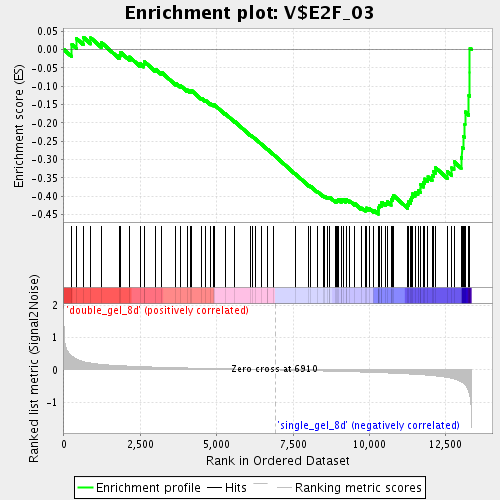
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
| GeneSet | V$E2F_03 |
| Enrichment Score (ES) | -0.44886976 |
| Normalized Enrichment Score (NES) | -1.6264611 |
| Nominal p-value | 0.0036968577 |
| FDR q-value | 0.10557116 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SF4 | SF4 Entrez, Source | splicing factor 4 | 261 | 0.427 | 0.0143 | No |
| 2 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 405 | 0.337 | 0.0303 | No |
| 3 | BMP7 | BMP7 Entrez, Source | bone morphogenetic protein 7 (osteogenic protein 1) | 635 | 0.257 | 0.0336 | No |
| 4 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 880 | 0.210 | 0.0318 | No |
| 5 | UXT | UXT Entrez, Source | ubiquitously-expressed transcript | 1225 | 0.169 | 0.0193 | No |
| 6 | SEZ6 | SEZ6 Entrez, Source | seizure related 6 homolog (mouse) | 1821 | 0.130 | -0.0153 | No |
| 7 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 1853 | 0.129 | -0.0074 | No |
| 8 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 2146 | 0.116 | -0.0201 | No |
| 9 | EGLN2 | EGLN2 Entrez, Source | egl nine homolog 2 (C. elegans) | 2499 | 0.103 | -0.0385 | No |
| 10 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 2621 | 0.098 | -0.0399 | No |
| 11 | PCSK4 | PCSK4 Entrez, Source | proprotein convertase subtilisin/kexin type 4 | 2628 | 0.098 | -0.0325 | No |
| 12 | CTDSP1 | CTDSP1 Entrez, Source | CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 1 | 3001 | 0.085 | -0.0538 | No |
| 13 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 3194 | 0.079 | -0.0620 | No |
| 14 | KCNA6 | KCNA6 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 6 | 3661 | 0.067 | -0.0919 | No |
| 15 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 3805 | 0.064 | -0.0976 | No |
| 16 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 4035 | 0.059 | -0.1102 | No |
| 17 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 4143 | 0.056 | -0.1138 | No |
| 18 | HTF9C | HTF9C Entrez, Source | - | 4181 | 0.055 | -0.1122 | No |
| 19 | GRIA4 | GRIA4 Entrez, Source | glutamate receptor, ionotrophic, AMPA 4 | 4512 | 0.048 | -0.1332 | No |
| 20 | KLHDC3 | KLHDC3 Entrez, Source | kelch domain containing 3 | 4632 | 0.046 | -0.1386 | No |
| 21 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 4801 | 0.042 | -0.1479 | No |
| 22 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 4884 | 0.040 | -0.1509 | No |
| 23 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 4925 | 0.039 | -0.1508 | No |
| 24 | CACNA1G | CACNA1G Entrez, Source | calcium channel, voltage-dependent, alpha 1G subunit | 5281 | 0.032 | -0.1750 | No |
| 25 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 5597 | 0.026 | -0.1968 | No |
| 26 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 6115 | 0.016 | -0.2345 | No |
| 27 | VAMP3 | VAMP3 Entrez, Source | vesicle-associated membrane protein 3 (cellubrevin) | 6158 | 0.015 | -0.2365 | No |
| 28 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 6264 | 0.013 | -0.2434 | No |
| 29 | GLRA3 | GLRA3 Entrez, Source | glycine receptor, alpha 3 | 6466 | 0.008 | -0.2579 | No |
| 30 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 6650 | 0.005 | -0.2713 | No |
| 31 | PLAGL1 | PLAGL1 Entrez, Source | pleiomorphic adenoma gene-like 1 | 6668 | 0.005 | -0.2722 | No |
| 32 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 6869 | 0.001 | -0.2873 | No |
| 33 | PDGFRA | PDGFRA Entrez, Source | platelet-derived growth factor receptor, alpha polypeptide | 7572 | -0.014 | -0.3391 | No |
| 34 | RPS20 | RPS20 Entrez, Source | ribosomal protein S20 | 8019 | -0.024 | -0.3709 | No |
| 35 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 8077 | -0.025 | -0.3732 | No |
| 36 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 8301 | -0.030 | -0.3877 | No |
| 37 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 8494 | -0.034 | -0.3994 | No |
| 38 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 8540 | -0.035 | -0.4000 | No |
| 39 | RPL28 | RPL28 Entrez, Source | ribosomal protein L28 | 8612 | -0.037 | -0.4024 | No |
| 40 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 8679 | -0.039 | -0.4043 | No |
| 41 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 8707 | -0.039 | -0.4033 | No |
| 42 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 8883 | -0.043 | -0.4130 | No |
| 43 | SLC25A11 | SLC25A11 Entrez, Source | solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11 | 8929 | -0.044 | -0.4129 | No |
| 44 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 8949 | -0.045 | -0.4108 | No |
| 45 | NUTF2 | NUTF2 Entrez, Source | nuclear transport factor 2 | 8981 | -0.045 | -0.4095 | No |
| 46 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 9075 | -0.048 | -0.4127 | No |
| 47 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 9082 | -0.048 | -0.4093 | No |
| 48 | PVRL1 | PVRL1 Entrez, Source | poliovirus receptor-related 1 (herpesvirus entry mediator C; nectin) | 9133 | -0.050 | -0.4091 | No |
| 49 | FMO4 | FMO4 Entrez, Source | flavin containing monooxygenase 4 | 9231 | -0.052 | -0.4123 | No |
| 50 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 9233 | -0.052 | -0.4082 | No |
| 51 | FBXL20 | FBXL20 Entrez, Source | F-box and leucine-rich repeat protein 20 | 9357 | -0.056 | -0.4130 | No |
| 52 | DIO3 | DIO3 Entrez, Source | deiodinase, iodothyronine, type III | 9522 | -0.060 | -0.4206 | No |
| 53 | SIAHBP1 | SIAHBP1 Entrez, Source | - | 9746 | -0.066 | -0.4322 | No |
| 54 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 9861 | -0.070 | -0.4352 | No |
| 55 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 9897 | -0.071 | -0.4321 | No |
| 56 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 9999 | -0.075 | -0.4338 | No |
| 57 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 10143 | -0.079 | -0.4383 | No |
| 58 | NR3C2 | NR3C2 Entrez, Source | nuclear receptor subfamily 3, group C, member 2 | 10284 | -0.084 | -0.4422 | Yes |
| 59 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 10287 | -0.084 | -0.4356 | Yes |
| 60 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 10306 | -0.085 | -0.4302 | Yes |
| 61 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 10317 | -0.085 | -0.4242 | Yes |
| 62 | XTP3TPA | XTP3TPA Entrez, Source | - | 10388 | -0.088 | -0.4225 | Yes |
| 63 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 10400 | -0.088 | -0.4163 | Yes |
| 64 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 10538 | -0.093 | -0.4192 | Yes |
| 65 | ATP6V1D | ATP6V1D Entrez, Source | ATPase, H+ transporting, lysosomal 34kDa, V1 subunit D | 10581 | -0.095 | -0.4149 | Yes |
| 66 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 10707 | -0.099 | -0.4164 | Yes |
| 67 | RET | RET Entrez, Source | ret proto-oncogene (multiple endocrine neoplasia and medullary thyroid carcinoma 1, Hirschsprung disease) | 10708 | -0.099 | -0.4086 | Yes |
| 68 | RHD | RHD Entrez, Source | Rh blood group, D antigen | 10757 | -0.101 | -0.4041 | Yes |
| 69 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 10774 | -0.102 | -0.3972 | Yes |
| 70 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 11253 | -0.124 | -0.4234 | Yes |
| 71 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 11280 | -0.126 | -0.4154 | Yes |
| 72 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 11345 | -0.129 | -0.4100 | Yes |
| 73 | ARF3 | ARF3 Entrez, Source | ADP-ribosylation factor 3 | 11384 | -0.130 | -0.4025 | Yes |
| 74 | DCK | DCK Entrez, Source | deoxycytidine kinase | 11406 | -0.131 | -0.3936 | Yes |
| 75 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 11492 | -0.137 | -0.3891 | Yes |
| 76 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 11589 | -0.143 | -0.3850 | Yes |
| 77 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 11672 | -0.148 | -0.3794 | Yes |
| 78 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 11673 | -0.148 | -0.3676 | Yes |
| 79 | KLF5 | KLF5 Entrez, Source | Kruppel-like factor 5 (intestinal) | 11774 | -0.155 | -0.3628 | Yes |
| 80 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 11811 | -0.158 | -0.3530 | Yes |
| 81 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 11914 | -0.167 | -0.3474 | Yes |
| 82 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 12050 | -0.177 | -0.3435 | Yes |
| 83 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 12094 | -0.181 | -0.3323 | Yes |
| 84 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 12163 | -0.188 | -0.3225 | Yes |
| 85 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 12545 | -0.235 | -0.3325 | Yes |
| 86 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 12695 | -0.263 | -0.3229 | Yes |
| 87 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 12767 | -0.278 | -0.3061 | Yes |
| 88 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 12999 | -0.364 | -0.2946 | Yes |
| 89 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13028 | -0.381 | -0.2664 | Yes |
| 90 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13077 | -0.411 | -0.2373 | Yes |
| 91 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 13120 | -0.443 | -0.2052 | Yes |
| 92 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13128 | -0.448 | -0.1701 | Yes |
| 93 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13252 | -0.684 | -0.1249 | Yes |
| 94 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13288 | -0.823 | -0.0620 | Yes |
| 95 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13289 | -0.828 | 0.0040 | Yes |

