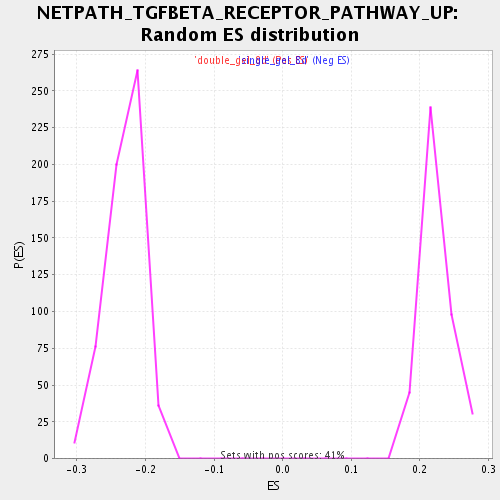
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | NETPATH_TGFBETA_RECEPTOR_PATHWAY_UP |
| Enrichment Score (ES) | -0.39395806 |
| Normalized Enrichment Score (NES) | -1.7173586 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07384236 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAP3K8 | MAP3K8 Entrez, Source | mitogen-activated protein kinase kinase kinase 8 | 34 | 0.817 | 0.0074 | No |
| 2 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 56 | 0.731 | 0.0147 | No |
| 3 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 64 | 0.710 | 0.0228 | No |
| 4 | INSIG1 | INSIG1 Entrez, Source | insulin induced gene 1 | 83 | 0.642 | 0.0293 | No |
| 5 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 84 | 0.641 | 0.0371 | No |
| 6 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 110 | 0.603 | 0.0426 | No |
| 7 | CTH | CTH Entrez, Source | cystathionase (cystathionine gamma-lyase) | 148 | 0.535 | 0.0462 | No |
| 8 | CBS | CBS Entrez, Source | cystathionine-beta-synthase | 158 | 0.523 | 0.0519 | No |
| 9 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 169 | 0.502 | 0.0573 | No |
| 10 | ACLY | ACLY Entrez, Source | ATP citrate lyase | 186 | 0.487 | 0.0620 | No |
| 11 | DBP | DBP Entrez, Source | D site of albumin promoter (albumin D-box) binding protein | 195 | 0.480 | 0.0673 | No |
| 12 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 199 | 0.477 | 0.0729 | No |
| 13 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 236 | 0.445 | 0.0755 | No |
| 14 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 237 | 0.445 | 0.0810 | No |
| 15 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 277 | 0.416 | 0.0830 | No |
| 16 | PCSK6 | PCSK6 Entrez, Source | proprotein convertase subtilisin/kexin type 6 | 279 | 0.416 | 0.0880 | No |
| 17 | ADRB2 | ADRB2 Entrez, Source | adrenergic, beta-2-, receptor, surface | 298 | 0.399 | 0.0915 | No |
| 18 | MASP2 | MASP2 Entrez, Source | mannan-binding lectin serine peptidase 2 | 308 | 0.392 | 0.0956 | No |
| 19 | TRIB3 | TRIB3 Entrez, Source | tribbles homolog 3 (Drosophila) | 317 | 0.384 | 0.0997 | No |
| 20 | EBP | EBP Entrez, Source | emopamil binding protein (sterol isomerase) | 324 | 0.381 | 0.1039 | No |
| 21 | AGTR1 | AGTR1 Entrez, Source | angiotensin II receptor, type 1 | 341 | 0.372 | 0.1072 | No |
| 22 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 383 | 0.346 | 0.1082 | No |
| 23 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 401 | 0.338 | 0.1111 | No |
| 24 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 402 | 0.338 | 0.1152 | No |
| 25 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 425 | 0.331 | 0.1175 | No |
| 26 | MUC4 | MUC4 Entrez, Source | mucin 4, cell surface associated | 427 | 0.330 | 0.1215 | No |
| 27 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 434 | 0.327 | 0.1250 | No |
| 28 | SLC22A7 | SLC22A7 Entrez, Source | solute carrier family 22 (organic anion transporter), member 7 | 437 | 0.326 | 0.1289 | No |
| 29 | INPP4B | INPP4B Entrez, Source | inositol polyphosphate-4-phosphatase, type II, 105kDa | 463 | 0.314 | 0.1308 | No |
| 30 | SREBF1 | SREBF1 Entrez, Source | sterol regulatory element binding transcription factor 1 | 467 | 0.313 | 0.1344 | No |
| 31 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 470 | 0.312 | 0.1380 | No |
| 32 | MVK | MVK Entrez, Source | mevalonate kinase (mevalonic aciduria) | 482 | 0.309 | 0.1410 | No |
| 33 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 511 | 0.296 | 0.1424 | No |
| 34 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 546 | 0.284 | 0.1432 | No |
| 35 | SULT1E1 | SULT1E1 Entrez, Source | sulfotransferase family 1E, estrogen-preferring, member 1 | 571 | 0.277 | 0.1448 | No |
| 36 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 581 | 0.272 | 0.1474 | No |
| 37 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 585 | 0.271 | 0.1505 | No |
| 38 | FOXO1A | FOXO1A Entrez, Source | forkhead box O1A (rhabdomyosarcoma) | 596 | 0.268 | 0.1530 | No |
| 39 | SC5DL | SC5DL Entrez, Source | sterol-C5-desaturase (ERG3 delta-5-desaturase homolog, fungal)-like | 617 | 0.261 | 0.1546 | No |
| 40 | CTF1 | CTF1 Entrez, Source | cardiotrophin 1 | 648 | 0.252 | 0.1554 | No |
| 41 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 670 | 0.247 | 0.1568 | No |
| 42 | GTF3A | GTF3A Entrez, Source | general transcription factor IIIA | 689 | 0.241 | 0.1583 | No |
| 43 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 720 | 0.235 | 0.1589 | No |
| 44 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 740 | 0.230 | 0.1602 | No |
| 45 | SLC6A11 | SLC6A11 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, GABA), member 11 | 756 | 0.226 | 0.1618 | No |
| 46 | ALOX5AP | ALOX5AP Entrez, Source | arachidonate 5-lipoxygenase-activating protein | 764 | 0.225 | 0.1640 | No |
| 47 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 837 | 0.215 | 0.1611 | No |
| 48 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 867 | 0.212 | 0.1614 | No |
| 49 | IRAK2 | IRAK2 Entrez, Source | interleukin-1 receptor-associated kinase 2 | 868 | 0.212 | 0.1640 | No |
| 50 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 880 | 0.210 | 0.1657 | No |
| 51 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 922 | 0.204 | 0.1650 | No |
| 52 | CDH11 | CDH11 Entrez, Source | cadherin 11, type 2, OB-cadherin (osteoblast) | 1034 | 0.189 | 0.1587 | No |
| 53 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 1037 | 0.188 | 0.1608 | No |
| 54 | ALDOC | ALDOC Entrez, Source | aldolase C, fructose-bisphosphate | 1047 | 0.187 | 0.1624 | No |
| 55 | APOA1 | APOA1 Entrez, Source | apolipoprotein A-I | 1052 | 0.187 | 0.1644 | No |
| 56 | CDC2L1 | CDC2L1 Entrez, Source | cell division cycle 2-like 1 (PITSLRE proteins) | 1182 | 0.172 | 0.1565 | No |
| 57 | SLC23A1 | SLC23A1 Entrez, Source | solute carrier family 23 (nucleobase transporters), member 1 | 1187 | 0.172 | 0.1583 | No |
| 58 | CAPN9 | CAPN9 Entrez, Source | calpain 9 | 1191 | 0.172 | 0.1602 | No |
| 59 | ODC1 | ODC1 Entrez, Source | ornithine decarboxylase 1 | 1205 | 0.170 | 0.1612 | No |
| 60 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 1217 | 0.169 | 0.1624 | No |
| 61 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 1238 | 0.168 | 0.1629 | No |
| 62 | GTPBP2 | GTPBP2 Entrez, Source | GTP binding protein 2 | 1276 | 0.165 | 0.1621 | No |
| 63 | FGF18 | FGF18 Entrez, Source | fibroblast growth factor 18 | 1301 | 0.164 | 0.1622 | No |
| 64 | ST8SIA4 | ST8SIA4 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 | 1308 | 0.164 | 0.1638 | No |
| 65 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 1319 | 0.163 | 0.1650 | No |
| 66 | CYP1A1 | CYP1A1 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 1 | 1331 | 0.162 | 0.1661 | No |
| 67 | MUCDHL | MUCDHL Entrez, Source | mucin and cadherin-like | 1335 | 0.162 | 0.1679 | No |
| 68 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 1338 | 0.162 | 0.1697 | No |
| 69 | EIF4EBP1 | EIF4EBP1 Entrez, Source | eukaryotic translation initiation factor 4E binding protein 1 | 1395 | 0.157 | 0.1673 | No |
| 70 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1414 | 0.156 | 0.1678 | No |
| 71 | ITIH3 | ITIH3 Entrez, Source | inter-alpha (globulin) inhibitor H3 | 1430 | 0.155 | 0.1685 | No |
| 72 | IL13RA1 | IL13RA1 Entrez, Source | interleukin 13 receptor, alpha 1 | 1446 | 0.153 | 0.1692 | No |
| 73 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 1558 | 0.145 | 0.1624 | No |
| 74 | FGF12 | FGF12 Entrez, Source | fibroblast growth factor 12 | 1563 | 0.145 | 0.1638 | No |
| 75 | TGM1 | TGM1 Entrez, Source | transglutaminase 1 (K polypeptide epidermal type I, protein-glutamine-gamma-glutamyltransferase) | 1589 | 0.144 | 0.1636 | No |
| 76 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 1691 | 0.137 | 0.1575 | No |
| 77 | EGF | EGF Entrez, Source | epidermal growth factor (beta-urogastrone) | 1695 | 0.137 | 0.1589 | No |
| 78 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 1711 | 0.137 | 0.1594 | No |
| 79 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 1808 | 0.131 | 0.1536 | No |
| 80 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 1875 | 0.127 | 0.1500 | No |
| 81 | ESM1 | ESM1 Entrez, Source | endothelial cell-specific molecule 1 | 1922 | 0.125 | 0.1480 | No |
| 82 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 1938 | 0.125 | 0.1483 | No |
| 83 | MPP3 | MPP3 Entrez, Source | membrane protein, palmitoylated 3 (MAGUK p55 subfamily member 3) | 2026 | 0.121 | 0.1430 | No |
| 84 | WDR45 | WDR45 Entrez, Source | WD repeat domain 45 | 2037 | 0.120 | 0.1437 | No |
| 85 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 2067 | 0.119 | 0.1429 | No |
| 86 | RABEP1 | RABEP1 Entrez, Source | rabaptin, RAB GTPase binding effector protein 1 | 2072 | 0.118 | 0.1441 | No |
| 87 | RAPGEF3 | RAPGEF3 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 3 | 2114 | 0.117 | 0.1423 | No |
| 88 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 2116 | 0.117 | 0.1437 | No |
| 89 | FKBP8 | FKBP8 Entrez, Source | FK506 binding protein 8, 38kDa | 2157 | 0.116 | 0.1420 | No |
| 90 | FASTK | FASTK Entrez, Source | Fas-activated serine/threonine kinase | 2218 | 0.114 | 0.1387 | No |
| 91 | PDZK1IP1 | PDZK1IP1 Entrez, Source | PDZK1 interacting protein 1 | 2299 | 0.110 | 0.1339 | No |
| 92 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 2409 | 0.106 | 0.1267 | No |
| 93 | ASNS | ASNS Entrez, Source | asparagine synthetase | 2424 | 0.106 | 0.1269 | No |
| 94 | ARNTL | ARNTL Entrez, Source | aryl hydrocarbon receptor nuclear translocator-like | 2555 | 0.101 | 0.1180 | No |
| 95 | MAPK10 | MAPK10 Entrez, Source | mitogen-activated protein kinase 10 | 2562 | 0.101 | 0.1188 | No |
| 96 | CGA | CGA Entrez, Source | glycoprotein hormones, alpha polypeptide | 2568 | 0.100 | 0.1196 | No |
| 97 | AKT2 | AKT2 Entrez, Source | v-akt murine thymoma viral oncogene homolog 2 | 2581 | 0.100 | 0.1199 | No |
| 98 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 2588 | 0.100 | 0.1207 | No |
| 99 | TIMM8B | TIMM8B Entrez, Source | translocase of inner mitochondrial membrane 8 homolog B (yeast) | 2592 | 0.100 | 0.1217 | No |
| 100 | EYA2 | EYA2 Entrez, Source | eyes absent homolog 2 (Drosophila) | 2749 | 0.094 | 0.1107 | No |
| 101 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 2781 | 0.092 | 0.1094 | No |
| 102 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 2846 | 0.090 | 0.1056 | No |
| 103 | GFER | GFER Entrez, Source | growth factor, augmenter of liver regeneration (ERV1 homolog, S. cerevisiae) | 2850 | 0.090 | 0.1064 | No |
| 104 | SCRG1 | SCRG1 Entrez, Source | - | 2851 | 0.089 | 0.1075 | No |
| 105 | CORIN | CORIN Entrez, Source | corin, serine peptidase | 2880 | 0.089 | 0.1064 | No |
| 106 | NEU3 | NEU3 Entrez, Source | sialidase 3 (membrane sialidase) | 2937 | 0.087 | 0.1031 | No |
| 107 | ATP2A1 | ATP2A1 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, fast twitch 1 | 2970 | 0.086 | 0.1017 | No |
| 108 | MMP3 | MMP3 Entrez, Source | matrix metallopeptidase 3 (stromelysin 1, progelatinase) | 2981 | 0.085 | 0.1020 | No |
| 109 | MPDU1 | MPDU1 Entrez, Source | mannose-P-dolichol utilization defect 1 | 3005 | 0.085 | 0.1012 | No |
| 110 | EIF1 | EIF1 Entrez, Source | eukaryotic translation initiation factor 1 | 3008 | 0.085 | 0.1021 | No |
| 111 | PGRMC2 | PGRMC2 Entrez, Source | progesterone receptor membrane component 2 | 3016 | 0.084 | 0.1026 | No |
| 112 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 3032 | 0.084 | 0.1025 | No |
| 113 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 3035 | 0.084 | 0.1033 | No |
| 114 | CASP4 | CASP4 Entrez, Source | caspase 4, apoptosis-related cysteine peptidase | 3096 | 0.082 | 0.0997 | No |
| 115 | CAST | CAST Entrez, Source | calpastatin | 3197 | 0.079 | 0.0929 | No |
| 116 | LIMK2 | LIMK2 Entrez, Source | LIM domain kinase 2 | 3234 | 0.078 | 0.0910 | No |
| 117 | BZRAP1 | BZRAP1 Entrez, Source | benzodiazapine receptor (peripheral) associated protein 1 | 3235 | 0.078 | 0.0920 | No |
| 118 | L1CAM | L1CAM Entrez, Source | L1 cell adhesion molecule | 3249 | 0.077 | 0.0919 | No |
| 119 | SHC1 | SHC1 Entrez, Source | SHC (Src homology 2 domain containing) transforming protein 1 | 3311 | 0.076 | 0.0881 | No |
| 120 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3320 | 0.076 | 0.0884 | No |
| 121 | CAMK2B | CAMK2B Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II beta | 3351 | 0.075 | 0.0870 | No |
| 122 | PTPRH | PTPRH Entrez, Source | protein tyrosine phosphatase, receptor type, H | 3376 | 0.074 | 0.0860 | No |
| 123 | HES1 | HES1 Entrez, Source | hairy and enhancer of split 1, (Drosophila) | 3410 | 0.073 | 0.0844 | No |
| 124 | IER3 | IER3 Entrez, Source | immediate early response 3 | 3649 | 0.068 | 0.0667 | No |
| 125 | MAP2K3 | MAP2K3 Entrez, Source | mitogen-activated protein kinase kinase 3 | 3755 | 0.065 | 0.0593 | No |
| 126 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 3759 | 0.065 | 0.0599 | No |
| 127 | FEZ2 | FEZ2 Entrez, Source | fasciculation and elongation protein zeta 2 (zygin II) | 3801 | 0.064 | 0.0575 | No |
| 128 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 3805 | 0.064 | 0.0580 | No |
| 129 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 3839 | 0.063 | 0.0562 | No |
| 130 | RAF1 | RAF1 Entrez, Source | v-raf-1 murine leukemia viral oncogene homolog 1 | 3847 | 0.063 | 0.0565 | No |
| 131 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 3892 | 0.062 | 0.0538 | No |
| 132 | NUDT21 | NUDT21 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 21 | 3927 | 0.061 | 0.0519 | No |
| 133 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 3936 | 0.061 | 0.0521 | No |
| 134 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 3998 | 0.059 | 0.0480 | No |
| 135 | GSR | GSR Entrez, Source | glutathione reductase | 4018 | 0.059 | 0.0473 | No |
| 136 | SLC35B1 | SLC35B1 Entrez, Source | solute carrier family 35, member B1 | 4043 | 0.058 | 0.0461 | No |
| 137 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 4054 | 0.058 | 0.0461 | No |
| 138 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 4056 | 0.058 | 0.0467 | No |
| 139 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 4062 | 0.058 | 0.0470 | No |
| 140 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 4096 | 0.057 | 0.0452 | No |
| 141 | FBN1 | FBN1 Entrez, Source | fibrillin 1 | 4122 | 0.057 | 0.0439 | No |
| 142 | NELL1 | NELL1 Entrez, Source | NEL-like 1 (chicken) | 4151 | 0.056 | 0.0424 | No |
| 143 | LHX2 | LHX2 Entrez, Source | LIM homeobox 2 | 4202 | 0.055 | 0.0392 | No |
| 144 | SOS2 | SOS2 Entrez, Source | son of sevenless homolog 2 (Drosophila) | 4217 | 0.055 | 0.0388 | No |
| 145 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 4254 | 0.054 | 0.0367 | No |
| 146 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 4291 | 0.053 | 0.0345 | No |
| 147 | WIF1 | WIF1 Entrez, Source | WNT inhibitory factor 1 | 4395 | 0.051 | 0.0271 | No |
| 148 | ARPC1A | ARPC1A Entrez, Source | actin related protein 2/3 complex, subunit 1A, 41kDa | 4419 | 0.050 | 0.0260 | No |
| 149 | BUD31 | BUD31 Entrez, Source | BUD31 homolog (yeast) | 4471 | 0.049 | 0.0226 | No |
| 150 | INSL6 | INSL6 Entrez, Source | insulin-like 6 | 4531 | 0.048 | 0.0186 | No |
| 151 | GPR30 | GPR30 Entrez, Source | G protein-coupled receptor 30 | 4567 | 0.047 | 0.0165 | No |
| 152 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 4574 | 0.047 | 0.0166 | No |
| 153 | DAP3 | DAP3 Entrez, Source | death associated protein 3 | 4590 | 0.047 | 0.0160 | No |
| 154 | ELN | ELN Entrez, Source | elastin (supravalvular aortic stenosis, Williams-Beuren syndrome) | 4595 | 0.046 | 0.0162 | No |
| 155 | RAMP1 | RAMP1 Entrez, Source | receptor (calcitonin) activity modifying protein 1 | 4600 | 0.046 | 0.0165 | No |
| 156 | WIPI2 | WIPI2 Entrez, Source | WD repeat domain, phosphoinositide interacting 2 | 4601 | 0.046 | 0.0171 | No |
| 157 | POLD3 | POLD3 Entrez, Source | polymerase (DNA-directed), delta 3, accessory subunit | 4626 | 0.046 | 0.0157 | No |
| 158 | HCN4 | HCN4 Entrez, Source | hyperpolarization activated cyclic nucleotide-gated potassium channel 4 | 4661 | 0.045 | 0.0137 | No |
| 159 | MT3 | MT3 Entrez, Source | metallothionein 3 (growth inhibitory factor (neurotrophic)) | 4680 | 0.044 | 0.0128 | No |
| 160 | IL12A | IL12A Entrez, Source | interleukin 12A (natural killer cell stimulatory factor 1, cytotoxic lymphocyte maturation factor 1, p35) | 4872 | 0.040 | -0.0015 | No |
| 161 | CLK1 | CLK1 Entrez, Source | CDC-like kinase 1 | 4880 | 0.040 | -0.0016 | No |
| 162 | RBP1 | RBP1 Entrez, Source | retinol binding protein 1, cellular | 4881 | 0.040 | -0.0011 | No |
| 163 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 4937 | 0.039 | -0.0049 | No |
| 164 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 4957 | 0.039 | -0.0059 | No |
| 165 | GPR37L1 | GPR37L1 Entrez, Source | G protein-coupled receptor 37 like 1 | 5017 | 0.038 | -0.0100 | No |
| 166 | WISP1 | WISP1 Entrez, Source | WNT1 inducible signaling pathway protein 1 | 5022 | 0.038 | -0.0099 | No |
| 167 | AKAP12 | AKAP12 Entrez, Source | A kinase (PRKA) anchor protein (gravin) 12 | 5084 | 0.036 | -0.0142 | No |
| 168 | C5AR1 | C5AR1 Entrez, Source | complement component 5a receptor 1 | 5189 | 0.034 | -0.0218 | No |
| 169 | PADI3 | PADI3 Entrez, Source | peptidyl arginine deiminase, type III | 5216 | 0.034 | -0.0234 | No |
| 170 | GABARAPL2 | GABARAPL2 Entrez, Source | GABA(A) receptor-associated protein-like 2 | 5307 | 0.032 | -0.0300 | No |
| 171 | TFPI2 | TFPI2 Entrez, Source | tissue factor pathway inhibitor 2 | 5317 | 0.032 | -0.0303 | No |
| 172 | COMP | COMP Entrez, Source | cartilage oligomeric matrix protein | 5328 | 0.031 | -0.0307 | No |
| 173 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 5428 | 0.029 | -0.0381 | No |
| 174 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 5484 | 0.028 | -0.0420 | No |
| 175 | RYR1 | RYR1 Entrez, Source | ryanodine receptor 1 (skeletal) | 5571 | 0.026 | -0.0484 | No |
| 176 | COL5A1 | COL5A1 Entrez, Source | collagen, type V, alpha 1 | 5666 | 0.024 | -0.0554 | No |
| 177 | IFITM3 | IFITM3 Entrez, Source | interferon induced transmembrane protein 3 (1-8U) | 5763 | 0.023 | -0.0625 | No |
| 178 | NUDT3 | NUDT3 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 3 | 5804 | 0.022 | -0.0654 | No |
| 179 | ACVR2A | ACVR2A Entrez, Source | activin A receptor, type IIA | 5863 | 0.021 | -0.0696 | No |
| 180 | ATG3 | ATG3 Entrez, Source | ATG3 autophagy related 3 homolog (S. cerevisiae) | 5867 | 0.021 | -0.0696 | No |
| 181 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 5893 | 0.020 | -0.0713 | No |
| 182 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 5913 | 0.020 | -0.0725 | No |
| 183 | THBS2 | THBS2 Entrez, Source | thrombospondin 2 | 6077 | 0.017 | -0.0850 | No |
| 184 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 6082 | 0.017 | -0.0851 | No |
| 185 | WBP4 | WBP4 Entrez, Source | WW domain binding protein 4 (formin binding protein 21) | 6097 | 0.016 | -0.0860 | No |
| 186 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 6103 | 0.016 | -0.0862 | No |
| 187 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 6115 | 0.016 | -0.0868 | No |
| 188 | RBL2 | RBL2 Entrez, Source | retinoblastoma-like 2 (p130) | 6125 | 0.016 | -0.0873 | No |
| 189 | BRE | BRE Entrez, Source | brain and reproductive organ-expressed (TNFRSF1A modulator) | 6169 | 0.015 | -0.0905 | No |
| 190 | PTPRR | PTPRR Entrez, Source | protein tyrosine phosphatase, receptor type, R | 6189 | 0.014 | -0.0918 | No |
| 191 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 6247 | 0.013 | -0.0961 | No |
| 192 | PLCB3 | PLCB3 Entrez, Source | phospholipase C, beta 3 (phosphatidylinositol-specific) | 6251 | 0.013 | -0.0961 | No |
| 193 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 6264 | 0.013 | -0.0969 | No |
| 194 | ARHGEF5 | ARHGEF5 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 5 | 6309 | 0.012 | -0.1002 | No |
| 195 | TCEA1 | TCEA1 Entrez, Source | transcription elongation factor A (SII), 1 | 6329 | 0.011 | -0.1015 | No |
| 196 | PLN | PLN Entrez, Source | phospholamban | 6355 | 0.011 | -0.1033 | No |
| 197 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 6356 | 0.010 | -0.1032 | No |
| 198 | STRAP | STRAP Entrez, Source | serine/threonine kinase receptor associated protein | 6387 | 0.010 | -0.1054 | No |
| 199 | EPS15 | EPS15 Entrez, Source | epidermal growth factor receptor pathway substrate 15 | 6394 | 0.010 | -0.1058 | No |
| 200 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 6420 | 0.009 | -0.1076 | No |
| 201 | STCH | STCH Entrez, Source | stress 70 protein chaperone, microsome-associated, 60kDa | 6424 | 0.009 | -0.1077 | No |
| 202 | VPS4A | VPS4A Entrez, Source | vacuolar protein sorting 4 homolog A (S. cerevisiae) | 6478 | 0.008 | -0.1117 | No |
| 203 | ENG | ENG Entrez, Source | endoglin (Osler-Rendu-Weber syndrome 1) | 6481 | 0.008 | -0.1118 | No |
| 204 | NOX4 | NOX4 Entrez, Source | NADPH oxidase 4 | 6694 | 0.004 | -0.1282 | No |
| 205 | SLC1A4 | SLC1A4 Entrez, Source | solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 | 6704 | 0.004 | -0.1289 | No |
| 206 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 6727 | 0.004 | -0.1305 | No |
| 207 | ABCB8 | ABCB8 Entrez, Source | ATP-binding cassette, sub-family B (MDR/TAP), member 8 | 6773 | 0.003 | -0.1340 | No |
| 208 | IL11 | IL11 Entrez, Source | interleukin 11 | 6783 | 0.003 | -0.1347 | No |
| 209 | COPS3 | COPS3 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 3 (Arabidopsis) | 6803 | 0.002 | -0.1361 | No |
| 210 | ARID5A | ARID5A Entrez, Source | AT rich interactive domain 5A (MRF1-like) | 6828 | 0.002 | -0.1379 | No |
| 211 | GABRB1 | GABRB1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 1 | 6864 | 0.001 | -0.1406 | No |
| 212 | MAP2K2 | MAP2K2 Entrez, Source | mitogen-activated protein kinase kinase 2 | 6934 | -0.001 | -0.1460 | No |
| 213 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 6978 | -0.002 | -0.1493 | No |
| 214 | SEC14L2 | SEC14L2 Entrez, Source | SEC14-like 2 (S. cerevisiae) | 7014 | -0.002 | -0.1520 | No |
| 215 | AKT3 | AKT3 Entrez, Source | v-akt murine thymoma viral oncogene homolog 3 (protein kinase B, gamma) | 7024 | -0.003 | -0.1527 | No |
| 216 | TUSC4 | TUSC4 Entrez, Source | tumor suppressor candidate 4 | 7068 | -0.003 | -0.1560 | No |
| 217 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 7069 | -0.004 | -0.1559 | No |
| 218 | PDE5A | PDE5A Entrez, Source | phosphodiesterase 5A, cGMP-specific | 7087 | -0.004 | -0.1572 | No |
| 219 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 7106 | -0.004 | -0.1586 | No |
| 220 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 7114 | -0.004 | -0.1590 | No |
| 221 | SERTAD1 | SERTAD1 Entrez, Source | SERTA domain containing 1 | 7125 | -0.005 | -0.1598 | No |
| 222 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 7149 | -0.005 | -0.1615 | No |
| 223 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 7160 | -0.005 | -0.1622 | No |
| 224 | NINJ2 | NINJ2 Entrez, Source | ninjurin 2 | 7217 | -0.007 | -0.1665 | No |
| 225 | SEC23IP | SEC23IP Entrez, Source | SEC23 interacting protein | 7260 | -0.008 | -0.1696 | No |
| 226 | KCNA3 | KCNA3 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 3 | 7275 | -0.008 | -0.1706 | No |
| 227 | CCNH | CCNH Entrez, Source | cyclin H | 7280 | -0.008 | -0.1708 | No |
| 228 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 7310 | -0.008 | -0.1730 | No |
| 229 | RALBP1 | RALBP1 Entrez, Source | ralA binding protein 1 | 7318 | -0.009 | -0.1734 | No |
| 230 | COL1A1 | COL1A1 Entrez, Source | collagen, type I, alpha 1 | 7328 | -0.009 | -0.1740 | No |
| 231 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 7375 | -0.010 | -0.1775 | No |
| 232 | EFNB1 | EFNB1 Entrez, Source | ephrin-B1 | 7378 | -0.010 | -0.1775 | No |
| 233 | GOLPH4 | GOLPH4 Entrez, Source | golgi phosphoprotein 4 | 7438 | -0.011 | -0.1820 | No |
| 234 | WDR39 | WDR39 Entrez, Source | WD repeat domain 39 | 7439 | -0.011 | -0.1818 | No |
| 235 | NUP50 | NUP50 Entrez, Source | nucleoporin 50kDa | 7554 | -0.014 | -0.1905 | No |
| 236 | CDKN2B | CDKN2B Entrez, Source | cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4) | 7589 | -0.014 | -0.1930 | No |
| 237 | LTBP2 | LTBP2 Entrez, Source | latent transforming growth factor beta binding protein 2 | 7613 | -0.015 | -0.1946 | No |
| 238 | XRCC4 | XRCC4 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 4 | 7672 | -0.016 | -0.1989 | No |
| 239 | FBLN5 | FBLN5 Entrez, Source | fibulin 5 | 7698 | -0.017 | -0.2006 | No |
| 240 | SST | SST Entrez, Source | somatostatin | 7737 | -0.017 | -0.2034 | No |
| 241 | IVL | IVL Entrez, Source | involucrin | 7745 | -0.018 | -0.2037 | No |
| 242 | PKIG | PKIG Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor gamma | 7811 | -0.019 | -0.2085 | No |
| 243 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 7849 | -0.020 | -0.2112 | No |
| 244 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7856 | -0.020 | -0.2114 | No |
| 245 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 7859 | -0.020 | -0.2113 | No |
| 246 | BMP2K | BMP2K Entrez, Source | BMP2 inducible kinase | 7875 | -0.020 | -0.2122 | No |
| 247 | EIF2C2 | EIF2C2 Entrez, Source | eukaryotic translation initiation factor 2C, 2 | 7880 | -0.021 | -0.2123 | No |
| 248 | PPIA | PPIA Entrez, Source | peptidylprolyl isomerase A (cyclophilin A) | 8000 | -0.023 | -0.2212 | No |
| 249 | DKK3 | DKK3 Entrez, Source | dickkopf homolog 3 (Xenopus laevis) | 8061 | -0.025 | -0.2256 | No |
| 250 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 8124 | -0.026 | -0.2301 | No |
| 251 | RPL4 | RPL4 Entrez, Source | ribosomal protein L4 | 8146 | -0.026 | -0.2314 | No |
| 252 | KIF4A | KIF4A Entrez, Source | kinesin family member 4A | 8274 | -0.029 | -0.2409 | No |
| 253 | TINF2 | TINF2 Entrez, Source | TERF1 (TRF1)-interacting nuclear factor 2 | 8295 | -0.030 | -0.2421 | No |
| 254 | NAV2 | NAV2 Entrez, Source | neuron navigator 2 | 8365 | -0.032 | -0.2471 | No |
| 255 | AXIN1 | AXIN1 Entrez, Source | axin 1 | 8385 | -0.032 | -0.2481 | No |
| 256 | TMEM49 | TMEM49 Entrez, Source | transmembrane protein 49 | 8469 | -0.034 | -0.2542 | No |
| 257 | GAS6 | GAS6 Entrez, Source | growth arrest-specific 6 | 8485 | -0.034 | -0.2549 | No |
| 258 | RPL5 | RPL5 Entrez, Source | ribosomal protein L5 | 8527 | -0.035 | -0.2577 | No |
| 259 | TOM1 | TOM1 Entrez, Source | target of myb1 (chicken) | 8538 | -0.035 | -0.2580 | No |
| 260 | TCF8 | TCF8 Entrez, Source | transcription factor 8 (represses interleukin 2 expression) | 8543 | -0.036 | -0.2579 | No |
| 261 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 8548 | -0.036 | -0.2578 | No |
| 262 | COL15A1 | COL15A1 Entrez, Source | collagen, type XV, alpha 1 | 8592 | -0.037 | -0.2607 | No |
| 263 | CDH6 | CDH6 Entrez, Source | cadherin 6, type 2, K-cadherin (fetal kidney) | 8608 | -0.037 | -0.2614 | No |
| 264 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 8626 | -0.037 | -0.2622 | No |
| 265 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 8677 | -0.038 | -0.2657 | No |
| 266 | CELSR3 | CELSR3 Entrez, Source | cadherin, EGF LAG seven-pass G-type receptor 3 (flamingo homolog, Drosophila) | 8696 | -0.039 | -0.2666 | No |
| 267 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 8700 | -0.039 | -0.2663 | No |
| 268 | POU6F1 | POU6F1 Entrez, Source | POU domain, class 6, transcription factor 1 | 8710 | -0.039 | -0.2666 | No |
| 269 | TGM4 | TGM4 Entrez, Source | transglutaminase 4 (prostate) | 8762 | -0.040 | -0.2700 | No |
| 270 | SEC22A | SEC22A Entrez, Source | SEC22 vesicle trafficking protein homolog A (S. cerevisiae) | 8808 | -0.041 | -0.2730 | No |
| 271 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 8842 | -0.042 | -0.2751 | No |
| 272 | STAT5B | STAT5B Entrez, Source | signal transducer and activator of transcription 5B | 8858 | -0.043 | -0.2757 | No |
| 273 | MAPKAPK2 | MAPKAPK2 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 2 | 8907 | -0.044 | -0.2789 | No |
| 274 | GATA4 | GATA4 Entrez, Source | GATA binding protein 4 | 8964 | -0.045 | -0.2827 | No |
| 275 | YWHAE | YWHAE Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide | 8995 | -0.046 | -0.2845 | No |
| 276 | TNFSF4 | TNFSF4 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 4 (tax-transcriptionally activated glycoprotein 1, 34kDa) | 9004 | -0.046 | -0.2845 | No |
| 277 | KCNE3 | KCNE3 Entrez, Source | potassium voltage-gated channel, Isk-related family, member 3 | 9024 | -0.046 | -0.2854 | No |
| 278 | ADAM19 | ADAM19 Entrez, Source | ADAM metallopeptidase domain 19 (meltrin beta) | 9048 | -0.047 | -0.2867 | No |
| 279 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 9075 | -0.048 | -0.2881 | No |
| 280 | GFPT2 | GFPT2 Entrez, Source | glutamine-fructose-6-phosphate transaminase 2 | 9121 | -0.049 | -0.2910 | No |
| 281 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 9124 | -0.049 | -0.2905 | No |
| 282 | PSMB6 | PSMB6 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 6 | 9235 | -0.052 | -0.2984 | No |
| 283 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 9236 | -0.052 | -0.2978 | No |
| 284 | RPS4X | RPS4X Entrez, Source | ribosomal protein S4, X-linked | 9239 | -0.052 | -0.2973 | No |
| 285 | PMP22 | PMP22 Entrez, Source | peripheral myelin protein 22 | 9250 | -0.053 | -0.2974 | No |
| 286 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 9345 | -0.056 | -0.3041 | No |
| 287 | BPGM | BPGM Entrez, Source | 2,3-bisphosphoglycerate mutase | 9399 | -0.057 | -0.3075 | No |
| 288 | BMP1 | BMP1 Entrez, Source | bone morphogenetic protein 1 | 9411 | -0.057 | -0.3076 | No |
| 289 | PTPN11 | PTPN11 Entrez, Source | protein tyrosine phosphatase, non-receptor type 11 (Noonan syndrome 1) | 9419 | -0.058 | -0.3075 | No |
| 290 | IGF2BP1 | IGF2BP1 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 1 | 9422 | -0.058 | -0.3069 | No |
| 291 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 9477 | -0.059 | -0.3104 | No |
| 292 | ARPC5 | ARPC5 Entrez, Source | actin related protein 2/3 complex, subunit 5, 16kDa | 9496 | -0.060 | -0.3111 | No |
| 293 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 9515 | -0.060 | -0.3117 | No |
| 294 | RAN | RAN Entrez, Source | RAN, member RAS oncogene family | 9534 | -0.061 | -0.3124 | No |
| 295 | SLC29A1 | SLC29A1 Entrez, Source | solute carrier family 29 (nucleoside transporters), member 1 | 9613 | -0.063 | -0.3177 | No |
| 296 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 9628 | -0.063 | -0.3180 | No |
| 297 | PPP2R1A | PPP2R1A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), alpha isoform | 9656 | -0.064 | -0.3193 | No |
| 298 | CCRK | CCRK Entrez, Source | cell cycle related kinase | 9684 | -0.065 | -0.3206 | No |
| 299 | APH1A | APH1A Entrez, Source | anterior pharynx defective 1 homolog A (C. elegans) | 9742 | -0.066 | -0.3242 | No |
| 300 | JUN | JUN Entrez, Source | jun oncogene | 9748 | -0.066 | -0.3238 | No |
| 301 | GTF2B | GTF2B Entrez, Source | general transcription factor IIB | 9753 | -0.067 | -0.3233 | No |
| 302 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 9762 | -0.067 | -0.3231 | No |
| 303 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 9782 | -0.067 | -0.3237 | No |
| 304 | CTBP1 | CTBP1 Entrez, Source | C-terminal binding protein 1 | 9803 | -0.068 | -0.3245 | No |
| 305 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 9897 | -0.071 | -0.3308 | No |
| 306 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 9912 | -0.072 | -0.3310 | No |
| 307 | FN1 | FN1 Entrez, Source | fibronectin 1 | 9917 | -0.072 | -0.3304 | No |
| 308 | SPINT2 | SPINT2 Entrez, Source | serine peptidase inhibitor, Kunitz type, 2 | 9927 | -0.073 | -0.3302 | No |
| 309 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 10036 | -0.076 | -0.3377 | No |
| 310 | MAPK3 | MAPK3 Entrez, Source | mitogen-activated protein kinase 3 | 10053 | -0.076 | -0.3380 | No |
| 311 | RAB4B | RAB4B Entrez, Source | RAB4B, member RAS oncogene family | 10064 | -0.077 | -0.3379 | No |
| 312 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 10069 | -0.077 | -0.3372 | No |
| 313 | HSF2 | HSF2 Entrez, Source | heat shock transcription factor 2 | 10081 | -0.077 | -0.3371 | No |
| 314 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 10098 | -0.078 | -0.3374 | No |
| 315 | BAG5 | BAG5 Entrez, Source | BCL2-associated athanogene 5 | 10166 | -0.080 | -0.3416 | No |
| 316 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 10171 | -0.080 | -0.3410 | No |
| 317 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 10198 | -0.081 | -0.3420 | No |
| 318 | CDIPT | CDIPT Entrez, Source | CDP-diacylglycerol--inositol 3-phosphatidyltransferase (phosphatidylinositol synthase) | 10202 | -0.081 | -0.3412 | No |
| 319 | FUT4 | FUT4 Entrez, Source | fucosyltransferase 4 (alpha (1,3) fucosyltransferase, myeloid-specific) | 10213 | -0.082 | -0.3410 | No |
| 320 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 10317 | -0.085 | -0.3480 | No |
| 321 | MAPK7 | MAPK7 Entrez, Source | mitogen-activated protein kinase 7 | 10319 | -0.085 | -0.3470 | No |
| 322 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10323 | -0.086 | -0.3462 | No |
| 323 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 10376 | -0.087 | -0.3492 | No |
| 324 | GRB7 | GRB7 Entrez, Source | growth factor receptor-bound protein 7 | 10441 | -0.090 | -0.3530 | No |
| 325 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 10545 | -0.093 | -0.3599 | No |
| 326 | ACTB | ACTB Entrez, Source | actin, beta | 10590 | -0.095 | -0.3621 | No |
| 327 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 10604 | -0.095 | -0.3620 | No |
| 328 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 10651 | -0.097 | -0.3644 | No |
| 329 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 10694 | -0.098 | -0.3664 | No |
| 330 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 10722 | -0.099 | -0.3673 | No |
| 331 | PPM1F | PPM1F Entrez, Source | protein phosphatase 1F (PP2C domain containing) | 10745 | -0.100 | -0.3678 | No |
| 332 | STK25 | STK25 Entrez, Source | serine/threonine kinase 25 (STE20 homolog, yeast) | 10787 | -0.102 | -0.3697 | No |
| 333 | RGL2 | RGL2 Entrez, Source | ral guanine nucleotide dissociation stimulator-like 2 | 10808 | -0.103 | -0.3700 | No |
| 334 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 10825 | -0.104 | -0.3700 | No |
| 335 | BGN | BGN Entrez, Source | biglycan | 10918 | -0.108 | -0.3758 | No |
| 336 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 10997 | -0.112 | -0.3805 | No |
| 337 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 10998 | -0.112 | -0.3792 | No |
| 338 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 11000 | -0.112 | -0.3779 | No |
| 339 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 11027 | -0.113 | -0.3785 | No |
| 340 | PDGFRL | PDGFRL Entrez, Source | platelet-derived growth factor receptor-like | 11031 | -0.113 | -0.3773 | No |
| 341 | MAP2K1 | MAP2K1 Entrez, Source | mitogen-activated protein kinase kinase 1 | 11069 | -0.115 | -0.3788 | No |
| 342 | PRKD2 | PRKD2 Entrez, Source | protein kinase D2 | 11081 | -0.116 | -0.3783 | No |
| 343 | LSP1 | LSP1 Entrez, Source | lymphocyte-specific protein 1 | 11156 | -0.119 | -0.3825 | No |
| 344 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 11173 | -0.120 | -0.3823 | No |
| 345 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 11178 | -0.120 | -0.3812 | No |
| 346 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 11180 | -0.120 | -0.3798 | No |
| 347 | RHOA | RHOA Entrez, Source | ras homolog gene family, member A | 11190 | -0.121 | -0.3790 | No |
| 348 | AMIGO2 | AMIGO2 Entrez, Source | adhesion molecule with Ig-like domain 2 | 11194 | -0.121 | -0.3777 | No |
| 349 | SPON2 | SPON2 Entrez, Source | spondin 2, extracellular matrix protein | 11200 | -0.121 | -0.3766 | No |
| 350 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 11212 | -0.122 | -0.3760 | No |
| 351 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 11278 | -0.125 | -0.3795 | No |
| 352 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 11280 | -0.126 | -0.3781 | No |
| 353 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 11298 | -0.126 | -0.3778 | No |
| 354 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 11316 | -0.127 | -0.3776 | No |
| 355 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 11352 | -0.129 | -0.3787 | No |
| 356 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 11363 | -0.129 | -0.3779 | No |
| 357 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 11420 | -0.132 | -0.3807 | No |
| 358 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 11444 | -0.134 | -0.3808 | No |
| 359 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 11475 | -0.136 | -0.3815 | No |
| 360 | MBD1 | MBD1 Entrez, Source | methyl-CpG binding domain protein 1 | 11486 | -0.137 | -0.3806 | No |
| 361 | TGFBR2 | TGFBR2 Entrez, Source | transforming growth factor, beta receptor II (70/80kDa) | 11501 | -0.137 | -0.3800 | No |
| 362 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 11523 | -0.139 | -0.3799 | No |
| 363 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11527 | -0.139 | -0.3785 | No |
| 364 | TSPAN2 | TSPAN2 Entrez, Source | tetraspanin 2 | 11553 | -0.140 | -0.3787 | No |
| 365 | EHMT2 | EHMT2 Entrez, Source | euchromatic histone-lysine N-methyltransferase 2 | 11568 | -0.141 | -0.3780 | No |
| 366 | ITGB6 | ITGB6 Entrez, Source | integrin, beta 6 | 11573 | -0.142 | -0.3766 | No |
| 367 | ACTR3 | ACTR3 Entrez, Source | ARP3 actin-related protein 3 homolog (yeast) | 11578 | -0.142 | -0.3752 | No |
| 368 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 11599 | -0.144 | -0.3750 | No |
| 369 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 11673 | -0.148 | -0.3788 | No |
| 370 | PAWR | PAWR Entrez, Source | PRKC, apoptosis, WT1, regulator | 11680 | -0.149 | -0.3775 | No |
| 371 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 11690 | -0.149 | -0.3764 | No |
| 372 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 11694 | -0.149 | -0.3748 | No |
| 373 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 11695 | -0.149 | -0.3729 | No |
| 374 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 11744 | -0.152 | -0.3748 | No |
| 375 | CLU | CLU Entrez, Source | clusterin | 11779 | -0.155 | -0.3756 | No |
| 376 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 11783 | -0.155 | -0.3739 | No |
| 377 | ACTG2 | ACTG2 Entrez, Source | actin, gamma 2, smooth muscle, enteric | 11806 | -0.157 | -0.3737 | No |
| 378 | ITGB4 | ITGB4 Entrez, Source | integrin, beta 4 | 11844 | -0.160 | -0.3746 | No |
| 379 | PCOLCE | PCOLCE Entrez, Source | procollagen C-endopeptidase enhancer | 11861 | -0.161 | -0.3739 | No |
| 380 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 11900 | -0.165 | -0.3748 | No |
| 381 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 11998 | -0.173 | -0.3802 | No |
| 382 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 11999 | -0.173 | -0.3781 | No |
| 383 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12039 | -0.176 | -0.3790 | No |
| 384 | FST | FST Entrez, Source | follistatin | 12041 | -0.177 | -0.3769 | No |
| 385 | PTPRF | PTPRF Entrez, Source | protein tyrosine phosphatase, receptor type, F | 12043 | -0.177 | -0.3748 | No |
| 386 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 12047 | -0.177 | -0.3729 | No |
| 387 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 12163 | -0.188 | -0.3795 | No |
| 388 | CBFB | CBFB Entrez, Source | core-binding factor, beta subunit | 12189 | -0.190 | -0.3791 | No |
| 389 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 12216 | -0.193 | -0.3788 | No |
| 390 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 12251 | -0.199 | -0.3790 | No |
| 391 | JUP | JUP Entrez, Source | junction plakoglobin | 12351 | -0.210 | -0.3841 | No |
| 392 | ST14 | ST14 Entrez, Source | suppression of tumorigenicity 14 (colon carcinoma) | 12479 | -0.224 | -0.3912 | Yes |
| 393 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 12496 | -0.228 | -0.3897 | Yes |
| 394 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 12522 | -0.231 | -0.3888 | Yes |
| 395 | TGFB1I1 | TGFB1I1 Entrez, Source | transforming growth factor beta 1 induced transcript 1 | 12535 | -0.233 | -0.3868 | Yes |
| 396 | TGFBR1 | TGFBR1 Entrez, Source | transforming growth factor, beta receptor I (activin A receptor type II-like kinase, 53kDa) | 12538 | -0.234 | -0.3841 | Yes |
| 397 | FSTL3 | FSTL3 Entrez, Source | follistatin-like 3 (secreted glycoprotein) | 12563 | -0.238 | -0.3831 | Yes |
| 398 | MYH9 | MYH9 Entrez, Source | myosin, heavy chain 9, non-muscle | 12585 | -0.241 | -0.3818 | Yes |
| 399 | S100A11 | S100A11 Entrez, Source | S100 calcium binding protein A11 | 12634 | -0.251 | -0.3824 | Yes |
| 400 | HK1 | HK1 Entrez, Source | hexokinase 1 | 12636 | -0.251 | -0.3794 | Yes |
| 401 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 12640 | -0.252 | -0.3766 | Yes |
| 402 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 12644 | -0.253 | -0.3737 | Yes |
| 403 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 12652 | -0.254 | -0.3712 | Yes |
| 404 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 12659 | -0.255 | -0.3685 | Yes |
| 405 | PLAU | PLAU Entrez, Source | plasminogen activator, urokinase | 12662 | -0.256 | -0.3655 | Yes |
| 406 | CLCF1 | CLCF1 Entrez, Source | cardiotrophin-like cytokine factor 1 | 12679 | -0.260 | -0.3636 | Yes |
| 407 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 12698 | -0.263 | -0.3618 | Yes |
| 408 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 12714 | -0.267 | -0.3597 | Yes |
| 409 | CD151 | CD151 Entrez, Source | CD151 molecule (Raph blood group) | 12718 | -0.268 | -0.3566 | Yes |
| 410 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 12719 | -0.269 | -0.3533 | Yes |
| 411 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 12725 | -0.270 | -0.3504 | Yes |
| 412 | SLC12A7 | SLC12A7 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 7 | 12774 | -0.279 | -0.3507 | Yes |
| 413 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12786 | -0.282 | -0.3481 | Yes |
| 414 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 12818 | -0.293 | -0.3469 | Yes |
| 415 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 12826 | -0.295 | -0.3439 | Yes |
| 416 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 12832 | -0.296 | -0.3406 | Yes |
| 417 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 12838 | -0.299 | -0.3374 | Yes |
| 418 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 12840 | -0.300 | -0.3338 | Yes |
| 419 | RAI14 | RAI14 Entrez, Source | retinoic acid induced 14 | 12860 | -0.306 | -0.3315 | Yes |
| 420 | HSPB1 | HSPB1 Entrez, Source | heat shock 27kDa protein 1 | 12892 | -0.318 | -0.3300 | Yes |
| 421 | IFI35 | IFI35 Entrez, Source | interferon-induced protein 35 | 12894 | -0.318 | -0.3262 | Yes |
| 422 | GRN | GRN Entrez, Source | granulin | 12920 | -0.329 | -0.3241 | Yes |
| 423 | CKLF | CKLF Entrez, Source | chemokine-like factor | 12938 | -0.335 | -0.3214 | Yes |
| 424 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 12949 | -0.338 | -0.3180 | Yes |
| 425 | TUBB | TUBB Entrez, Source | tubulin, beta | 12961 | -0.344 | -0.3146 | Yes |
| 426 | CKB | CKB Entrez, Source | creatine kinase, brain | 12974 | -0.349 | -0.3113 | Yes |
| 427 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 12986 | -0.357 | -0.3078 | Yes |
| 428 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 12997 | -0.362 | -0.3041 | Yes |
| 429 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 12998 | -0.364 | -0.2997 | Yes |
| 430 | PODXL | PODXL Entrez, Source | podocalyxin-like | 13001 | -0.365 | -0.2954 | Yes |
| 431 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 13016 | -0.374 | -0.2919 | Yes |
| 432 | HIVEP2 | HIVEP2 Entrez, Source | human immunodeficiency virus type I enhancer binding protein 2 | 13020 | -0.376 | -0.2875 | Yes |
| 433 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 13032 | -0.382 | -0.2837 | Yes |
| 434 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 13047 | -0.395 | -0.2799 | Yes |
| 435 | EPHB6 | EPHB6 Entrez, Source | EPH receptor B6 | 13050 | -0.399 | -0.2752 | Yes |
| 436 | TSPAN6 | TSPAN6 Entrez, Source | tetraspanin 6 | 13052 | -0.400 | -0.2704 | Yes |
| 437 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 13064 | -0.406 | -0.2663 | Yes |
| 438 | JAK2 | JAK2 Entrez, Source | Janus kinase 2 (a protein tyrosine kinase) | 13065 | -0.406 | -0.2613 | Yes |
| 439 | GPC1 | GPC1 Entrez, Source | glypican 1 | 13081 | -0.412 | -0.2574 | Yes |
| 440 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 13094 | -0.421 | -0.2532 | Yes |
| 441 | SPRY2 | SPRY2 Entrez, Source | sprouty homolog 2 (Drosophila) | 13095 | -0.423 | -0.2480 | Yes |
| 442 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 13098 | -0.424 | -0.2430 | Yes |
| 443 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 13099 | -0.425 | -0.2378 | Yes |
| 444 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 13120 | -0.443 | -0.2339 | Yes |
| 445 | LMO7 | LMO7 Entrez, Source | LIM domain 7 | 13121 | -0.443 | -0.2285 | Yes |
| 446 | CNN1 | CNN1 Entrez, Source | calponin 1, basic, smooth muscle | 13134 | -0.456 | -0.2239 | Yes |
| 447 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13162 | -0.500 | -0.2198 | Yes |
| 448 | VIM | VIM Entrez, Source | vimentin | 13164 | -0.501 | -0.2138 | Yes |
| 449 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 13183 | -0.530 | -0.2087 | Yes |
| 450 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 13196 | -0.560 | -0.2028 | Yes |
| 451 | PPIC | PPIC Entrez, Source | peptidylprolyl isomerase C (cyclophilin C) | 13206 | -0.575 | -0.1964 | Yes |
| 452 | TIMP1 | TIMP1 Entrez, Source | TIMP metallopeptidase inhibitor 1 | 13213 | -0.582 | -0.1898 | Yes |
| 453 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 13219 | -0.601 | -0.1828 | Yes |
| 454 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 13226 | -0.619 | -0.1757 | Yes |
| 455 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13247 | -0.677 | -0.1690 | Yes |
| 456 | CST6 | CST6 Entrez, Source | cystatin E/M | 13263 | -0.730 | -0.1612 | Yes |
| 457 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 13269 | -0.760 | -0.1523 | Yes |
| 458 | MMP11 | MMP11 Entrez, Source | matrix metallopeptidase 11 (stromelysin 3) | 13273 | -0.768 | -0.1431 | Yes |
| 459 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 13277 | -0.783 | -0.1338 | Yes |
| 460 | HS3ST2 | HS3ST2 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 2 | 13280 | -0.791 | -0.1242 | Yes |
| 461 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 13287 | -0.812 | -0.1148 | Yes |
| 462 | LOX | LOX Entrez, Source | lysyl oxidase | 13308 | -0.991 | -0.1042 | Yes |
| 463 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 13309 | -0.997 | -0.0920 | Yes |
| 464 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 13312 | -1.018 | -0.0797 | Yes |
| 465 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 13316 | -1.048 | -0.0671 | Yes |
| 466 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13324 | -1.119 | -0.0539 | Yes |
| 467 | TAGLN | TAGLN Entrez, Source | transgelin | 13326 | -1.131 | -0.0402 | Yes |
| 468 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 13340 | -1.581 | -0.0218 | Yes |
| 469 | CPE | CPE Entrez, Source | carboxypeptidase E | 13342 | -1.792 | -0.0000 | Yes |