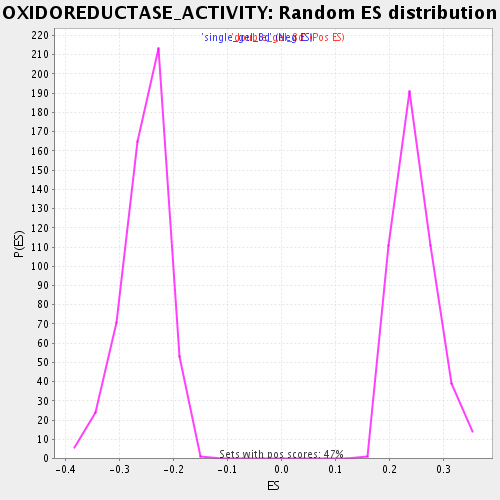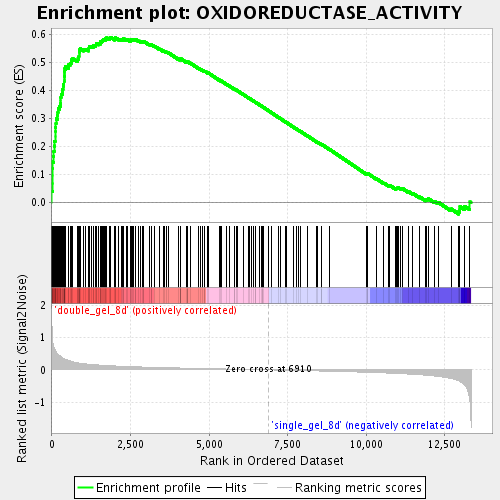
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | OXIDOREDUCTASE_ACTIVITY |
| Enrichment Score (ES) | 0.5889707 |
| Normalized Enrichment Score (NES) | 2.3803282 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 1 | 1.475 | 0.0413 | Yes |
| 2 | CYP8B1 | CYP8B1 Entrez, Source | cytochrome P450, family 8, subfamily B, polypeptide 1 | 14 | 1.030 | 0.0692 | Yes |
| 3 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 16 | 0.933 | 0.0953 | Yes |
| 4 | HGD | HGD Entrez, Source | homogentisate 1,2-dioxygenase (homogentisate oxidase) | 17 | 0.930 | 0.1213 | Yes |
| 5 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 24 | 0.864 | 0.1451 | Yes |
| 6 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 50 | 0.751 | 0.1642 | Yes |
| 7 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 60 | 0.721 | 0.1838 | Yes |
| 8 | HSD17B7 | HSD17B7 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 7 | 74 | 0.685 | 0.2020 | Yes |
| 9 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 84 | 0.641 | 0.2192 | Yes |
| 10 | MIOX | MIOX Entrez, Source | myo-inositol oxygenase | 103 | 0.612 | 0.2350 | Yes |
| 11 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 104 | 0.612 | 0.2522 | Yes |
| 12 | ABP1 | ABP1 Entrez, Source | amiloride binding protein 1 (amine oxidase (copper-containing)) | 109 | 0.606 | 0.2688 | Yes |
| 13 | DMGDH | DMGDH Entrez, Source | dimethylglycine dehydrogenase | 128 | 0.569 | 0.2834 | Yes |
| 14 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 135 | 0.556 | 0.2986 | Yes |
| 15 | KMO | KMO Entrez, Source | kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) | 174 | 0.497 | 0.3096 | Yes |
| 16 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 189 | 0.483 | 0.3221 | Yes |
| 17 | HAAO | HAAO Entrez, Source | 3-hydroxyanthranilate 3,4-dioxygenase | 196 | 0.479 | 0.3351 | Yes |
| 18 | SARDH | SARDH Entrez, Source | sarcosine dehydrogenase | 245 | 0.438 | 0.3437 | Yes |
| 19 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 271 | 0.419 | 0.3535 | Yes |
| 20 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 282 | 0.412 | 0.3643 | Yes |
| 21 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 284 | 0.409 | 0.3757 | Yes |
| 22 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 293 | 0.402 | 0.3863 | Yes |
| 23 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 333 | 0.375 | 0.3939 | Yes |
| 24 | SRD5A1 | SRD5A1 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 1 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 1) | 343 | 0.371 | 0.4036 | Yes |
| 25 | TDO2 | TDO2 Entrez, Source | tryptophan 2,3-dioxygenase | 355 | 0.360 | 0.4129 | Yes |
| 26 | ETFDH | ETFDH Entrez, Source | electron-transferring-flavoprotein dehydrogenase | 363 | 0.357 | 0.4223 | Yes |
| 27 | POR | POR Entrez, Source | P450 (cytochrome) oxidoreductase | 384 | 0.346 | 0.4305 | Yes |
| 28 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 389 | 0.344 | 0.4399 | Yes |
| 29 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 391 | 0.343 | 0.4494 | Yes |
| 30 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 401 | 0.338 | 0.4582 | Yes |
| 31 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 403 | 0.337 | 0.4676 | Yes |
| 32 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 411 | 0.336 | 0.4764 | Yes |
| 33 | MECR | MECR Entrez, Source | mitochondrial trans-2-enoyl-CoA reductase | 435 | 0.326 | 0.4838 | Yes |
| 34 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 524 | 0.292 | 0.4853 | Yes |
| 35 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 537 | 0.288 | 0.4925 | Yes |
| 36 | CYP27A1 | CYP27A1 Entrez, Source | cytochrome P450, family 27, subfamily A, polypeptide 1 | 578 | 0.274 | 0.4971 | Yes |
| 37 | SC5DL | SC5DL Entrez, Source | sterol-C5-desaturase (ERG3 delta-5-desaturase homolog, fungal)-like | 617 | 0.261 | 0.5015 | Yes |
| 38 | ADI1 | ADI1 Entrez, Source | acireductone dioxygenase 1 | 623 | 0.260 | 0.5084 | Yes |
| 39 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 663 | 0.250 | 0.5125 | Yes |
| 40 | COX4I2 | COX4I2 Entrez, Source | cytochrome c oxidase subunit IV isoform 2 (lung) | 811 | 0.219 | 0.5074 | Yes |
| 41 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 829 | 0.216 | 0.5122 | Yes |
| 42 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 849 | 0.214 | 0.5168 | Yes |
| 43 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 858 | 0.213 | 0.5221 | Yes |
| 44 | PDIA5 | PDIA5 Entrez, Source | protein disulfide isomerase family A, member 5 | 864 | 0.212 | 0.5277 | Yes |
| 45 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 865 | 0.212 | 0.5336 | Yes |
| 46 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 870 | 0.211 | 0.5392 | Yes |
| 47 | CAT | CAT Entrez, Source | catalase | 884 | 0.209 | 0.5441 | Yes |
| 48 | XDH | XDH Entrez, Source | xanthine dehydrogenase | 896 | 0.208 | 0.5491 | Yes |
| 49 | TXN2 | TXN2 Entrez, Source | thioredoxin 2 | 1019 | 0.190 | 0.5452 | Yes |
| 50 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 1074 | 0.185 | 0.5462 | Yes |
| 51 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1158 | 0.175 | 0.5448 | Yes |
| 52 | NQO2 | NQO2 Entrez, Source | NAD(P)H dehydrogenase, quinone 2 | 1174 | 0.173 | 0.5486 | Yes |
| 53 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 1180 | 0.172 | 0.5530 | Yes |
| 54 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 1183 | 0.172 | 0.5577 | Yes |
| 55 | PRDX3 | PRDX3 Entrez, Source | peroxiredoxin 3 | 1274 | 0.166 | 0.5555 | Yes |
| 56 | DECR1 | DECR1 Entrez, Source | 2,4-dienoyl CoA reductase 1, mitochondrial | 1315 | 0.163 | 0.5570 | Yes |
| 57 | MSRB2 | MSRB2 Entrez, Source | methionine sulfoxide reductase B2 | 1320 | 0.163 | 0.5613 | Yes |
| 58 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1399 | 0.157 | 0.5597 | Yes |
| 59 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 1403 | 0.157 | 0.5639 | Yes |
| 60 | SDHD | SDHD Entrez, Source | succinate dehydrogenase complex, subunit D, integral membrane protein | 1404 | 0.157 | 0.5683 | Yes |
| 61 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 1496 | 0.150 | 0.5656 | Yes |
| 62 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 1535 | 0.147 | 0.5668 | Yes |
| 63 | NOX1 | NOX1 Entrez, Source | NADPH oxidase 1 | 1538 | 0.147 | 0.5708 | Yes |
| 64 | HMOX2 | HMOX2 Entrez, Source | heme oxygenase (decycling) 2 | 1546 | 0.146 | 0.5743 | Yes |
| 65 | SUOX | SUOX Entrez, Source | sulfite oxidase | 1592 | 0.143 | 0.5749 | Yes |
| 66 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1615 | 0.142 | 0.5772 | Yes |
| 67 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 1628 | 0.141 | 0.5803 | Yes |
| 68 | BDH1 | BDH1 Entrez, Source | 3-hydroxybutyrate dehydrogenase, type 1 | 1686 | 0.138 | 0.5798 | Yes |
| 69 | EGLN1 | EGLN1 Entrez, Source | egl nine homolog 1 (C. elegans) | 1704 | 0.137 | 0.5824 | Yes |
| 70 | MAOB | MAOB Entrez, Source | monoamine oxidase B | 1712 | 0.137 | 0.5856 | Yes |
| 71 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 1747 | 0.134 | 0.5868 | Yes |
| 72 | NOS3 | NOS3 Entrez, Source | nitric oxide synthase 3 (endothelial cell) | 1840 | 0.129 | 0.5834 | Yes |
| 73 | COX7B | COX7B Entrez, Source | cytochrome c oxidase subunit VIIb | 1849 | 0.129 | 0.5865 | Yes |
| 74 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 1864 | 0.128 | 0.5890 | Yes |
| 75 | DHRS9 | DHRS9 Entrez, Source | dehydrogenase/reductase (SDR family) member 9 | 1998 | 0.122 | 0.5823 | No |
| 76 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 2018 | 0.121 | 0.5842 | No |
| 77 | ACADL | ACADL Entrez, Source | acyl-Coenzyme A dehydrogenase, long chain | 2024 | 0.121 | 0.5872 | No |
| 78 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 2134 | 0.117 | 0.5822 | No |
| 79 | TXNRD2 | TXNRD2 Entrez, Source | thioredoxin reductase 2 | 2200 | 0.114 | 0.5805 | No |
| 80 | IVD | IVD Entrez, Source | isovaleryl Coenzyme A dehydrogenase | 2248 | 0.113 | 0.5801 | No |
| 81 | CYGB | CYGB Entrez, Source | cytoglobin | 2265 | 0.112 | 0.5820 | No |
| 82 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 2281 | 0.111 | 0.5839 | No |
| 83 | MDH1 | MDH1 Entrez, Source | malate dehydrogenase 1, NAD (soluble) | 2361 | 0.108 | 0.5810 | No |
| 84 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 2402 | 0.107 | 0.5809 | No |
| 85 | EGLN2 | EGLN2 Entrez, Source | egl nine homolog 2 (C. elegans) | 2499 | 0.103 | 0.5765 | No |
| 86 | COX8A | COX8A Entrez, Source | cytochrome c oxidase subunit 8A (ubiquitous) | 2501 | 0.102 | 0.5793 | No |
| 87 | CP | CP Entrez, Source | ceruloplasmin (ferroxidase) | 2523 | 0.102 | 0.5805 | No |
| 88 | SPR | SPR Entrez, Source | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) | 2551 | 0.101 | 0.5813 | No |
| 89 | CYB5A | CYB5A Entrez, Source | cytochrome b5 type A (microsomal) | 2608 | 0.099 | 0.5798 | No |
| 90 | ETFA | ETFA Entrez, Source | electron-transfer-flavoprotein, alpha polypeptide (glutaric aciduria II) | 2650 | 0.098 | 0.5795 | No |
| 91 | NDUFS7 | NDUFS7 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 7, 20kDa (NADH-coenzyme Q reductase) | 2674 | 0.096 | 0.5804 | No |
| 92 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 2771 | 0.093 | 0.5757 | No |
| 93 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 2832 | 0.090 | 0.5737 | No |
| 94 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2886 | 0.088 | 0.5721 | No |
| 95 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 2909 | 0.088 | 0.5729 | No |
| 96 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 2926 | 0.087 | 0.5742 | No |
| 97 | SURF1 | SURF1 Entrez, Source | surfeit 1 | 3098 | 0.082 | 0.5634 | No |
| 98 | RDH10 | RDH10 Entrez, Source | retinol dehydrogenase 10 (all-trans) | 3161 | 0.080 | 0.5610 | No |
| 99 | GAPDHS | GAPDHS Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase, spermatogenic | 3168 | 0.079 | 0.5627 | No |
| 100 | AKR7A2 | AKR7A2 Entrez, Source | aldo-keto reductase family 7, member A2 (aflatoxin aldehyde reductase) | 3278 | 0.077 | 0.5566 | No |
| 101 | CYB5R2 | CYB5R2 Entrez, Source | cytochrome b5 reductase 2 | 3428 | 0.073 | 0.5473 | No |
| 102 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 3545 | 0.070 | 0.5405 | No |
| 103 | GLRX2 | GLRX2 Entrez, Source | glutaredoxin 2 | 3592 | 0.069 | 0.5389 | No |
| 104 | MTRR | MTRR Entrez, Source | 5-methyltetrahydrofolate-homocysteine methyltransferase reductase | 3648 | 0.068 | 0.5366 | No |
| 105 | LDHC | LDHC Entrez, Source | lactate dehydrogenase C | 3707 | 0.066 | 0.5341 | No |
| 106 | GSR | GSR Entrez, Source | glutathione reductase | 4018 | 0.059 | 0.5121 | No |
| 107 | FMO2 | FMO2 Entrez, Source | flavin containing monooxygenase 2 (non-functional) | 4078 | 0.058 | 0.5093 | No |
| 108 | CYP1A2 | CYP1A2 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 2 | 4085 | 0.058 | 0.5104 | No |
| 109 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 4097 | 0.057 | 0.5112 | No |
| 110 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 4100 | 0.057 | 0.5127 | No |
| 111 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 4291 | 0.053 | 0.4997 | No |
| 112 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 4305 | 0.053 | 0.5002 | No |
| 113 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4310 | 0.053 | 0.5014 | No |
| 114 | COX5B | COX5B Entrez, Source | cytochrome c oxidase subunit Vb | 4325 | 0.052 | 0.5018 | No |
| 115 | ALOX5 | ALOX5 Entrez, Source | arachidonate 5-lipoxygenase | 4403 | 0.050 | 0.4973 | No |
| 116 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 4665 | 0.045 | 0.4787 | No |
| 117 | ALOX15 | ALOX15 Entrez, Source | arachidonate 15-lipoxygenase | 4728 | 0.043 | 0.4752 | No |
| 118 | NOS1 | NOS1 Entrez, Source | nitric oxide synthase 1 (neuronal) | 4800 | 0.042 | 0.4710 | No |
| 119 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 4861 | 0.041 | 0.4676 | No |
| 120 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 4870 | 0.040 | 0.4681 | No |
| 121 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 4966 | 0.039 | 0.4620 | No |
| 122 | BLVRA | BLVRA Entrez, Source | biliverdin reductase A | 4973 | 0.038 | 0.4626 | No |
| 123 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 5320 | 0.032 | 0.4372 | No |
| 124 | COX4I1 | COX4I1 Entrez, Source | cytochrome c oxidase subunit IV isoform 1 | 5380 | 0.031 | 0.4335 | No |
| 125 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 5395 | 0.030 | 0.4333 | No |
| 126 | PRODH | PRODH Entrez, Source | proline dehydrogenase (oxidase) 1 | 5569 | 0.026 | 0.4209 | No |
| 127 | CYP26A1 | CYP26A1 Entrez, Source | cytochrome P450, family 26, subfamily A, polypeptide 1 | 5664 | 0.024 | 0.4145 | No |
| 128 | COX5A | COX5A Entrez, Source | cytochrome c oxidase subunit Va | 5800 | 0.022 | 0.4048 | No |
| 129 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 5822 | 0.022 | 0.4038 | No |
| 130 | CYP3A4 | CYP3A4 Entrez, Source | cytochrome P450, family 3, subfamily A, polypeptide 4 | 5860 | 0.021 | 0.4016 | No |
| 131 | AKR7A3 | AKR7A3 Entrez, Source | aldo-keto reductase family 7, member A3 (aflatoxin aldehyde reductase) | 5906 | 0.020 | 0.3987 | No |
| 132 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 6103 | 0.016 | 0.3843 | No |
| 133 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 6242 | 0.013 | 0.3742 | No |
| 134 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6279 | 0.012 | 0.3718 | No |
| 135 | DUOX2 | DUOX2 Entrez, Source | dual oxidase 2 | 6346 | 0.011 | 0.3671 | No |
| 136 | CRYZL1 | CRYZL1 Entrez, Source | crystallin, zeta (quinone reductase)-like 1 | 6417 | 0.009 | 0.3620 | No |
| 137 | NDUFS4 | NDUFS4 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) | 6477 | 0.008 | 0.3578 | No |
| 138 | SRD5A2 | SRD5A2 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 2 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 2) | 6493 | 0.008 | 0.3568 | No |
| 139 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 6602 | 0.006 | 0.3488 | No |
| 140 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 6665 | 0.005 | 0.3442 | No |
| 141 | NOX4 | NOX4 Entrez, Source | NADPH oxidase 4 | 6694 | 0.004 | 0.3422 | No |
| 142 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 6707 | 0.004 | 0.3414 | No |
| 143 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 6721 | 0.004 | 0.3405 | No |
| 144 | RETSAT | RETSAT Entrez, Source | retinol saturase (all-trans-retinol 13,14-reductase) | 6880 | 0.001 | 0.3285 | No |
| 145 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 6980 | -0.002 | 0.3211 | No |
| 146 | TXNL1 | TXNL1 Entrez, Source | thioredoxin-like 1 | 7208 | -0.006 | 0.3040 | No |
| 147 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 7268 | -0.008 | 0.2997 | No |
| 148 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 7432 | -0.011 | 0.2876 | No |
| 149 | TH | TH Entrez, Source | tyrosine hydroxylase | 7455 | -0.011 | 0.2863 | No |
| 150 | CYP11B1 | CYP11B1 Entrez, Source | cytochrome P450, family 11, subfamily B, polypeptide 1 | 7700 | -0.017 | 0.2682 | No |
| 151 | FMO1 | FMO1 Entrez, Source | flavin containing monooxygenase 1 | 7782 | -0.018 | 0.2626 | No |
| 152 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7856 | -0.020 | 0.2576 | No |
| 153 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 7908 | -0.021 | 0.2543 | No |
| 154 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 8124 | -0.026 | 0.2387 | No |
| 155 | CPOX | CPOX Entrez, Source | coproporphyrinogen oxidase | 8432 | -0.033 | 0.2163 | No |
| 156 | AOX1 | AOX1 Entrez, Source | aldehyde oxidase 1 | 8449 | -0.033 | 0.2160 | No |
| 157 | IDH3G | IDH3G Entrez, Source | isocitrate dehydrogenase 3 (NAD+) gamma | 8587 | -0.037 | 0.2066 | No |
| 158 | GPX2 | GPX2 Entrez, Source | glutathione peroxidase 2 (gastrointestinal) | 8597 | -0.037 | 0.2069 | No |
| 159 | CYB5R3 | CYB5R3 Entrez, Source | cytochrome b5 reductase 3 | 8840 | -0.042 | 0.1897 | No |
| 160 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 10024 | -0.075 | 0.1019 | No |
| 161 | CYBRD1 | CYBRD1 Entrez, Source | cytochrome b reductase 1 | 10038 | -0.076 | 0.1030 | No |
| 162 | CYB5R4 | CYB5R4 Entrez, Source | cytochrome b5 reductase 4 | 10050 | -0.076 | 0.1043 | No |
| 163 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10323 | -0.086 | 0.0861 | No |
| 164 | TXNDC1 | TXNDC1 Entrez, Source | thioredoxin domain containing 1 | 10569 | -0.094 | 0.0701 | No |
| 165 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 10723 | -0.099 | 0.0612 | No |
| 166 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 10743 | -0.100 | 0.0626 | No |
| 167 | PLOD3 | PLOD3 Entrez, Source | procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3 | 10940 | -0.109 | 0.0507 | No |
| 168 | FDX1 | FDX1 Entrez, Source | ferredoxin 1 | 10969 | -0.110 | 0.0517 | No |
| 169 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 11000 | -0.112 | 0.0526 | No |
| 170 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 11033 | -0.113 | 0.0533 | No |
| 171 | P4HA1 | P4HA1 Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide I | 11102 | -0.117 | 0.0514 | No |
| 172 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 11173 | -0.120 | 0.0494 | No |
| 173 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 11363 | -0.129 | 0.0387 | No |
| 174 | NUDT5 | NUDT5 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 5 | 11488 | -0.137 | 0.0331 | No |
| 175 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 11707 | -0.150 | 0.0207 | No |
| 176 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 11903 | -0.166 | 0.0106 | No |
| 177 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 11937 | -0.168 | 0.0128 | No |
| 178 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 11983 | -0.171 | 0.0141 | No |
| 179 | ALDH3B1 | ALDH3B1 Entrez, Source | aldehyde dehydrogenase 3 family, member B1 | 12169 | -0.189 | 0.0054 | No |
| 180 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 12291 | -0.203 | 0.0019 | No |
| 181 | COX15 | COX15 Entrez, Source | COX15 homolog, cytochrome c oxidase assembly protein (yeast) | 12707 | -0.266 | -0.0222 | No |
| 182 | GMPR | GMPR Entrez, Source | guanosine monophosphate reductase | 12951 | -0.339 | -0.0312 | No |
| 183 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 12985 | -0.357 | -0.0237 | No |
| 184 | CYBA | CYBA Entrez, Source | cytochrome b-245, alpha polypeptide | 12989 | -0.359 | -0.0139 | No |
| 185 | GPX3 | GPX3 Entrez, Source | glutathione peroxidase 3 (plasma) | 13136 | -0.457 | -0.0122 | No |
| 186 | LOX | LOX Entrez, Source | lysyl oxidase | 13308 | -0.991 | 0.0026 | No |