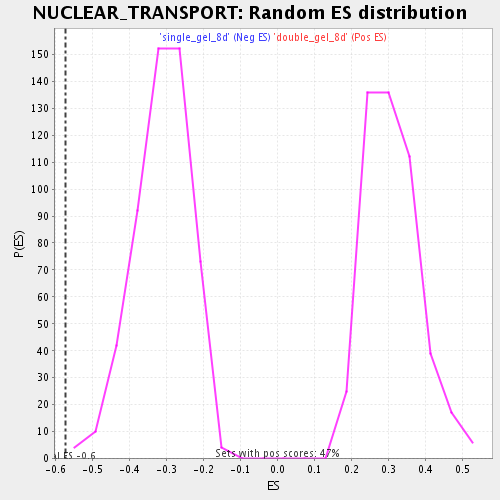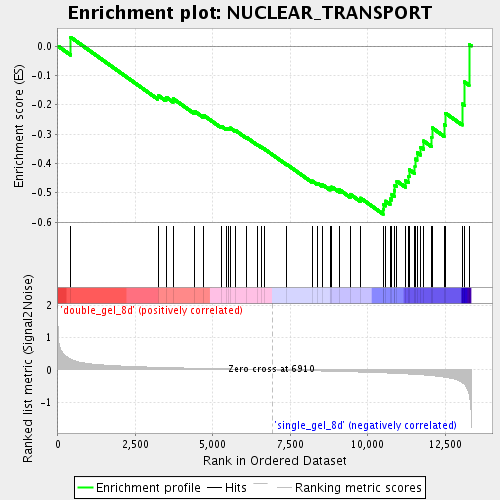
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | NUCLEAR_TRANSPORT |
| Enrichment Score (ES) | -0.5727093 |
| Normalized Enrichment Score (NES) | -1.8404652 |
| Nominal p-value | 0.0018903592 |
| FDR q-value | 0.03599247 |
| FWER p-Value | 0.764 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | F2 | F2 Entrez, Source | coagulation factor II (thrombin) | 413 | 0.335 | 0.0299 | No |
| 2 | PTTG1IP | PTTG1IP Entrez, Source | pituitary tumor-transforming 1 interacting protein | 3230 | 0.078 | -0.1678 | No |
| 3 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 3489 | 0.071 | -0.1742 | No |
| 4 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 3717 | 0.066 | -0.1792 | No |
| 5 | KPNA1 | KPNA1 Entrez, Source | karyopherin alpha 1 (importin alpha 5) | 4415 | 0.050 | -0.2225 | No |
| 6 | FAF1 | FAF1 Entrez, Source | Fas (TNFRSF6) associated factor 1 | 4712 | 0.044 | -0.2368 | No |
| 7 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 5278 | 0.032 | -0.2734 | No |
| 8 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 5443 | 0.029 | -0.2804 | No |
| 9 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 5506 | 0.028 | -0.2800 | No |
| 10 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 5558 | 0.026 | -0.2791 | No |
| 11 | NFKBIL1 | NFKBIL1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1 | 5719 | 0.023 | -0.2869 | No |
| 12 | GLI3 | GLI3 Entrez, Source | GLI-Kruppel family member GLI3 (Greig cephalopolysyndactyly syndrome) | 6095 | 0.016 | -0.3121 | No |
| 13 | DDX25 | DDX25 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 25 | 6455 | 0.009 | -0.3375 | No |
| 14 | CALR | CALR Entrez, Source | calreticulin | 6558 | 0.007 | -0.3440 | No |
| 15 | UHMK1 | UHMK1 Entrez, Source | U2AF homology motif (UHM) kinase 1 | 6651 | 0.005 | -0.3500 | No |
| 16 | XPO6 | XPO6 Entrez, Source | exportin 6 | 7390 | -0.010 | -0.4036 | No |
| 17 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 8204 | -0.028 | -0.4597 | No |
| 18 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 8376 | -0.032 | -0.4668 | No |
| 19 | MYBBP1A | MYBBP1A Entrez, Source | MYB binding protein (P160) 1a | 8546 | -0.036 | -0.4730 | No |
| 20 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 8782 | -0.040 | -0.4833 | No |
| 21 | RAE1 | RAE1 Entrez, Source | RAE1 RNA export 1 homolog (S. pombe) | 8844 | -0.042 | -0.4803 | No |
| 22 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 9082 | -0.048 | -0.4893 | No |
| 23 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 9435 | -0.058 | -0.5052 | No |
| 24 | XPO7 | XPO7 Entrez, Source | exportin 7 | 9765 | -0.067 | -0.5178 | No |
| 25 | TPR | TPR Entrez, Source | translocated promoter region (to activated MET oncogene) | 10496 | -0.091 | -0.5561 | Yes |
| 26 | NUP107 | NUP107 Entrez, Source | nucleoporin 107kDa | 10509 | -0.092 | -0.5403 | Yes |
| 27 | KPNA3 | KPNA3 Entrez, Source | karyopherin alpha 3 (importin alpha 4) | 10575 | -0.094 | -0.5281 | Yes |
| 28 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 10722 | -0.099 | -0.5210 | Yes |
| 29 | DDX39 | DDX39 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 | 10777 | -0.102 | -0.5065 | Yes |
| 30 | NUDT4 | NUDT4 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 4 | 10857 | -0.105 | -0.4933 | Yes |
| 31 | DDX19B | DDX19B Entrez, Source | DEAD (Asp-Glu-Ala-As) box polypeptide 19B | 10871 | -0.106 | -0.4750 | Yes |
| 32 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 10938 | -0.109 | -0.4602 | Yes |
| 33 | PDIA3 | PDIA3 Entrez, Source | protein disulfide isomerase family A, member 3 | 11217 | -0.122 | -0.4589 | Yes |
| 34 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 11321 | -0.127 | -0.4434 | Yes |
| 35 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 11345 | -0.129 | -0.4217 | Yes |
| 36 | KPNA4 | KPNA4 Entrez, Source | karyopherin alpha 4 (importin alpha 3) | 11522 | -0.139 | -0.4097 | Yes |
| 37 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11527 | -0.139 | -0.3847 | Yes |
| 38 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 11590 | -0.143 | -0.3634 | Yes |
| 39 | TBRG1 | TBRG1 Entrez, Source | transforming growth factor beta regulator 1 | 11701 | -0.150 | -0.3444 | Yes |
| 40 | AKT1 | AKT1 Entrez, Source | v-akt murine thymoma viral oncogene homolog 1 | 11801 | -0.156 | -0.3234 | Yes |
| 41 | KHDRBS1 | KHDRBS1 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 1 | 12056 | -0.178 | -0.3101 | Yes |
| 42 | TNPO1 | TNPO1 Entrez, Source | transportin 1 | 12075 | -0.179 | -0.2788 | Yes |
| 43 | RERE | RERE Entrez, Source | arginine-glutamic acid dipeptide (RE) repeats | 12466 | -0.223 | -0.2675 | Yes |
| 44 | LYK5 | LYK5 Entrez, Source | - | 12506 | -0.230 | -0.2286 | Yes |
| 45 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 13047 | -0.395 | -0.1973 | Yes |
| 46 | CEP57 | CEP57 Entrez, Source | centrosomal protein 57kDa | 13119 | -0.442 | -0.1221 | Yes |
| 47 | F2R | F2R Entrez, Source | coagulation factor II (thrombin) receptor | 13270 | -0.762 | 0.0054 | Yes |