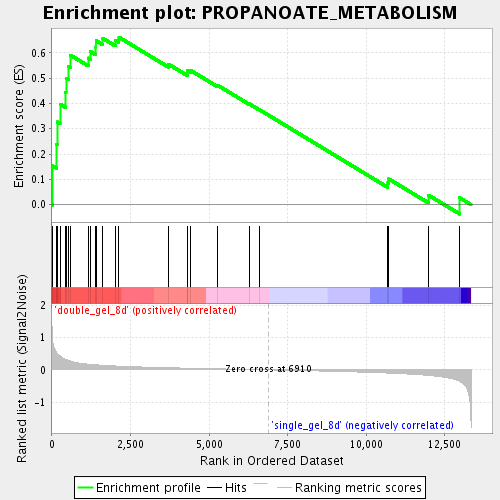
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | PROPANOATE_METABOLISM |
| Enrichment Score (ES) | 0.66144425 |
| Normalized Enrichment Score (NES) | 1.89641 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.005839789 |
| FWER p-Value | 0.478 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 24 | 0.864 | 0.1535 | Yes |
| 2 | ACACA | ACACA Entrez, Source | acetyl-Coenzyme A carboxylase alpha | 154 | 0.527 | 0.2385 | Yes |
| 3 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 172 | 0.499 | 0.3269 | Yes |
| 4 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 260 | 0.427 | 0.3971 | Yes |
| 5 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 430 | 0.329 | 0.4436 | Yes |
| 6 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 450 | 0.320 | 0.4997 | Yes |
| 7 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 524 | 0.292 | 0.5467 | Yes |
| 8 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 584 | 0.271 | 0.5910 | Yes |
| 9 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1158 | 0.175 | 0.5795 | Yes |
| 10 | SDS | SDS Entrez, Source | serine dehydratase | 1222 | 0.169 | 0.6051 | Yes |
| 11 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1399 | 0.157 | 0.6200 | Yes |
| 12 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 1403 | 0.157 | 0.6480 | Yes |
| 13 | PCCB | PCCB Entrez, Source | propionyl Coenzyme A carboxylase, beta polypeptide | 1620 | 0.142 | 0.6572 | Yes |
| 14 | ACADL | ACADL Entrez, Source | acyl-Coenzyme A dehydrogenase, long chain | 2024 | 0.121 | 0.6487 | Yes |
| 15 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 2134 | 0.117 | 0.6614 | Yes |
| 16 | LDHC | LDHC Entrez, Source | lactate dehydrogenase C | 3707 | 0.066 | 0.5553 | No |
| 17 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 4305 | 0.053 | 0.5199 | No |
| 18 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4310 | 0.053 | 0.5291 | No |
| 19 | SUCLG2 | SUCLG2 Entrez, Source | succinate-CoA ligase, GDP-forming, beta subunit | 4416 | 0.050 | 0.5302 | No |
| 20 | SUCLG1 | SUCLG1 Entrez, Source | succinate-CoA ligase, GDP-forming, alpha subunit | 5260 | 0.033 | 0.4728 | No |
| 21 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6279 | 0.012 | 0.3985 | No |
| 22 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 6602 | 0.006 | 0.3754 | No |
| 23 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 10696 | -0.098 | 0.0857 | No |
| 24 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 10723 | -0.099 | 0.1016 | No |
| 25 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 11983 | -0.171 | 0.0379 | No |
| 26 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 12985 | -0.357 | 0.0268 | No |

