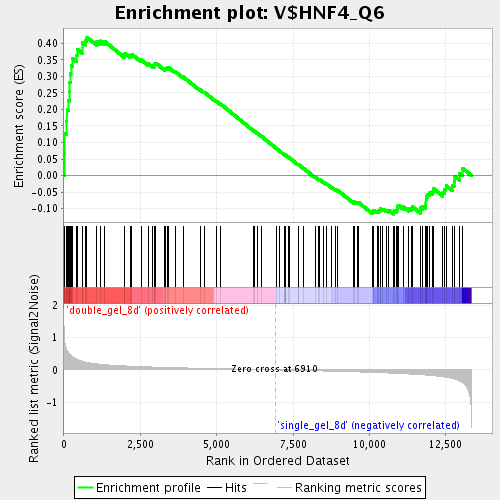
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$HNF4_Q6 |
| Enrichment Score (ES) | 0.4184282 |
| Normalized Enrichment Score (NES) | 1.5633863 |
| Nominal p-value | 0.008733625 |
| FDR q-value | 0.08666921 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 10 | 1.121 | 0.0660 | Yes |
| 2 | HFE2 | HFE2 Entrez, Source | hemochromatosis type 2 (juvenile) | 13 | 1.053 | 0.1286 | Yes |
| 3 | LCAT | LCAT Entrez, Source | lecithin-cholesterol acyltransferase | 76 | 0.677 | 0.1642 | Yes |
| 4 | RARB | RARB Entrez, Source | retinoic acid receptor, beta | 101 | 0.614 | 0.1990 | Yes |
| 5 | SERPIND1 | SERPIND1 Entrez, Source | serpin peptidase inhibitor, clade D (heparin cofactor), member 1 | 144 | 0.544 | 0.2282 | Yes |
| 6 | PKLR | PKLR Entrez, Source | pyruvate kinase, liver and RBC | 184 | 0.487 | 0.2543 | Yes |
| 7 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 189 | 0.483 | 0.2828 | Yes |
| 8 | NR5A2 | NR5A2 Entrez, Source | nuclear receptor subfamily 5, group A, member 2 | 220 | 0.457 | 0.3077 | Yes |
| 9 | AGXT2 | AGXT2 Entrez, Source | alanine-glyoxylate aminotransferase 2 | 233 | 0.448 | 0.3335 | Yes |
| 10 | ITGA1 | ITGA1 Entrez, Source | integrin, alpha 1 | 278 | 0.416 | 0.3549 | Yes |
| 11 | AGMAT | AGMAT Entrez, Source | agmatine ureohydrolase (agmatinase) | 424 | 0.331 | 0.3637 | Yes |
| 12 | SLC22A7 | SLC22A7 Entrez, Source | solute carrier family 22 (organic anion transporter), member 7 | 437 | 0.326 | 0.3822 | Yes |
| 13 | ERBB3 | ERBB3 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 3 (avian) | 595 | 0.269 | 0.3864 | Yes |
| 14 | FOXO1A | FOXO1A Entrez, Source | forkhead box O1A (rhabdomyosarcoma) | 596 | 0.268 | 0.4024 | Yes |
| 15 | FXC1 | FXC1 Entrez, Source | fracture callus 1 homolog (rat) | 696 | 0.239 | 0.4091 | Yes |
| 16 | GAMT | GAMT Entrez, Source | guanidinoacetate N-methyltransferase | 753 | 0.227 | 0.4184 | Yes |
| 17 | NFE2 | NFE2 Entrez, Source | nuclear factor (erythroid-derived 2), 45kDa | 1071 | 0.185 | 0.4055 | No |
| 18 | XPNPEP2 | XPNPEP2 Entrez, Source | X-prolyl aminopeptidase (aminopeptidase P) 2, membrane-bound | 1185 | 0.172 | 0.4072 | No |
| 19 | NOSTRIN | NOSTRIN Entrez, Source | nitric oxide synthase trafficker | 1330 | 0.162 | 0.4060 | No |
| 20 | TCF1 | TCF1 Entrez, Source | transcription factor 1, hepatic; LF-B1, hepatic nuclear factor (HNF1), albumin proximal factor | 1988 | 0.122 | 0.3636 | No |
| 21 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 1995 | 0.122 | 0.3704 | No |
| 22 | C9 | C9 Entrez, Source | complement component 9 | 2167 | 0.116 | 0.3644 | No |
| 23 | PROS1 | PROS1 Entrez, Source | protein S (alpha) | 2217 | 0.114 | 0.3675 | No |
| 24 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 2527 | 0.102 | 0.3502 | No |
| 25 | SLC25A5 | SLC25A5 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 | 2756 | 0.094 | 0.3385 | No |
| 26 | INSR | INSR Entrez, Source | insulin receptor | 2897 | 0.088 | 0.3332 | No |
| 27 | HPCA | HPCA Entrez, Source | hippocalcin | 2962 | 0.086 | 0.3335 | No |
| 28 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 2976 | 0.086 | 0.3376 | No |
| 29 | CACNA2D3 | CACNA2D3 Entrez, Source | calcium channel, voltage-dependent, alpha 2/delta 3 subunit | 3006 | 0.085 | 0.3404 | No |
| 30 | USH1C | USH1C Entrez, Source | Usher syndrome 1C (autosomal recessive, severe) | 3303 | 0.076 | 0.3226 | No |
| 31 | PDZK1 | PDZK1 Entrez, Source | PDZ domain containing 1 | 3333 | 0.075 | 0.3249 | No |
| 32 | PTPRH | PTPRH Entrez, Source | protein tyrosine phosphatase, receptor type, H | 3376 | 0.074 | 0.3261 | No |
| 33 | SLC7A8 | SLC7A8 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 8 | 3436 | 0.073 | 0.3260 | No |
| 34 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 3644 | 0.068 | 0.3144 | No |
| 35 | SFRS15 | SFRS15 Entrez, Source | splicing factor, arginine/serine-rich 15 | 3902 | 0.062 | 0.2987 | No |
| 36 | CREB3L2 | CREB3L2 Entrez, Source | cAMP responsive element binding protein 3-like 2 | 4457 | 0.049 | 0.2598 | No |
| 37 | DAP3 | DAP3 Entrez, Source | death associated protein 3 | 4590 | 0.047 | 0.2526 | No |
| 38 | SLN | SLN Entrez, Source | sarcolipin | 4980 | 0.038 | 0.2255 | No |
| 39 | LRRC17 | LRRC17 Entrez, Source | leucine rich repeat containing 17 | 5108 | 0.036 | 0.2180 | No |
| 40 | RNF40 | RNF40 Entrez, Source | ring finger protein 40 | 6216 | 0.014 | 0.1352 | No |
| 41 | CAB39L | CAB39L Entrez, Source | calcium binding protein 39-like | 6232 | 0.013 | 0.1348 | No |
| 42 | PELO | PELO Entrez, Source | pelota homolog (Drosophila) | 6323 | 0.011 | 0.1287 | No |
| 43 | AQP3 | AQP3 Entrez, Source | aquaporin 3 (Gill blood group) | 6462 | 0.009 | 0.1188 | No |
| 44 | RND1 | RND1 Entrez, Source | Rho family GTPase 1 | 6944 | -0.001 | 0.0825 | No |
| 45 | NRG2 | NRG2 Entrez, Source | neuregulin 2 | 7064 | -0.003 | 0.0737 | No |
| 46 | TREH | TREH Entrez, Source | trehalase (brush-border membrane glycoprotein) | 7209 | -0.006 | 0.0632 | No |
| 47 | ARFIP2 | ARFIP2 Entrez, Source | ADP-ribosylation factor interacting protein 2 (arfaptin 2) | 7224 | -0.007 | 0.0626 | No |
| 48 | SLC5A2 | SLC5A2 Entrez, Source | solute carrier family 5 (sodium/glucose cotransporter), member 2 | 7237 | -0.007 | 0.0621 | No |
| 49 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 7239 | -0.007 | 0.0625 | No |
| 50 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 7256 | -0.008 | 0.0617 | No |
| 51 | MRPL2 | MRPL2 Entrez, Source | mitochondrial ribosomal protein L2 | 7336 | -0.009 | 0.0563 | No |
| 52 | EFNB1 | EFNB1 Entrez, Source | ephrin-B1 | 7378 | -0.010 | 0.0537 | No |
| 53 | PAK1 | PAK1 Entrez, Source | p21/Cdc42/Rac1-activated kinase 1 (STE20 homolog, yeast) | 7664 | -0.016 | 0.0332 | No |
| 54 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 7666 | -0.016 | 0.0340 | No |
| 55 | FUT2 | FUT2 Entrez, Source | fucosyltransferase 2 (secretor status included) | 7831 | -0.019 | 0.0228 | No |
| 56 | ZNF403 | ZNF403 Entrez, Source | zinc finger protein 403 | 8229 | -0.028 | -0.0055 | No |
| 57 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 8336 | -0.031 | -0.0117 | No |
| 58 | NAV2 | NAV2 Entrez, Source | neuron navigator 2 | 8365 | -0.032 | -0.0119 | No |
| 59 | MAGED1 | MAGED1 Entrez, Source | melanoma antigen family D, 1 | 8492 | -0.034 | -0.0194 | No |
| 60 | GPX2 | GPX2 Entrez, Source | glutathione peroxidase 2 (gastrointestinal) | 8597 | -0.037 | -0.0251 | No |
| 61 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 8768 | -0.040 | -0.0355 | No |
| 62 | LHX1 | LHX1 Entrez, Source | LIM homeobox 1 | 8884 | -0.043 | -0.0417 | No |
| 63 | GATA4 | GATA4 Entrez, Source | GATA binding protein 4 | 8964 | -0.045 | -0.0449 | No |
| 64 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 9483 | -0.059 | -0.0805 | No |
| 65 | CBX7 | CBX7 Entrez, Source | chromobox homolog 7 | 9510 | -0.060 | -0.0789 | No |
| 66 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 9602 | -0.063 | -0.0821 | No |
| 67 | MOBP | MOBP Entrez, Source | myelin-associated oligodendrocyte basic protein | 9647 | -0.064 | -0.0816 | No |
| 68 | ASB2 | ASB2 Entrez, Source | ankyrin repeat and SOCS box-containing 2 | 10090 | -0.078 | -0.1104 | No |
| 69 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 10123 | -0.079 | -0.1081 | No |
| 70 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 10142 | -0.079 | -0.1047 | No |
| 71 | HTATIP | HTATIP Entrez, Source | HIV-1 Tat interacting protein, 60kDa | 10248 | -0.083 | -0.1077 | No |
| 72 | APC | APC Entrez, Source | adenomatosis polyposis coli | 10296 | -0.085 | -0.1062 | No |
| 73 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 10345 | -0.086 | -0.1047 | No |
| 74 | PTMS | PTMS Entrez, Source | parathymosin | 10355 | -0.087 | -0.1002 | No |
| 75 | GRB7 | GRB7 Entrez, Source | growth factor receptor-bound protein 7 | 10441 | -0.090 | -0.1013 | No |
| 76 | RASSF5 | RASSF5 Entrez, Source | Ras association (RalGDS/AF-6) domain family 5 | 10560 | -0.094 | -0.1046 | No |
| 77 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 10638 | -0.097 | -0.1047 | No |
| 78 | JUB | JUB Entrez, Source | jub, ajuba homolog (Xenopus laevis) | 10796 | -0.103 | -0.1104 | No |
| 79 | KIAA1443 | KIAA1443 Entrez, Source | KIAA1443 | 10804 | -0.103 | -0.1048 | No |
| 80 | FUT11 | FUT11 Entrez, Source | fucosyltransferase 11 (alpha (1,3) fucosyltransferase) | 10884 | -0.106 | -0.1045 | No |
| 81 | RAB3D | RAB3D Entrez, Source | RAB3D, member RAS oncogene family | 10905 | -0.107 | -0.0996 | No |
| 82 | SLC9A3R1 | SLC9A3R1 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 1 | 10921 | -0.108 | -0.0943 | No |
| 83 | BMF | BMF Entrez, Source | Bcl2 modifying factor | 10943 | -0.109 | -0.0894 | No |
| 84 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 11105 | -0.117 | -0.0946 | No |
| 85 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 11278 | -0.125 | -0.1002 | No |
| 86 | SPINT1 | SPINT1 Entrez, Source | serine peptidase inhibitor, Kunitz type 1 | 11368 | -0.129 | -0.0992 | No |
| 87 | SP140 | SP140 Entrez, Source | SP140 nuclear body protein | 11402 | -0.131 | -0.0939 | No |
| 88 | ART3 | ART3 Entrez, Source | ADP-ribosyltransferase 3 | 11663 | -0.148 | -0.1047 | No |
| 89 | RALY | RALY Entrez, Source | RNA binding protein, autoantigenic (hnRNP-associated with lethal yellow homolog (mouse)) | 11665 | -0.148 | -0.0960 | No |
| 90 | NUMB | NUMB Entrez, Source | numb homolog (Drosophila) | 11741 | -0.151 | -0.0926 | No |
| 91 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 11830 | -0.159 | -0.0898 | No |
| 92 | MMP16 | MMP16 Entrez, Source | matrix metallopeptidase 16 (membrane-inserted) | 11849 | -0.160 | -0.0816 | No |
| 93 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 11850 | -0.160 | -0.0721 | No |
| 94 | ZDHHC3 | ZDHHC3 Entrez, Source | zinc finger, DHHC-type containing 3 | 11867 | -0.162 | -0.0637 | No |
| 95 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 11900 | -0.165 | -0.0563 | No |
| 96 | KIFC3 | KIFC3 Entrez, Source | kinesin family member C3 | 11973 | -0.171 | -0.0515 | No |
| 97 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 12050 | -0.177 | -0.0467 | No |
| 98 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 12100 | -0.181 | -0.0396 | No |
| 99 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12405 | -0.215 | -0.0497 | No |
| 100 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 12463 | -0.223 | -0.0408 | No |
| 101 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 12504 | -0.229 | -0.0302 | No |
| 102 | TAGLN2 | TAGLN2 Entrez, Source | transgelin 2 | 12708 | -0.266 | -0.0297 | No |
| 103 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 12770 | -0.278 | -0.0177 | No |
| 104 | ADM | ADM Entrez, Source | adrenomedullin | 12796 | -0.285 | -0.0027 | No |
| 105 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12954 | -0.340 | 0.0057 | No |
| 106 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 13047 | -0.395 | 0.0223 | No |

