Yes |
| 34 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1680 | 0.138 | 0.7257 | No |
| 35 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 1747 | 0.134 | 0.7289 | No |
| 36 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1770 | 0.133 | 0.7353 | No |
| 37 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 1920 | 0.125 | 0.7317 | No |
| 38 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 2249 | 0.113 | 0.7139 | No |
| 39 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 2294 | 0.111 | 0.7173 | No |
| 40 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 2352 | 0.108 | 0.7196 | No |
| 41 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 2527 | 0.102 | 0.7127 | No |
| 42 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 2832 | 0.090 | 0.6953 | No |
| 43 | ARPP-19 | ARPP-19 Entrez, Source | - | 3013 | 0.084 | 0.6869 | No |
| 44 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 3114 | 0.081 | 0.6843 | No |
| 45 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 3231 | 0.078 | 0.6803 | No |
| 46 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 4291 | 0.053 | 0.6038 | No |
| 47 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4310 | 0.053 | 0.6056 | No |
| 48 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 4613 | 0.046 | 0.5857 | No |
| 49 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 5784 | 0.022 | 0.4989 | No |
| 50 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 6103 | 0.016 | 0.4760 | No |
| 51 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 7432 | -0.011 | 0.3767 | No |
| 52 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 7451 | -0.011 | 0.3760 | No |
| 53 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 7996 | -0.023 | 0.3364 | No |
| 54 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8334 | -0.031 | 0.3129 | No |
| 55 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 10024 | -0.075 | 0.1903 | No |
| 56 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 10142 | -0.079 | 0.1864 | No |
| 57 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 10907 | -0.107 | 0.1354 | No |
| 58 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 10944 | -0.109 | 0.1393 | No |
| 59 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 11033 | -0.113 | 0.1396 | No |
| 60 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 11523 | -0.139 | 0.1112 | No |
| 61 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 12246 | -0.198 | 0.0689 | No |
| 62 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 12469 | -0.223 | 0.0657 | No |