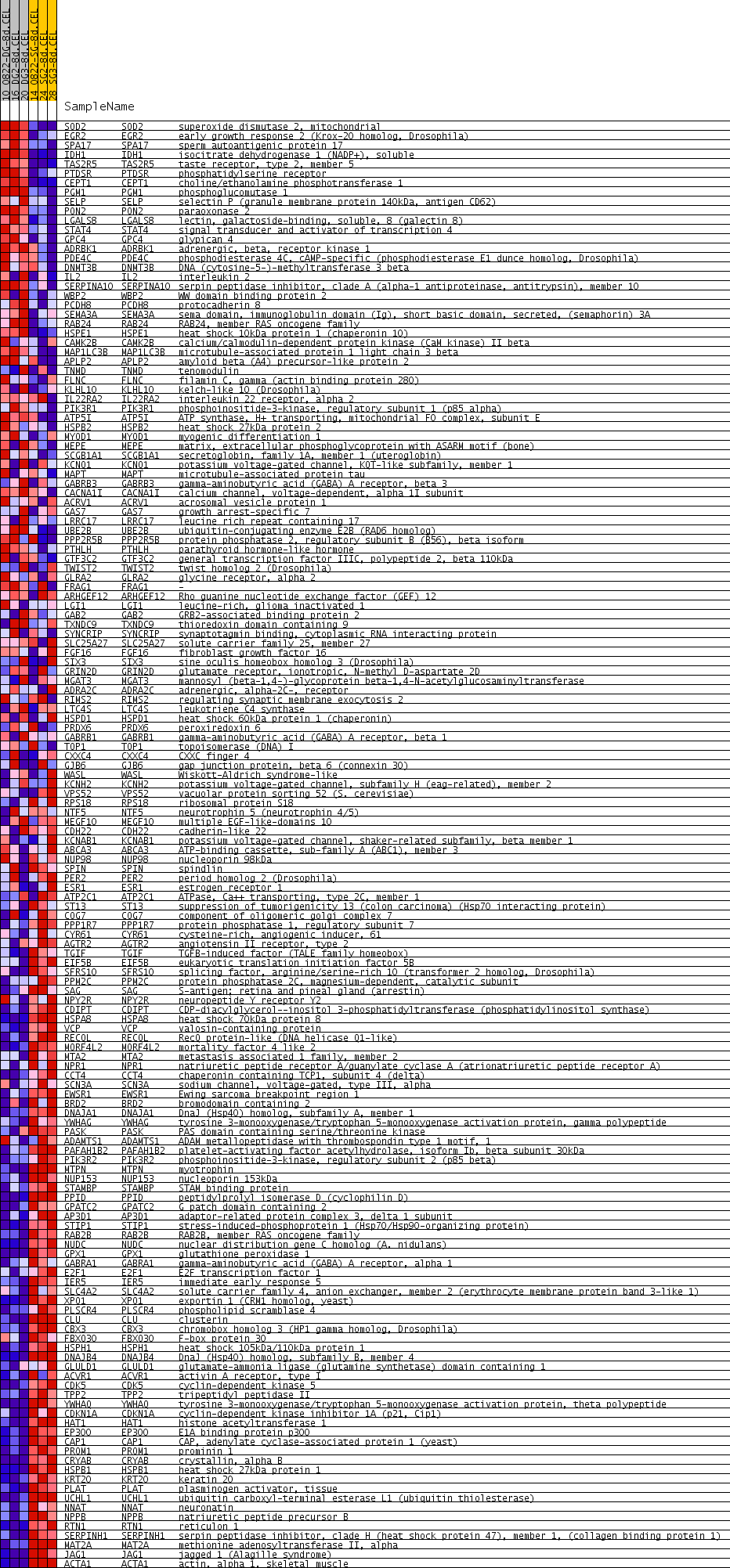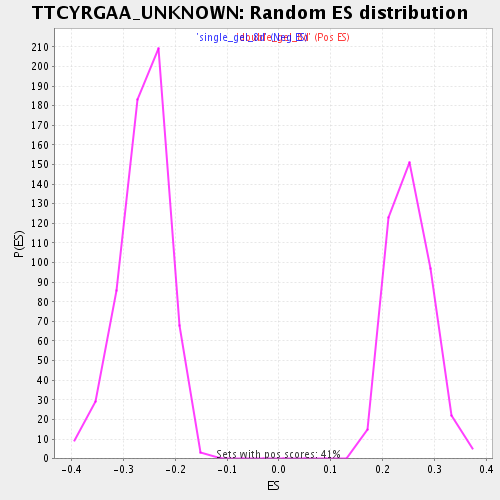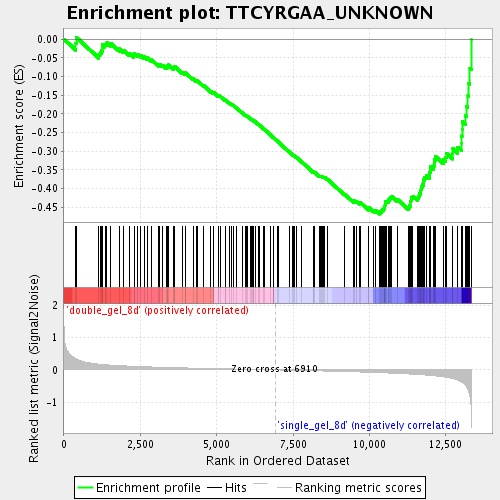
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | TTCYRGAA_UNKNOWN |
| Enrichment Score (ES) | -0.46836916 |
| Normalized Enrichment Score (NES) | -1.7992496 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04816173 |
| FWER p-Value | 0.916 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 391 | 0.343 | -0.0118 | No |
| 2 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 402 | 0.338 | 0.0051 | No |
| 3 | SPA17 | SPA17 Entrez, Source | sperm autoantigenic protein 17 | 1145 | 0.177 | -0.0419 | No |
| 4 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 1183 | 0.172 | -0.0358 | No |
| 5 | TAS2R5 | TAS2R5 Entrez, Source | taste receptor, type 2, member 5 | 1224 | 0.169 | -0.0300 | No |
| 6 | PTDSR | PTDSR Entrez, Source | phosphatidylserine receptor | 1255 | 0.167 | -0.0235 | No |
| 7 | CEPT1 | CEPT1 Entrez, Source | choline/ethanolamine phosphotransferase 1 | 1257 | 0.167 | -0.0149 | No |
| 8 | PGM1 | PGM1 Entrez, Source | phosphoglucomutase 1 | 1344 | 0.161 | -0.0130 | No |
| 9 | SELP | SELP Entrez, Source | selectin P (granule membrane protein 140kDa, antigen CD62) | 1408 | 0.156 | -0.0096 | No |
| 10 | PON2 | PON2 Entrez, Source | paraoxonase 2 | 1531 | 0.147 | -0.0112 | No |
| 11 | LGALS8 | LGALS8 Entrez, Source | lectin, galactoside-binding, soluble, 8 (galectin 8) | 1816 | 0.130 | -0.0259 | No |
| 12 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 1945 | 0.124 | -0.0292 | No |
| 13 | GPC4 | GPC4 Entrez, Source | glypican 4 | 2133 | 0.117 | -0.0372 | No |
| 14 | ADRBK1 | ADRBK1 Entrez, Source | adrenergic, beta, receptor kinase 1 | 2293 | 0.111 | -0.0435 | No |
| 15 | PDE4C | PDE4C Entrez, Source | phosphodiesterase 4C, cAMP-specific (phosphodiesterase E1 dunce homolog, Drosophila) | 2298 | 0.110 | -0.0381 | No |
| 16 | DNMT3B | DNMT3B Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 beta | 2399 | 0.107 | -0.0401 | No |
| 17 | IL2 | IL2 Entrez, Source | interleukin 2 | 2505 | 0.102 | -0.0427 | No |
| 18 | SERPINA10 | SERPINA10 Entrez, Source | serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 | 2629 | 0.098 | -0.0469 | No |
| 19 | WBP2 | WBP2 Entrez, Source | WW domain binding protein 2 | 2728 | 0.095 | -0.0494 | No |
| 20 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 2869 | 0.089 | -0.0554 | No |
| 21 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 3092 | 0.082 | -0.0680 | No |
| 22 | RAB24 | RAB24 Entrez, Source | RAB24, member RAS oncogene family | 3137 | 0.080 | -0.0671 | No |
| 23 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 3227 | 0.078 | -0.0698 | No |
| 24 | CAMK2B | CAMK2B Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II beta | 3351 | 0.075 | -0.0752 | No |
| 25 | MAP1LC3B | MAP1LC3B Entrez, Source | microtubule-associated protein 1 light chain 3 beta | 3380 | 0.074 | -0.0735 | No |
| 26 | APLP2 | APLP2 Entrez, Source | amyloid beta (A4) precursor-like protein 2 | 3389 | 0.074 | -0.0702 | No |
| 27 | TNMD | TNMD Entrez, Source | tenomodulin | 3422 | 0.073 | -0.0688 | No |
| 28 | FLNC | FLNC Entrez, Source | filamin C, gamma (actin binding protein 280) | 3577 | 0.069 | -0.0769 | No |
| 29 | KLHL10 | KLHL10 Entrez, Source | kelch-like 10 (Drosophila) | 3602 | 0.069 | -0.0751 | No |
| 30 | IL22RA2 | IL22RA2 Entrez, Source | interleukin 22 receptor, alpha 2 | 3612 | 0.068 | -0.0722 | No |
| 31 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 3876 | 0.062 | -0.0889 | No |
| 32 | ATP5I | ATP5I Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit E | 3963 | 0.061 | -0.0923 | No |
| 33 | HSPB2 | HSPB2 Entrez, Source | heat shock 27kDa protein 2 | 3966 | 0.060 | -0.0893 | No |
| 34 | MYOD1 | MYOD1 Entrez, Source | myogenic differentiation 1 | 4224 | 0.054 | -0.1059 | No |
| 35 | MEPE | MEPE Entrez, Source | matrix, extracellular phosphoglycoprotein with ASARM motif (bone) | 4322 | 0.052 | -0.1106 | No |
| 36 | SCGB1A1 | SCGB1A1 Entrez, Source | secretoglobin, family 1A, member 1 (uteroglobin) | 4373 | 0.051 | -0.1117 | No |
| 37 | KCNQ1 | KCNQ1 Entrez, Source | potassium voltage-gated channel, KQT-like subfamily, member 1 | 4581 | 0.047 | -0.1249 | No |
| 38 | MAPT | MAPT Entrez, Source | microtubule-associated protein tau | 4812 | 0.042 | -0.1402 | No |
| 39 | GABRB3 | GABRB3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 3 | 4900 | 0.040 | -0.1447 | No |
| 40 | CACNA1I | CACNA1I Entrez, Source | calcium channel, voltage-dependent, alpha 1I subunit | 4910 | 0.040 | -0.1433 | No |
| 41 | ACRV1 | ACRV1 Entrez, Source | acrosomal vesicle protein 1 | 5049 | 0.037 | -0.1519 | No |
| 42 | GAS7 | GAS7 Entrez, Source | growth arrest-specific 7 | 5073 | 0.036 | -0.1517 | No |
| 43 | LRRC17 | LRRC17 Entrez, Source | leucine rich repeat containing 17 | 5108 | 0.036 | -0.1524 | No |
| 44 | UBE2B | UBE2B Entrez, Source | ubiquitin-conjugating enzyme E2B (RAD6 homolog) | 5274 | 0.032 | -0.1632 | No |
| 45 | PPP2R5B | PPP2R5B Entrez, Source | protein phosphatase 2, regulatory subunit B (B56), beta isoform | 5292 | 0.032 | -0.1629 | No |
| 46 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 5428 | 0.029 | -0.1716 | No |
| 47 | GTF3C2 | GTF3C2 Entrez, Source | general transcription factor IIIC, polypeptide 2, beta 110kDa | 5481 | 0.028 | -0.1740 | No |
| 48 | TWIST2 | TWIST2 Entrez, Source | twist homolog 2 (Drosophila) | 5544 | 0.027 | -0.1773 | No |
| 49 | GLRA2 | GLRA2 Entrez, Source | glycine receptor, alpha 2 | 5658 | 0.024 | -0.1846 | No |
| 50 | FRAG1 | FRAG1 Entrez, Source | - | 5837 | 0.021 | -0.1970 | No |
| 51 | ARHGEF12 | ARHGEF12 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 12 | 5936 | 0.020 | -0.2034 | No |
| 52 | LGI1 | LGI1 Entrez, Source | leucine-rich, glioma inactivated 1 | 5982 | 0.019 | -0.2059 | No |
| 53 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 6013 | 0.018 | -0.2072 | No |
| 54 | TXNDC9 | TXNDC9 Entrez, Source | thioredoxin domain containing 9 | 6096 | 0.016 | -0.2126 | No |
| 55 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 6146 | 0.015 | -0.2155 | No |
| 56 | SLC25A27 | SLC25A27 Entrez, Source | solute carrier family 25, member 27 | 6177 | 0.015 | -0.2170 | No |
| 57 | FGF16 | FGF16 Entrez, Source | fibroblast growth factor 16 | 6207 | 0.014 | -0.2184 | No |
| 58 | SIX3 | SIX3 Entrez, Source | sine oculis homeobox homolog 3 (Drosophila) | 6220 | 0.014 | -0.2186 | No |
| 59 | GRIN2D | GRIN2D Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2D | 6262 | 0.013 | -0.2211 | No |
| 60 | MGAT3 | MGAT3 Entrez, Source | mannosyl (beta-1,4-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase | 6360 | 0.010 | -0.2279 | No |
| 61 | ADRA2C | ADRA2C Entrez, Source | adrenergic, alpha-2C-, receptor | 6396 | 0.010 | -0.2300 | No |
| 62 | RIMS2 | RIMS2 Entrez, Source | regulating synaptic membrane exocytosis 2 | 6544 | 0.007 | -0.2408 | No |
| 63 | LTC4S | LTC4S Entrez, Source | leukotriene C4 synthase | 6566 | 0.007 | -0.2421 | No |
| 64 | HSPD1 | HSPD1 Entrez, Source | heat shock 60kDa protein 1 (chaperonin) | 6772 | 0.003 | -0.2575 | No |
| 65 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 6850 | 0.001 | -0.2632 | No |
| 66 | GABRB1 | GABRB1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 1 | 6864 | 0.001 | -0.2642 | No |
| 67 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 6869 | 0.001 | -0.2644 | No |
| 68 | CXXC4 | CXXC4 Entrez, Source | CXXC finger 4 | 6999 | -0.002 | -0.2741 | No |
| 69 | GJB6 | GJB6 Entrez, Source | gap junction protein, beta 6 (connexin 30) | 7028 | -0.003 | -0.2761 | No |
| 70 | WASL | WASL Entrez, Source | Wiskott-Aldrich syndrome-like | 7397 | -0.010 | -0.3034 | No |
| 71 | KCNH2 | KCNH2 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 2 | 7491 | -0.012 | -0.3099 | No |
| 72 | VPS52 | VPS52 Entrez, Source | vacuolar protein sorting 52 (S. cerevisiae) | 7507 | -0.013 | -0.3103 | No |
| 73 | RPS18 | RPS18 Entrez, Source | ribosomal protein S18 | 7553 | -0.014 | -0.3130 | No |
| 74 | NTF5 | NTF5 Entrez, Source | neurotrophin 5 (neurotrophin 4/5) | 7611 | -0.015 | -0.3166 | No |
| 75 | MEGF10 | MEGF10 Entrez, Source | multiple EGF-like-domains 10 | 7779 | -0.018 | -0.3283 | No |
| 76 | CDH22 | CDH22 Entrez, Source | cadherin-like 22 | 8169 | -0.027 | -0.3564 | No |
| 77 | KCNAB1 | KCNAB1 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, beta member 1 | 8172 | -0.027 | -0.3552 | No |
| 78 | ABCA3 | ABCA3 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 3 | 8214 | -0.028 | -0.3568 | No |
| 79 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 8376 | -0.032 | -0.3674 | No |
| 80 | SPIN | SPIN Entrez, Source | spindlin | 8397 | -0.032 | -0.3672 | No |
| 81 | PER2 | PER2 Entrez, Source | period homolog 2 (Drosophila) | 8421 | -0.033 | -0.3672 | No |
| 82 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 8446 | -0.033 | -0.3673 | No |
| 83 | ATP2C1 | ATP2C1 Entrez, Source | ATPase, Ca++ transporting, type 2C, member 1 | 8499 | -0.034 | -0.3694 | No |
| 84 | ST13 | ST13 Entrez, Source | suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) | 8530 | -0.035 | -0.3699 | No |
| 85 | COG7 | COG7 Entrez, Source | component of oligomeric golgi complex 7 | 8620 | -0.037 | -0.3747 | No |
| 86 | PPP1R7 | PPP1R7 Entrez, Source | protein phosphatase 1, regulatory subunit 7 | 9184 | -0.051 | -0.4147 | No |
| 87 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 9477 | -0.059 | -0.4338 | No |
| 88 | AGTR2 | AGTR2 Entrez, Source | angiotensin II receptor, type 2 | 9505 | -0.060 | -0.4327 | No |
| 89 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 9563 | -0.061 | -0.4338 | No |
| 90 | EIF5B | EIF5B Entrez, Source | eukaryotic translation initiation factor 5B | 9668 | -0.064 | -0.4383 | No |
| 91 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 9695 | -0.065 | -0.4369 | No |
| 92 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 9971 | -0.074 | -0.4539 | No |
| 93 | SAG | SAG Entrez, Source | S-antigen; retina and pineal gland (arrestin) | 9972 | -0.074 | -0.4501 | No |
| 94 | NPY2R | NPY2R Entrez, Source | neuropeptide Y receptor Y2 | 10141 | -0.079 | -0.4587 | No |
| 95 | CDIPT | CDIPT Entrez, Source | CDP-diacylglycerol--inositol 3-phosphatidyltransferase (phosphatidylinositol synthase) | 10202 | -0.081 | -0.4590 | No |
| 96 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 10327 | -0.086 | -0.4639 | Yes |
| 97 | VCP | VCP Entrez, Source | valosin-containing protein | 10359 | -0.087 | -0.4617 | Yes |
| 98 | RECQL | RECQL Entrez, Source | RecQ protein-like (DNA helicase Q1-like) | 10390 | -0.088 | -0.4594 | Yes |
| 99 | MORF4L2 | MORF4L2 Entrez, Source | mortality factor 4 like 2 | 10413 | -0.089 | -0.4564 | Yes |
| 100 | MTA2 | MTA2 Entrez, Source | metastasis associated 1 family, member 2 | 10450 | -0.090 | -0.4545 | Yes |
| 101 | NPR1 | NPR1 Entrez, Source | natriuretic peptide receptor A/guanylate cyclase A (atrionatriuretic peptide receptor A) | 10478 | -0.091 | -0.4518 | Yes |
| 102 | CCT4 | CCT4 Entrez, Source | chaperonin containing TCP1, subunit 4 (delta) | 10483 | -0.091 | -0.4474 | Yes |
| 103 | SCN3A | SCN3A Entrez, Source | sodium channel, voltage-gated, type III, alpha | 10506 | -0.091 | -0.4442 | Yes |
| 104 | EWSR1 | EWSR1 Entrez, Source | Ewing sarcoma breakpoint region 1 | 10512 | -0.092 | -0.4398 | Yes |
| 105 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 10513 | -0.092 | -0.4350 | Yes |
| 106 | DNAJA1 | DNAJA1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 1 | 10557 | -0.094 | -0.4334 | Yes |
| 107 | YWHAG | YWHAG Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide | 10619 | -0.096 | -0.4330 | Yes |
| 108 | PASK | PASK Entrez, Source | PAS domain containing serine/threonine kinase | 10632 | -0.096 | -0.4289 | Yes |
| 109 | ADAMTS1 | ADAMTS1 Entrez, Source | ADAM metallopeptidase with thrombospondin type 1 motif, 1 | 10664 | -0.097 | -0.4262 | Yes |
| 110 | PAFAH1B2 | PAFAH1B2 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, beta subunit 30kDa | 10691 | -0.098 | -0.4230 | Yes |
| 111 | PIK3R2 | PIK3R2 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 2 (p85 beta) | 10731 | -0.100 | -0.4208 | Yes |
| 112 | MTPN | MTPN Entrez, Source | myotrophin | 10904 | -0.107 | -0.4283 | Yes |
| 113 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 11280 | -0.126 | -0.4501 | Yes |
| 114 | STAMBP | STAMBP Entrez, Source | STAM binding protein | 11308 | -0.127 | -0.4456 | Yes |
| 115 | PPID | PPID Entrez, Source | peptidylprolyl isomerase D (cyclophilin D) | 11332 | -0.128 | -0.4406 | Yes |
| 116 | GPATC2 | GPATC2 Entrez, Source | G patch domain containing 2 | 11333 | -0.128 | -0.4340 | Yes |
| 117 | AP3D1 | AP3D1 Entrez, Source | adaptor-related protein complex 3, delta 1 subunit | 11360 | -0.129 | -0.4292 | Yes |
| 118 | STIP1 | STIP1 Entrez, Source | stress-induced-phosphoprotein 1 (Hsp70/Hsp90-organizing protein) | 11377 | -0.130 | -0.4236 | Yes |
| 119 | RAB2B | RAB2B Entrez, Source | RAB2B, member RAS oncogene family | 11421 | -0.132 | -0.4200 | Yes |
| 120 | NUDC | NUDC Entrez, Source | nuclear distribution gene C homolog (A. nidulans) | 11586 | -0.143 | -0.4250 | Yes |
| 121 | GPX1 | GPX1 Entrez, Source | glutathione peroxidase 1 | 11620 | -0.145 | -0.4199 | Yes |
| 122 | GABRA1 | GABRA1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 1 | 11632 | -0.146 | -0.4131 | Yes |
| 123 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 11673 | -0.148 | -0.4084 | Yes |
| 124 | IER5 | IER5 Entrez, Source | immediate early response 5 | 11685 | -0.149 | -0.4015 | Yes |
| 125 | SLC4A2 | SLC4A2 Entrez, Source | solute carrier family 4, anion exchanger, member 2 (erythrocyte membrane protein band 3-like 1) | 11703 | -0.150 | -0.3950 | Yes |
| 126 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 11725 | -0.151 | -0.3887 | Yes |
| 127 | PLSCR4 | PLSCR4 Entrez, Source | phospholipid scramblase 4 | 11764 | -0.153 | -0.3836 | Yes |
| 128 | CLU | CLU Entrez, Source | clusterin | 11779 | -0.155 | -0.3766 | Yes |
| 129 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 11802 | -0.156 | -0.3701 | Yes |
| 130 | FBXO30 | FBXO30 Entrez, Source | F-box protein 30 | 11851 | -0.160 | -0.3653 | Yes |
| 131 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 11957 | -0.169 | -0.3645 | Yes |
| 132 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 11966 | -0.170 | -0.3562 | Yes |
| 133 | GLULD1 | GLULD1 Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) domain containing 1 | 11986 | -0.172 | -0.3487 | Yes |
| 134 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 11998 | -0.173 | -0.3405 | Yes |
| 135 | CDK5 | CDK5 Entrez, Source | cyclin-dependent kinase 5 | 12107 | -0.182 | -0.3392 | Yes |
| 136 | TPP2 | TPP2 Entrez, Source | tripeptidyl peptidase II | 12117 | -0.183 | -0.3303 | Yes |
| 137 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 12130 | -0.184 | -0.3216 | Yes |
| 138 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 12163 | -0.188 | -0.3143 | Yes |
| 139 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 12419 | -0.217 | -0.3223 | Yes |
| 140 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 12496 | -0.228 | -0.3161 | Yes |
| 141 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 12522 | -0.231 | -0.3059 | Yes |
| 142 | PROM1 | PROM1 Entrez, Source | prominin 1 | 12722 | -0.269 | -0.3069 | Yes |
| 143 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 12728 | -0.271 | -0.2932 | Yes |
| 144 | HSPB1 | HSPB1 Entrez, Source | heat shock 27kDa protein 1 | 12892 | -0.318 | -0.2890 | Yes |
| 145 | KRT20 | KRT20 Entrez, Source | keratin 20 | 13006 | -0.369 | -0.2783 | Yes |
| 146 | PLAT | PLAT Entrez, Source | plasminogen activator, tissue | 13010 | -0.371 | -0.2591 | Yes |
| 147 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 13032 | -0.382 | -0.2408 | Yes |
| 148 | NNAT | NNAT Entrez, Source | neuronatin | 13037 | -0.385 | -0.2210 | Yes |
| 149 | NPPB | NPPB Entrez, Source | natriuretic peptide precursor B | 13138 | -0.457 | -0.2047 | Yes |
| 150 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 13181 | -0.527 | -0.1803 | Yes |
| 151 | SERPINH1 | SERPINH1 Entrez, Source | serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) | 13222 | -0.606 | -0.1518 | Yes |
| 152 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13252 | -0.684 | -0.1183 | Yes |
| 153 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 13287 | -0.812 | -0.0784 | Yes |
| 154 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 13340 | -1.581 | 0.0002 | Yes |