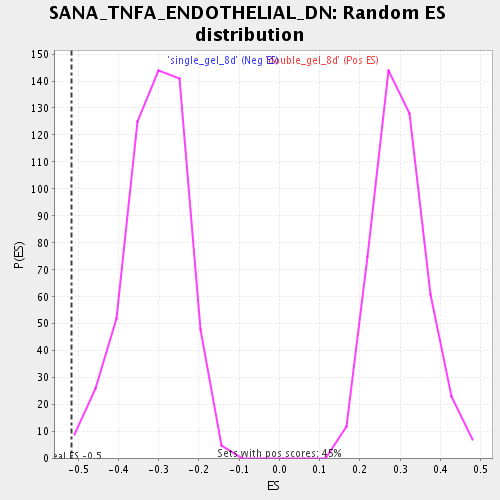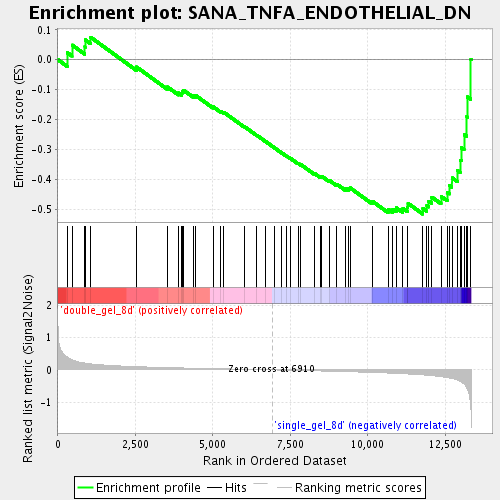
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | SANA_TNFA_ENDOTHELIAL_DN |
| Enrichment Score (ES) | -0.5163454 |
| Normalized Enrichment Score (NES) | -1.6705161 |
| Nominal p-value | 0.0072727273 |
| FDR q-value | 0.08672215 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 303 | 0.396 | 0.0235 | No |
| 2 | IGFBP4 | IGFBP4 Entrez, Source | insulin-like growth factor binding protein 4 | 456 | 0.317 | 0.0491 | No |
| 3 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 871 | 0.211 | 0.0426 | No |
| 4 | PCTP | PCTP Entrez, Source | phosphatidylcholine transfer protein | 882 | 0.210 | 0.0664 | No |
| 5 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 1058 | 0.187 | 0.0750 | No |
| 6 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 2535 | 0.102 | -0.0242 | No |
| 7 | CALCRL | CALCRL Entrez, Source | calcitonin receptor-like | 3533 | 0.070 | -0.0910 | No |
| 8 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 3895 | 0.062 | -0.1109 | No |
| 9 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 4003 | 0.059 | -0.1121 | No |
| 10 | ACE | ACE Entrez, Source | angiotensin I converting enzyme (peptidyl-dipeptidase A) 1 | 4009 | 0.059 | -0.1055 | No |
| 11 | DNAH11 | DNAH11 Entrez, Source | dynein, axonemal, heavy chain 11 | 4055 | 0.058 | -0.1021 | No |
| 12 | VWF | VWF Entrez, Source | von Willebrand factor | 4369 | 0.051 | -0.1197 | No |
| 13 | EMCN | EMCN Entrez, Source | endomucin | 4455 | 0.049 | -0.1203 | No |
| 14 | PGAM2 | PGAM2 Entrez, Source | phosphoglycerate mutase 2 (muscle) | 5008 | 0.038 | -0.1574 | No |
| 15 | APLN | APLN Entrez, Source | apelin, AGTRL1 ligand | 5262 | 0.033 | -0.1727 | No |
| 16 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 5345 | 0.031 | -0.1752 | No |
| 17 | JAG2 | JAG2 Entrez, Source | jagged 2 | 6015 | 0.018 | -0.2234 | No |
| 18 | ANTXR1 | ANTXR1 Entrez, Source | anthrax toxin receptor 1 | 6416 | 0.009 | -0.2524 | No |
| 19 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 6707 | 0.004 | -0.2738 | No |
| 20 | TPT1 | TPT1 Entrez, Source | tumor protein, translationally-controlled 1 | 6974 | -0.002 | -0.2936 | No |
| 21 | ARL5A | ARL5A Entrez, Source | ADP-ribosylation factor-like 5A | 7225 | -0.007 | -0.3117 | No |
| 22 | HTR1B | HTR1B Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 1B | 7386 | -0.010 | -0.3225 | No |
| 23 | KCNH2 | KCNH2 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 2 | 7491 | -0.012 | -0.3289 | No |
| 24 | PRKCH | PRKCH Entrez, Source | protein kinase C, eta | 7749 | -0.018 | -0.3462 | No |
| 25 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 7842 | -0.020 | -0.3509 | No |
| 26 | KDR | KDR Entrez, Source | kinase insert domain receptor (a type III receptor tyrosine kinase) | 8279 | -0.029 | -0.3802 | No |
| 27 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 8464 | -0.033 | -0.3902 | No |
| 28 | RPS3A | RPS3A Entrez, Source | ribosomal protein S3A | 8514 | -0.035 | -0.3898 | No |
| 29 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 8756 | -0.040 | -0.4033 | No |
| 30 | RHOJ | RHOJ Entrez, Source | ras homolog gene family, member J | 8997 | -0.046 | -0.4160 | No |
| 31 | NT5E | NT5E Entrez, Source | 5'-nucleotidase, ecto (CD73) | 9276 | -0.054 | -0.4306 | No |
| 32 | EFEMP1 | EFEMP1 Entrez, Source | EGF-containing fibulin-like extracellular matrix protein 1 | 9375 | -0.057 | -0.4314 | No |
| 33 | ARL6IP1 | ARL6IP1 Entrez, Source | ADP-ribosylation factor-like 6 interacting protein 1 | 9427 | -0.058 | -0.4285 | No |
| 34 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 10142 | -0.079 | -0.4729 | No |
| 35 | CAPNS1 | CAPNS1 Entrez, Source | calpain, small subunit 1 | 10660 | -0.097 | -0.5005 | Yes |
| 36 | TMSB4X | TMSB4X Entrez, Source | thymosin, beta 4, X-linked | 10806 | -0.103 | -0.4993 | Yes |
| 37 | BGN | BGN Entrez, Source | biglycan | 10918 | -0.108 | -0.4951 | Yes |
| 38 | PDLIM4 | PDLIM4 Entrez, Source | PDZ and LIM domain 4 | 11133 | -0.119 | -0.4974 | Yes |
| 39 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 11278 | -0.125 | -0.4935 | Yes |
| 40 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 11298 | -0.126 | -0.4802 | Yes |
| 41 | CLU | CLU Entrez, Source | clusterin | 11779 | -0.155 | -0.4982 | Yes |
| 42 | SMYD2 | SMYD2 Entrez, Source | SET and MYND domain containing 2 | 11907 | -0.166 | -0.4884 | Yes |
| 43 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 11966 | -0.170 | -0.4729 | Yes |
| 44 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 12064 | -0.179 | -0.4593 | Yes |
| 45 | SDPR | SDPR Entrez, Source | serum deprivation response (phosphatidylserine binding protein) | 12367 | -0.211 | -0.4574 | Yes |
| 46 | PALMD | PALMD Entrez, Source | palmdelphin | 12567 | -0.238 | -0.4445 | Yes |
| 47 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 12630 | -0.250 | -0.4200 | Yes |
| 48 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 12719 | -0.269 | -0.3952 | Yes |
| 49 | PEA15 | PEA15 Entrez, Source | phosphoprotein enriched in astrocytes 15 | 12899 | -0.321 | -0.3711 | Yes |
| 50 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 12985 | -0.357 | -0.3358 | Yes |
| 51 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 13022 | -0.377 | -0.2945 | Yes |
| 52 | DDR1 | DDR1 Entrez, Source | discoidin domain receptor family, member 1 | 13111 | -0.436 | -0.2502 | Yes |
| 53 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 13196 | -0.560 | -0.1911 | Yes |
| 54 | CRIP2 | CRIP2 Entrez, Source | cysteine-rich protein 2 | 13214 | -0.582 | -0.1243 | Yes |
| 55 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13327 | -1.146 | 0.0011 | Yes |