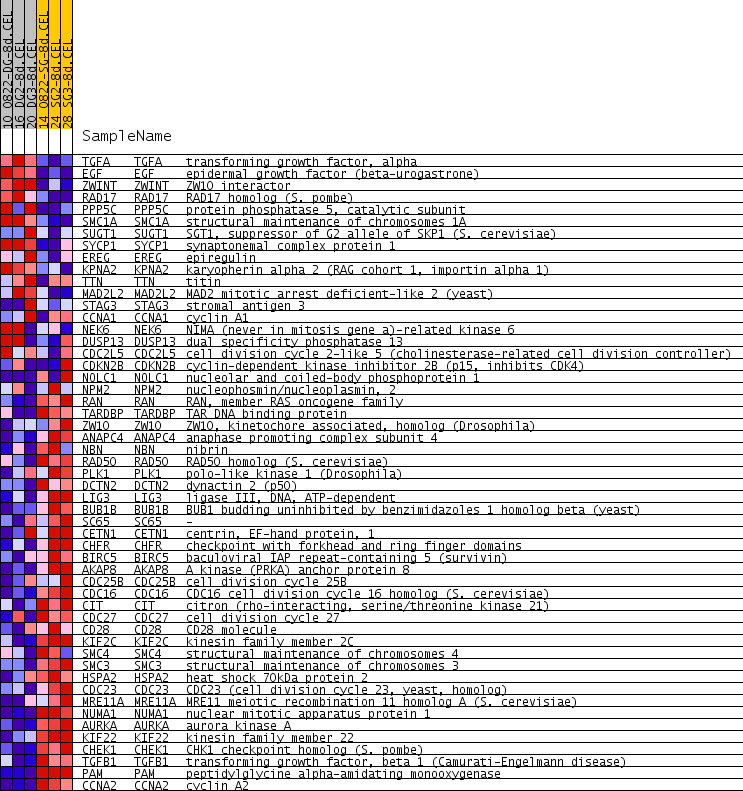
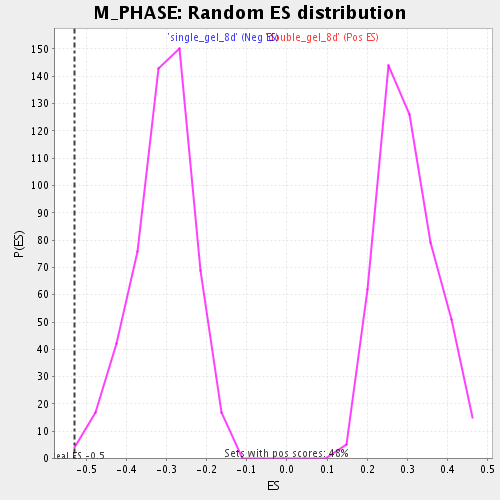
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | M_PHASE |
| Enrichment Score (ES) | -0.52875507 |
| Normalized Enrichment Score (NES) | -1.7119504 |
| Nominal p-value | 0.0038610038 |
| FDR q-value | 0.07549958 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 740 | 0.230 | -0.0235 | No |
| 2 | EGF | EGF Entrez, Source | epidermal growth factor (beta-urogastrone) | 1695 | 0.137 | -0.0761 | No |
| 3 | ZWINT | ZWINT Entrez, Source | ZW10 interactor | 2055 | 0.119 | -0.0864 | No |
| 4 | RAD17 | RAD17 Entrez, Source | RAD17 homolog (S. pombe) | 2601 | 0.099 | -0.1135 | No |
| 5 | PPP5C | PPP5C Entrez, Source | protein phosphatase 5, catalytic subunit | 2945 | 0.087 | -0.1272 | No |
| 6 | SMC1A | SMC1A Entrez, Source | structural maintenance of chromosomes 1A | 3109 | 0.081 | -0.1281 | No |
| 7 | SUGT1 | SUGT1 Entrez, Source | SGT1, suppressor of G2 allele of SKP1 (S. cerevisiae) | 3198 | 0.079 | -0.1236 | No |
| 8 | SYCP1 | SYCP1 Entrez, Source | synaptonemal complex protein 1 | 3323 | 0.075 | -0.1224 | No |
| 9 | EREG | EREG Entrez, Source | epiregulin | 3625 | 0.068 | -0.1355 | No |
| 10 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 3717 | 0.066 | -0.1331 | No |
| 11 | TTN | TTN Entrez, Source | titin | 4401 | 0.050 | -0.1774 | No |
| 12 | MAD2L2 | MAD2L2 Entrez, Source | MAD2 mitotic arrest deficient-like 2 (yeast) | 4562 | 0.047 | -0.1829 | No |
| 13 | STAG3 | STAG3 Entrez, Source | stromal antigen 3 | 5167 | 0.035 | -0.2235 | No |
| 14 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 5345 | 0.031 | -0.2324 | No |
| 15 | NEK6 | NEK6 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 6 | 5393 | 0.030 | -0.2317 | No |
| 16 | DUSP13 | DUSP13 Entrez, Source | dual specificity phosphatase 13 | 5702 | 0.024 | -0.2516 | No |
| 17 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 5925 | 0.020 | -0.2655 | No |
| 18 | CDKN2B | CDKN2B Entrez, Source | cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4) | 7589 | -0.014 | -0.3886 | No |
| 19 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 9128 | -0.050 | -0.4974 | No |
| 20 | NPM2 | NPM2 Entrez, Source | nucleophosmin/nucleoplasmin, 2 | 9187 | -0.051 | -0.4946 | No |
| 21 | RAN | RAN Entrez, Source | RAN, member RAS oncogene family | 9534 | -0.061 | -0.5122 | No |
| 22 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 9617 | -0.063 | -0.5095 | No |
| 23 | ZW10 | ZW10 Entrez, Source | ZW10, kinetochore associated, homolog (Drosophila) | 9627 | -0.063 | -0.5013 | No |
| 24 | ANAPC4 | ANAPC4 Entrez, Source | anaphase promoting complex subunit 4 | 9814 | -0.069 | -0.5057 | No |
| 25 | NBN | NBN Entrez, Source | nibrin | 10122 | -0.079 | -0.5177 | Yes |
| 26 | RAD50 | RAD50 Entrez, Source | RAD50 homolog (S. cerevisiae) | 10151 | -0.080 | -0.5086 | Yes |
| 27 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10276 | -0.084 | -0.5062 | Yes |
| 28 | DCTN2 | DCTN2 Entrez, Source | dynactin 2 (p50) | 10303 | -0.085 | -0.4963 | Yes |
| 29 | LIG3 | LIG3 Entrez, Source | ligase III, DNA, ATP-dependent | 10312 | -0.085 | -0.4850 | Yes |
| 30 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 10391 | -0.088 | -0.4786 | Yes |
| 31 | SC65 | SC65 Entrez, Source | - | 10443 | -0.090 | -0.4698 | Yes |
| 32 | CETN1 | CETN1 Entrez, Source | centrin, EF-hand protein, 1 | 10599 | -0.095 | -0.4682 | Yes |
| 33 | CHFR | CHFR Entrez, Source | checkpoint with forkhead and ring finger domains | 10712 | -0.099 | -0.4628 | Yes |
| 34 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 11117 | -0.118 | -0.4767 | Yes |
| 35 | AKAP8 | AKAP8 Entrez, Source | A kinase (PRKA) anchor protein 8 | 11248 | -0.124 | -0.4691 | Yes |
| 36 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 11352 | -0.129 | -0.4588 | Yes |
| 37 | CDC16 | CDC16 Entrez, Source | CDC16 cell division cycle 16 homolog (S. cerevisiae) | 11467 | -0.135 | -0.4484 | Yes |
| 38 | CIT | CIT Entrez, Source | citron (rho-interacting, serine/threonine kinase 21) | 11639 | -0.146 | -0.4408 | Yes |
| 39 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 11658 | -0.148 | -0.4215 | Yes |
| 40 | CD28 | CD28 Entrez, Source | CD28 molecule | 11745 | -0.152 | -0.4067 | Yes |
| 41 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 11889 | -0.164 | -0.3946 | Yes |
| 42 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 11916 | -0.167 | -0.3732 | Yes |
| 43 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 12021 | -0.175 | -0.3565 | Yes |
| 44 | HSPA2 | HSPA2 Entrez, Source | heat shock 70kDa protein 2 | 12146 | -0.186 | -0.3398 | Yes |
| 45 | CDC23 | CDC23 Entrez, Source | CDC23 (cell division cycle 23, yeast, homolog) | 12182 | -0.190 | -0.3159 | Yes |
| 46 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 12243 | -0.198 | -0.2927 | Yes |
| 47 | NUMA1 | NUMA1 Entrez, Source | nuclear mitotic apparatus protein 1 | 12531 | -0.233 | -0.2817 | Yes |
| 48 | AURKA | AURKA Entrez, Source | aurora kinase A | 12548 | -0.235 | -0.2500 | Yes |
| 49 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 12713 | -0.267 | -0.2249 | Yes |
| 50 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12821 | -0.294 | -0.1918 | Yes |
| 51 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 13047 | -0.395 | -0.1535 | Yes |
| 52 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 13208 | -0.576 | -0.0848 | Yes |
| 53 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13247 | -0.677 | 0.0071 | Yes |