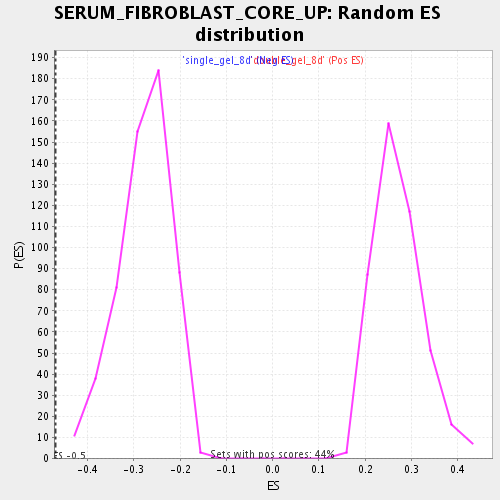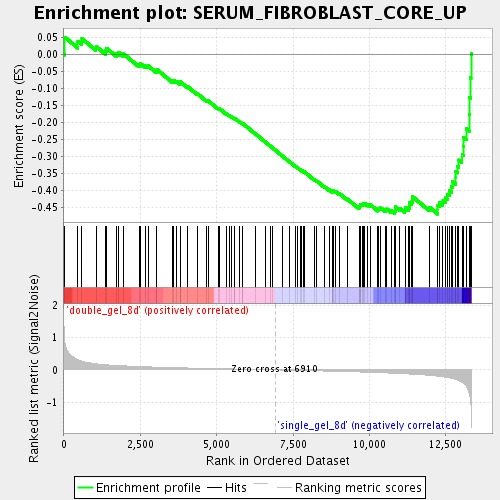
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | SERUM_FIBROBLAST_CORE_UP |
| Enrichment Score (ES) | -0.47053644 |
| Normalized Enrichment Score (NES) | -1.6934742 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0777283 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAP3K8 | MAP3K8 Entrez, Source | mitogen-activated protein kinase kinase kinase 8 | 34 | 0.817 | 0.0499 | No |
| 2 | DHFR | DHFR Entrez, Source | dihydrofolate reductase | 457 | 0.317 | 0.0383 | No |
| 3 | SAR1B | SAR1B Entrez, Source | SAR1 gene homolog B (S. cerevisiae) | 588 | 0.271 | 0.0459 | No |
| 4 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 1050 | 0.187 | 0.0230 | No |
| 5 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 1363 | 0.160 | 0.0097 | No |
| 6 | EIF4EBP1 | EIF4EBP1 Entrez, Source | eukaryotic translation initiation factor 4E binding protein 1 | 1395 | 0.157 | 0.0175 | No |
| 7 | MKKS | MKKS Entrez, Source | McKusick-Kaufman syndrome | 1728 | 0.135 | 0.0011 | No |
| 8 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 1785 | 0.132 | 0.0053 | No |
| 9 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 1940 | 0.124 | 0.0017 | No |
| 10 | LYAR | LYAR Entrez, Source | - | 2462 | 0.104 | -0.0310 | No |
| 11 | MCTS1 | MCTS1 Entrez, Source | malignant T cell amplified sequence 1 | 2515 | 0.102 | -0.0284 | No |
| 12 | TPRKB | TPRKB Entrez, Source | TP53RK binding protein | 2675 | 0.096 | -0.0342 | No |
| 13 | SLC25A5 | SLC25A5 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 | 2756 | 0.094 | -0.0342 | No |
| 14 | MRPL37 | MRPL37 Entrez, Source | mitochondrial ribosomal protein L37 | 3031 | 0.084 | -0.0495 | No |
| 15 | NCLN | NCLN Entrez, Source | nicalin homolog (zebrafish) | 3045 | 0.084 | -0.0452 | No |
| 16 | FARSLB | FARSLB Entrez, Source | phenylalanine-tRNA synthetase-like, beta subunit | 3561 | 0.069 | -0.0796 | No |
| 17 | FLNC | FLNC Entrez, Source | filamin C, gamma (actin binding protein 280) | 3577 | 0.069 | -0.0763 | No |
| 18 | SSR3 | SSR3 Entrez, Source | signal sequence receptor, gamma (translocon-associated protein gamma) | 3688 | 0.067 | -0.0803 | No |
| 19 | HNRPA3 | HNRPA3 Entrez, Source | heterogeneous nuclear ribonucleoprotein A3 | 3804 | 0.064 | -0.0849 | No |
| 20 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 3808 | 0.064 | -0.0810 | No |
| 21 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 4054 | 0.058 | -0.0958 | No |
| 22 | PDIA4 | PDIA4 Entrez, Source | protein disulfide isomerase family A, member 4 | 4372 | 0.051 | -0.1165 | No |
| 23 | MT3 | MT3 Entrez, Source | metallothionein 3 (growth inhibitory factor (neurotrophic)) | 4680 | 0.044 | -0.1368 | No |
| 24 | NUDT1 | NUDT1 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 1 | 4723 | 0.043 | -0.1372 | No |
| 25 | CENPJ | CENPJ Entrez, Source | centromere protein J | 5047 | 0.037 | -0.1592 | No |
| 26 | SNRPA | SNRPA Entrez, Source | small nuclear ribonucleoprotein polypeptide A | 5088 | 0.036 | -0.1599 | No |
| 27 | TFPI2 | TFPI2 Entrez, Source | tissue factor pathway inhibitor 2 | 5317 | 0.032 | -0.1751 | No |
| 28 | TDP1 | TDP1 Entrez, Source | tyrosyl-DNA phosphodiesterase 1 | 5407 | 0.030 | -0.1799 | No |
| 29 | PCSK7 | PCSK7 Entrez, Source | proprotein convertase subtilisin/kexin type 7 | 5492 | 0.028 | -0.1845 | No |
| 30 | COPB2 | COPB2 Entrez, Source | coatomer protein complex, subunit beta 2 (beta prime) | 5578 | 0.026 | -0.1892 | No |
| 31 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 5597 | 0.026 | -0.1889 | No |
| 32 | RAB3B | RAB3B Entrez, Source | RAB3B, member RAS oncogene family | 5758 | 0.023 | -0.1996 | No |
| 33 | RPN1 | RPN1 Entrez, Source | ribophorin I | 5839 | 0.021 | -0.2042 | No |
| 34 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 5852 | 0.021 | -0.2038 | No |
| 35 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 6264 | 0.013 | -0.2340 | No |
| 36 | TOMM40 | TOMM40 Entrez, Source | translocase of outer mitochondrial membrane 40 homolog (yeast) | 6598 | 0.006 | -0.2588 | No |
| 37 | PSMA7 | PSMA7 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 7 | 6760 | 0.003 | -0.2708 | No |
| 38 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 6837 | 0.002 | -0.2764 | No |
| 39 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 7149 | -0.005 | -0.2996 | No |
| 40 | GPLD1 | GPLD1 Entrez, Source | glycosylphosphatidylinositol specific phospholipase D1 | 7374 | -0.010 | -0.3159 | No |
| 41 | LYPLA2 | LYPLA2 Entrez, Source | lysophospholipase II | 7583 | -0.014 | -0.3307 | No |
| 42 | NUP35 | NUP35 Entrez, Source | nucleoporin 35kDa | 7632 | -0.015 | -0.3333 | No |
| 43 | EMP2 | EMP2 Entrez, Source | epithelial membrane protein 2 | 7732 | -0.017 | -0.3397 | No |
| 44 | NLN | NLN Entrez, Source | neurolysin (metallopeptidase M3 family) | 7790 | -0.018 | -0.3428 | No |
| 45 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 7849 | -0.020 | -0.3459 | No |
| 46 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 7859 | -0.020 | -0.3453 | No |
| 47 | CDCA4 | CDCA4 Entrez, Source | cell division cycle associated 4 | 8197 | -0.027 | -0.3690 | No |
| 48 | HNRPAB | HNRPAB Entrez, Source | heterogeneous nuclear ribonucleoprotein A/B | 8280 | -0.029 | -0.3733 | No |
| 49 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 8540 | -0.035 | -0.3906 | No |
| 50 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 8677 | -0.038 | -0.3984 | No |
| 51 | WDR77 | WDR77 Entrez, Source | WD repeat domain 77 | 8777 | -0.040 | -0.4033 | No |
| 52 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 8796 | -0.041 | -0.4021 | No |
| 53 | POLE3 | POLE3 Entrez, Source | polymerase (DNA directed), epsilon 3 (p17 subunit) | 8809 | -0.041 | -0.4003 | No |
| 54 | EBNA1BP2 | EBNA1BP2 Entrez, Source | EBNA1 binding protein 2 | 8885 | -0.043 | -0.4032 | No |
| 55 | PSMD12 | PSMD12 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 12 | 9015 | -0.046 | -0.4100 | No |
| 56 | PSMD2 | PSMD2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 2 | 9272 | -0.054 | -0.4259 | No |
| 57 | RNF138 | RNF138 Entrez, Source | ring finger protein 138 | 9660 | -0.064 | -0.4510 | No |
| 58 | GNG11 | GNG11 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 11 | 9666 | -0.064 | -0.4473 | No |
| 59 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 9695 | -0.065 | -0.4452 | No |
| 60 | ACTC1 | ACTC1 Entrez, Source | actin, alpha, cardiac muscle 1 | 9699 | -0.065 | -0.4413 | No |
| 61 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 9762 | -0.067 | -0.4416 | No |
| 62 | NUP93 | NUP93 Entrez, Source | nucleoporin 93kDa | 9797 | -0.068 | -0.4399 | No |
| 63 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 9840 | -0.070 | -0.4386 | No |
| 64 | NPTN | NPTN Entrez, Source | neuroplastin | 9949 | -0.073 | -0.4420 | No |
| 65 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 10026 | -0.075 | -0.4429 | No |
| 66 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 10274 | -0.084 | -0.4562 | No |
| 67 | TXNL2 | TXNL2 Entrez, Source | thioredoxin-like 2 | 10307 | -0.085 | -0.4532 | No |
| 68 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 10369 | -0.087 | -0.4522 | No |
| 69 | NUP107 | NUP107 Entrez, Source | nucleoporin 107kDa | 10509 | -0.092 | -0.4569 | No |
| 70 | RNASEH2A | RNASEH2A Entrez, Source | ribonuclease H2, subunit A | 10571 | -0.094 | -0.4554 | No |
| 71 | RNF41 | RNF41 Entrez, Source | ring finger protein 41 | 10715 | -0.099 | -0.4599 | No |
| 72 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 10832 | -0.104 | -0.4620 | Yes |
| 73 | CCT5 | CCT5 Entrez, Source | chaperonin containing TCP1, subunit 5 (epsilon) | 10843 | -0.105 | -0.4560 | Yes |
| 74 | CEP78 | CEP78 Entrez, Source | centrosomal protein 78kDa | 10845 | -0.105 | -0.4494 | Yes |
| 75 | PSMC3 | PSMC3 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 3 | 10989 | -0.111 | -0.4530 | Yes |
| 76 | PFN1 | PFN1 Entrez, Source | profilin 1 | 11162 | -0.120 | -0.4584 | Yes |
| 77 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 11178 | -0.120 | -0.4518 | Yes |
| 78 | DCBLD2 | DCBLD2 Entrez, Source | discoidin, CUB and LCCL domain containing 2 | 11288 | -0.126 | -0.4519 | Yes |
| 79 | SNRPB | SNRPB Entrez, Source | small nuclear ribonucleoprotein polypeptides B and B1 | 11304 | -0.127 | -0.4449 | Yes |
| 80 | HNRPR | HNRPR Entrez, Source | heterogeneous nuclear ribonucleoprotein R | 11314 | -0.127 | -0.4375 | Yes |
| 81 | RPP40 | RPP40 Entrez, Source | ribonuclease P 40kDa subunit | 11387 | -0.130 | -0.4345 | Yes |
| 82 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 11400 | -0.131 | -0.4270 | Yes |
| 83 | DCK | DCK Entrez, Source | deoxycytidine kinase | 11406 | -0.131 | -0.4190 | Yes |
| 84 | NUPL1 | NUPL1 Entrez, Source | nucleoporin like 1 | 11959 | -0.169 | -0.4498 | Yes |
| 85 | MAPRE1 | MAPRE1 Entrez, Source | microtubule-associated protein, RP/EB family, member 1 | 12234 | -0.197 | -0.4579 | Yes |
| 86 | ACTL6A | ACTL6A Entrez, Source | actin-like 6A | 12236 | -0.197 | -0.4453 | Yes |
| 87 | CCDC5 | CCDC5 Entrez, Source | coiled-coil domain containing 5 (spindle associated) | 12285 | -0.202 | -0.4360 | Yes |
| 88 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12405 | -0.215 | -0.4311 | Yes |
| 89 | DCLRE1B | DCLRE1B Entrez, Source | DNA cross-link repair 1B (PSO2 homolog, S. cerevisiae) | 12486 | -0.225 | -0.4227 | Yes |
| 90 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 12545 | -0.235 | -0.4120 | Yes |
| 91 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 12611 | -0.247 | -0.4011 | Yes |
| 92 | CLCF1 | CLCF1 Entrez, Source | cardiotrophin-like cytokine factor 1 | 12679 | -0.260 | -0.3895 | Yes |
| 93 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 12719 | -0.269 | -0.3751 | Yes |
| 94 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12821 | -0.294 | -0.3639 | Yes |
| 95 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12824 | -0.294 | -0.3452 | Yes |
| 96 | STX3 | STX3 Entrez, Source | syntaxin 3 | 12884 | -0.315 | -0.3294 | Yes |
| 97 | MET | MET Entrez, Source | met proto-oncogene (hepatocyte growth factor receptor) | 12926 | -0.331 | -0.3113 | Yes |
| 98 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13028 | -0.381 | -0.2945 | Yes |
| 99 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13077 | -0.411 | -0.2718 | Yes |
| 100 | SMS | SMS Entrez, Source | spermine synthase | 13083 | -0.414 | -0.2456 | Yes |
| 101 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 13170 | -0.508 | -0.2195 | Yes |
| 102 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13271 | -0.763 | -0.1781 | Yes |
| 103 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 13282 | -0.795 | -0.1278 | Yes |
| 104 | MSN | MSN Entrez, Source | moesin | 13302 | -0.941 | -0.0688 | Yes |
| 105 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13324 | -1.119 | 0.0014 | Yes |